| Research Article | ||
Open Vet. J.. 2025; 15(12): 6287-6297 Open Veterinary Journal, (2025), Vol. 15(12): 6287-6297 Research Article Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in IndonesiaSheila Marty Yanestria1,2, Tri Untari3*, Sidna Artanto3, Mustofa Helmi Effendi4, Wiwiek Tyasningsih5 and Aswin Rafif Khairullah61Department of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Wijaya Kusuma Surabaya, Surabaya, Indonesia 2Post Doctoral Program in Veterinary Science, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia 3Department of Microbiology, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia 4Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 5Division of Veterinary Microbiology, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 6Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor, Indonesia *Corresponding Author: Tri Untari. Departement of Microbiology Microbiology, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Yogyakarta, Daerah Istimewa Yogyakarta, Indonesia. Email: triuntariugm [at] gmail.com Submitted: 23/08/2025 Revised: 15/11/2025 Accepted: 26/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
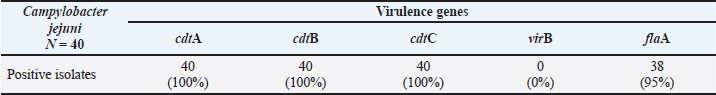
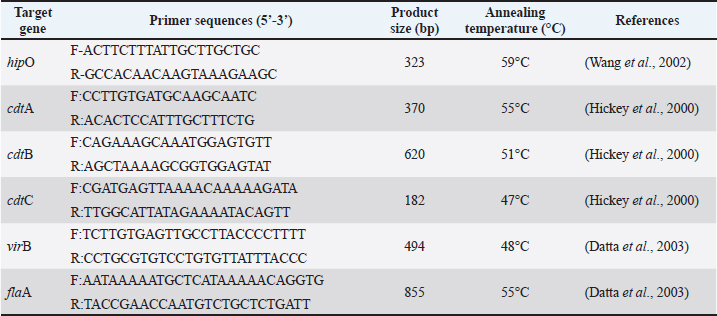
AbstractBackground: Campylobacter jejuni is a major zoonotic pathogen that causes gastroenteritis in humans, with broiler chickens as its primary reservoir. The pathogenicity of this bacterium is highly dependent on virulence factors encoded by specific genes, such as the cytolethal distending toxin (CDT) operon (cdtA, cdtB, and cdtC), the motility gene (flaA), and the invasion-related gene (virB). Aim: This study aimed to identify and determine the prevalence of the virulence genes cdtA, cdtB, cdtC, flaA, and virB in C. jejuni isolates from the small intestine of broiler chickens in Indonesia. Methods: This study used a cross-sectional design on 200 small intestine jejunum samples randomly collected from 200 Cobb broiler chickens in Pasuruan Regency, East Java, between October and December 2023. Bacterial isolation was performed through an enrichment stage using Bolton Broth, followed by cultivation on mCCDA under microaerophilic conditions at 42°C. Confirmation of the C. jejuni species was performed using conventional PCR targeting the hipO gene. Isolates that had been confirmed positive were then further analyzed to detect the presence of five virulence genes (cdtA, cdtB, cdtC, flaA, and virB) using polymerase chain reaction. Results: Isolation and identification results showed that 40 (20%) isolates of C. jejuni were found out of 200 small intestine samples from Cobb broiler chickens at several farms and poultry slaughterhouses in Pasuruan Regency, East Java, Indonesia. Specifically, 100% of the tested isolates carried the three genes that make up the CDT operon: cdtA, cdtB, and cdtC. Furthermore, the flaA gene was almost universal in the isolates studied, with 38 (95%) of the 40 isolates identified as positive. All isolates (100%) tested negative for the virB gene. Conclusion: Campylobacter jejuni isolates from the intestines of broiler chickens in Indonesia contained the cdtA, cdtB, cdtC, and flaA virulence genes, indicating a public health risk. An integrated approach of surveillance, biosecurity, antibiotic management, and research and development of natural therapies is a priority strategy to protect public health and the Indonesian poultry industry’s sustainability. Keywords: Campylobacter, Gene, Virulence, Chicken, Public health. IntroductionGlobally, Campylobacter jejuni is one of the leading causes of human gastroenteritis, with poultry (especially broiler chickens) as its primary reservoir (Khairullah et al., 2024). This infection generally occurs due to the consumption of contaminated poultry products, making the prevalence and virulence of C. jejuni in animal-derived food products a critical issue in food safety and public health (Li et al., 2020a). The prevalence of C. jejuni in broiler chickens has been reported to be very high in various countries, with prevalence rates varying depending on detection methods and geographic location. In Eastern China, Campylobacter prevalence in broiler production reached over 60% (Tang et al., 2020). In Saudi Arabia, the prevalence of C. jejuni in broiler meat reached 26.4%, with isolates showing high levels of MDR (Aljasir and Allam, 2025). Early detection and monitoring of virulence factors in this bacterium are crucial for mitigating the risk of human transmission (Ramandinianto et al., 2020; Amaliah et al., 2023). Virulence factors play a crucial role in the pathogenicity mechanism of C. jejuni. The cdtA, cdtB, and cdtC genes, which form the cytolethal distending toxin (CDT) operon, and the flaA gene, which plays a role in bacterial motility and colonization in the poultry gastrointestinal tract, are important virulence genes (Sierra-Arguello et al., 2021). Furthermore, the virB gene, typically found on plasmids, plays a role in the invasion process and increases the ability of bacteria to cause disease (Elsayed et al., 2019). The combination of these virulence genes detected in broiler chicken isolates can increase the risk of zoonosis and make control more challenging (Hadiyan et al., 2022). Molecular detection techniques, particularly polymerase chain reaction (PCR), are now the primary method for identifying C. jejuni and detecting associated virulence genes. PCR not only offers high sensitivity and specificity but also accelerates the diagnostic time compared with conventional methods (Vizzini et al., 2021; Wanja et al., 2024). Several global surveys have documented the high prevalence of cdtA, cdtB, and cdtC genes in chicken C. jejuni isolates. Many studies have also reported the detection of flaA in nearly all C. jejuni isolates, indicating its important role in bacterial colonization and infectivity (Goualié et al., 2019). However, the prevalence of the virB gene can vary between studies, with some reporting low or even no detection in broiler isolates, reflecting genetic variation in the field (Divya et al., 2025). The distribution pattern of virulence genes is an important indicator for assessing public health threats and the potential for outbreaks of zoonotic infections (Hızlısoy et al., 2020). Virulence gene studies in C. jejuni isolated directly from the small intestine of broiler chickens in Indonesia are not yet widely available. The results of this analysis are expected to provide current and relevant information to enrich local data as a basis for formulating zoonotic campylobacteriosis control policies in Indonesia, as well as supporting efforts to improve food security and public health. However, previous research has been limited to detecting antibiotic resistance in C. jejuni, without considering the virulence potential (Yanestria et al., 2024). Furthermore, this research is expected to contribute to the global literature on the molecular epidemiology of C. jejuni at the regional and international levels (Montero et al., 2024). Therefore, this study aimed to identify and determine the prevalence of important virulence genes, namely cdtA, cdtB, cdtC, flaA, and virB, in these C. jejuni isolates. Materials and MethodsEthical approvalEthical approval was obtained from the Ethical Clearance Committee of the Faculty of Veterinary Medicine, Universitas Wijaya Kusuma Surabaya, Indonesia (Ethics number: 176-KKE/2023). Sample collectionThis study used a cross-sectional design with random sampling of the small intestine jejunum from Cobb broiler chickens. This study was conducted from October 2023 to December 2023. A total of 200 small intestine samples were taken from 200 Cobb broiler chickens at several farms and poultry slaughterhouses in Pasuruan Regency, East Java, Indonesia. Each sample (10 g) was collected in sterile sample bottles containing Nutrient Broth No. 2 (Oxoid, England) and transported to the laboratory. All samples were immediately stored at 4°C and processed to isolate Campylobacter sp. Isolation and identification of Campylobacter sp.The contents of the small intestine amounting to 1 g were homogenized in a dark bottle containing 40 ml of Bolton Broth (Oxoid, England) with 5% lysed sheep blood and Preston supplement (Oxoid, England), ferrous sulfate, sodium metabisulfite, and sodium pyruvate. The samples were incubated at 37°C for 4 hours and then at 42°C for 24 hours using an anaerobic tube containing a CampyGen sachet, which produces 10% CO2, 5% O2, and 85% N2 (Gharst et al., 2013). The small intestine of each chicken was processed using standard methods. The intestine was cultured on selective modified charcoal cefoperazone deoxycholate agar/mCCDA (Oxoid, England) media supplemented with selective antibiotics (Oxoid, England) and incubated microaerophilically at 42°C for 48 hours (Hadiyan et al., 2022). Colonies suspected of being Campylobacter sp. were identified morphologically (macroscopically and microscopically) and biochemically (catalase and oxidase tests) (Alarjani et al., 2021). Molecular characterization of C. jejuniGenomic DNA was extracted from confirmed colonies using a bacterial DNA extraction kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. PCR primers were synthesized by Integrated DNA Technologies (Iowa) (Table 1). Table 1. Primers of virulence genes, annealing temperatures, and PCR product sizes.
Confirmation of the C. jejuni species was performed using conventional PCR to detect the hipO gene, which is a specific gene for C. jejuni. The primer details and cycling conditions are explained in Table 1. The PCR master mix contained 5 μl DNA template, 1 μl each primer, 0.5 μl nuclease-free water, and 12.5 μl PCR master mix (Promega, USA) containing Taq DNA polymerase, dNTPs, MgCl2, and reaction buffer. The final volume of the reaction mixture was 20 μl. The amplification program consisted of primary denaturation at 95°C for 0.5 minutes and 35 cycles with secondary denaturation at 95°C for 0.5 minutes, annealing for 0.5 minutes, extension for 0.5 minutes at 72°C, and final extension for 7 minutes at 72°C. Table 1 lists the annealing temperature and used primer set (Wang et al., 2002). This test also includes positive (K+) and negative (K−) control. The amplification products were analyzed by 1.5% agarose gel electrophoresis (Invitrogen, USA), stained with RedSafe Nucleic Acid Staining Solution gel dye (Intron, South Korea), and documented with a UV transilluminator. Confirmed C. jejuni isolates were characterized for in vitro virulence gene detection by PCR for five known virulence genes: cdtA, cdtB, cdtC, virB, and flaA. The amplification program consisted of 35 cycles of denaturation at 95°C for 1 minute, annealing for 1 min, and elongation for 1 min at 72°C (Gharbi et al., 2022). Table 2 lists the annealing temperature for each gene and the primer sets used. Table 2. Prevalence of virulence genes in Campylobacter jejuni.
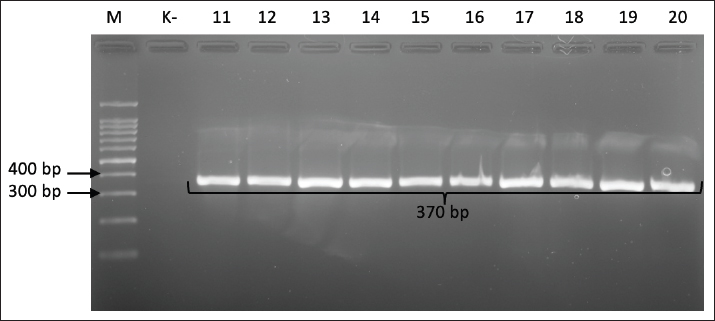
Data analysisThe prevalence of virulence genes in C. jejuni isolates was determined by calculating the proportion of isolates positive for a specific target gene out of the total isolates tested, then multiplying by 100% to obtain a percentage prevalence value. This calculation was applied to each virulence gene studied, such as cdtA, cdtB, cdtC, virB, and flaA, allowing the identification of the most dominant virulence genes and variations in their distribution within the study isolate population. ResultThe results of the isolation of Campylobacter sp. in mCCDA were macroscopically characterized by grayish, moist, and spreading colonies. Microscopically, the Campylobacter sp. isolate was characterized by the characteristics of red, spiral-shaped Gram-negative bacteria. Biochemical tests confirmed that the Campylobacter sp. isolate was positive for catalase and oxidase. Final confirmation using PCR targeting the hipO gene specific for C. jejuni produced a 323 bp DNA fragment, which definitively identified the isolate as C. jejuni. The results of the isolation and identification showed that 40 isolates of C. jejuni were present in the 200 samples of broiler chicken small intestine examined. These results are based on molecular detection of genes encoding CDT, a toxin that causes cell damage and inflammation in the host. Specifically, 100% of the isolates tested carried all three genes that make up the CDT operon: cdtA, cdtB, and cdtC (Table 2). A positive result for each gene is indicated by an amplification product (amplicon) of the predicted size. The presence of the cdtA gene, which encodes the toxin-binding subunit to target cells, was confirmed by the appearance of a 370 bp PCR product (Fig. 1).
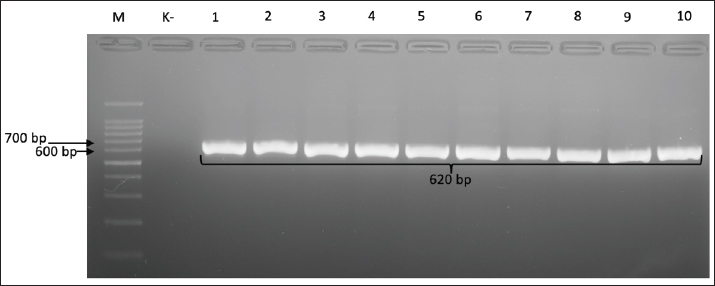
Fig. 1. Gel electrophoresis results for cdtA gene detection (370 bp). Note: Lanes: M, 100-bp marker; K-, control negative; 11-20, representative of Campylobacter jejuni isolates The presence of the cdtB gene, which is the active subunit responsible for host cell DNA damage, was confirmed by the detection of a 620-bp PCR product (Fig. 2).
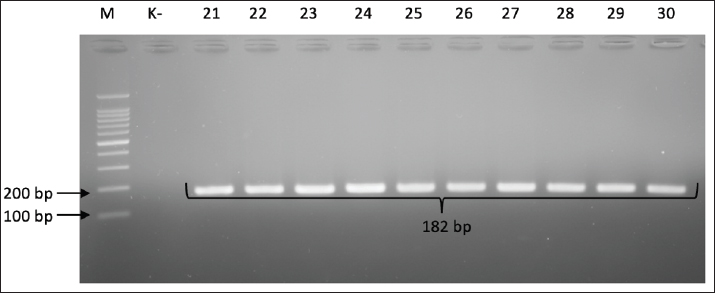
Fig. 2. Gel electrophoresis results for cdtB gene detection (620 bp). Note: Lanes: M, 100-bp marker; K-, control negative; 1-10, representative of Campylobacter jejuni isolates. The presence of the cdtC gene, which also functions as a binding subunit, was confirmed by the presence of a 182-bp PCR product (Fig. 3).
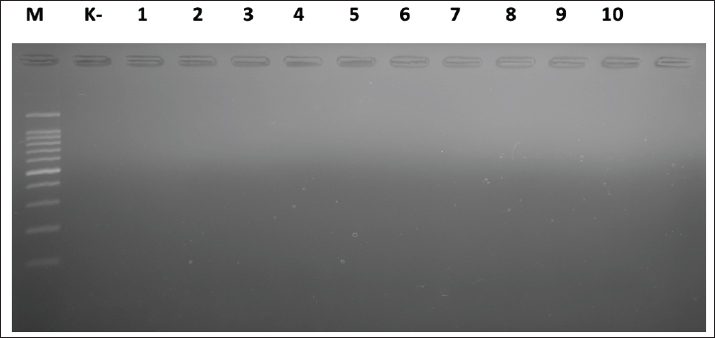
Fig. 3. Gel electrophoresis results for cdtC gene detection (182 bp). Note: Lanes: M, 100-bp marker; K-, control negative; 21-30, representative of Campylobacter jejuni isolates. The test results showed that of the 40 C. jejuni isolates examined, all (100%) gave negative results for the presence of the virB gene. The absence of this virB gene was proven through PCR, where the absence of a DNA amplification product at a specific size (494 bp) indicated the absence of the target gene from the bacterial genome. Figure 4 visually confirms the absence of the expected DNA band for all tested isolates, indicating that this pathogenic pathway may not be present in the circulating strains in the sample source.
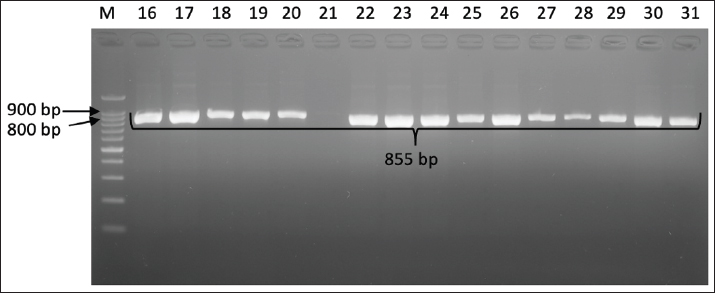
Fig. 4. Gel electrophoresis results for virB gene detection (494 bp) Note: Lanes: M, 100-bp marker; K-, control negative; 1-10, representative of Campylobacter jejuni isolates The results showed that the flaA gene was universal in the isolates studied, with 38 (95%) of the 40 isolates identified as positive. This positive confirmation was based on successful DNA amplification using PCR techniques, which specifically produced an 855 bp DNA fragment for each isolate carrying the gene (Fig. 5). This high prevalence rate confirms that flagella-mediated motility is a dominant virulence strategy and is maintained by almost all C. jejuni populations studied.
Fig. 5. Gel electrophoresis results for flaA gene (855 bp) detection Note: Lanes: M, 100-bp marker; K-, control negative; 16-31, representative of Campylobacter jejuni isolates DiscussionForty isolates of C. jejuni were identified from 200 broiler small intestine samples examined. This indicates a 20% prevalence rate of C. jejuni in broilers in the study area. This finding aligns with a previous study in Indonesia, which reported a C. jejuni prevalence of approximately 23% in broilers based on the hipO gene PCR test, underscoring the importance of surveillance in the Indonesian poultry sector (Yanestria et al., 2023). This prevalence is also within the range found in other regions such as Egypt, where the C. jejuni isolation rate in poultry samples was reported to be approximately 37% (Barakat et al., 2020). This percentage reflects a significant prevalence rate of pathogenic bacteria that potentially pose a zoonotic risk to humans, especially considering that poultry is the primary reservoir of C. jejuni, the leading cause of campylobacteriosis in humans (Amjad, 2023). Several other studies from Algeria and Kenya have reported a high prevalence of C. jejuni in broiler chickens, emphasizing variations in prevalence rates due to differences in detection methods, sample sizes, and geographic factors, with annual prevalence rates reaching over 70% at some sampling points (Baali et al., 2020; Wanja et al., 2024). Campylobacter sp. is a very important zoonotic bacterium that is a major source of foodborne illness worldwide, and its contamination is often found in undercooked poultry products (Shaltout and Shaltout, 2024). In several studies, PCR targeting the hipO gene has been widely used to identify C. jejuni in both chicken samples and its derivatives (Syarifah et al., 2020; Alarjani et al., 2021). The use of selective media and microaerophilic atmospheric conditions during the isolation process has been shown to be effective for the growth of this bacterium, thus strengthening the validity of the findings of similar studies (Prayudi et al., 2023). Confirmation of the specific identity of C. jejuni through PCR targeting the hipO gene is crucial because this gene encodes the enzyme hippurate oxidase. Amplification of a fragment size specific to hipO (approximately 323 bp or varying depending on the primer used) has been used as a standard in many molecular studies (Alarjani et al., 2021; Liu et al., 2022). The use of specific primers for this gene allows for the rapid and accurate detection of isolates obtained from poultry laboratory cultures (Shams et al., 2017). All tested C. jejuni isolates (100%) were positive for the cdtA, cdtB, and cdtC genes. These genes are part of the CDT operon, which produces a toxin that can cause DNA damage and apoptosis in host cells. CDT is an important virulence factor that causes cell distension, cell cycle block, and DNA damage that can ultimately lead to cell death (Pons et al., 2019). The simultaneous presence of all three cdt genes indicates the high pathogenic potential of the C. jejuni isolates found in this study, given the key roles of the cdtA, cdtB, and cdtC genes in the synthesis and function of the CDT toxin. CDT toxin has been shown to contribute to various clinical manifestations of Campylobacter sp. infection, including diarrhea, Guillain-Barré syndrome, and reactive arthritis, which are serious systemic complications associated with Campylobacter sp. infection (Wanja et al., 2024). Recent studies have also shown that CDT toxin can trigger a chronic inflammatory response, exacerbate tissue damage, and induce a cellular senescence phenotype that worsens the clinical condition (Faïs et al., 2016; Shenker et al., 2024). This consistent cdt virulence profile highlights the potential public health hazard of Campylobacter sp. strains circulating in Indonesian poultry, especially since poultry are the main reservoir and source of Campylobacter transmission to humans (Wysok et al., 2020). These findings are consistent with those of other studies in Asia and globally, where approximately 90%–100% of chicken isolates were reported to carry the cdtA, cdtB, and cdtC genes. For example, studies from China, Japan, and Korea reported a near-universal prevalence of the cdt gene in chicken isolates (Kim et al., 2019a; Li et al., 2020b; Saito et al., 2021). This confirms that these toxin genes are not only genetically conservative but also can serve as key markers of C. jejuni virulence. The presence of the cdt gene can be a key indicator in epidemiological studies and the development of infection mitigation strategies (Zhang et al., 2022; Nguyen et al., 2023). In contrast, virB was not detected in any of the C. jejuni isolates. The virB gene is a key component of the T4SS, which transfers DNA and protein molecules from bacteria to host cells or other bacteria for conjugation and modulation of the microbial environment (Drehkopf et al., 2023). This system also plays a crucial role in pathogenicity mechanisms in various Gram-negative pathogenic bacteria, including some Campylobacter strains, by facilitating the transfer of genetic elements or virulence effectors that can influence the host immune response (Matys et al., 2020). However, in this study, all C. jejuni isolates examined did not show the presence of the virB gene. The absence or abnormality of the virB gene in C. jejuni isolates may indicate several factors. First, other pathogenicity mechanisms, such as the type III secretion system (T3SS) or type VI secretion system (T6SS), adhesion factors, motility factors, and toxin factors, may be more dominant in these isolates (Kim et al., 2019b). This is also supported by research findings showing that not all virulence in C. jejuni depends on the presence of the T4SS or virB but also involves a combination of various other genes, such as flaA, cadF, cdtA, cdtB, and cdtC (Lima et al., 2022). Second, there is the potential for virB gene sequence variations that prevent detection by conventional PCR primers, necessitating the development of primers with broader coverage or alternative detection methods such as whole-genome sequencing (Awad et al., 2023). Previous studies have also shown that the virB gene distribution in Campylobacter sp. isolates varies significantly across locations and isolate sources. For example, Awad et al. (2023) found virB11 in only a small proportion of Campylobacter isolates from poultry farms, suggesting that this factor is not uniformly distributed and may not be a major determinant of virulence (Awad et al., 2023). A similar finding was also noted by Kim et al. (2019a), who did not consistently find virB11 in all virulent isolates, indicating the importance of alternative factors in pathogenicity. These findings are consistent with those of Montero et al. (2024), who stated that the presence of the T4SS system is not always essential for pathogenicity in all C. jejuni strains, although it has the potential to support horizontal transfer and genetic adaptation. A total of 38 of 40 C. jejuni isolates, 38 (95%) were positive for flaA. The flaA gene encodes flagellin, the main protein of flagella, which is essential for Campylobacter motility and attachment to host cells. Flagella also play a role in biofilm formation and intestinal colonization (Abbar et al., 2019). The high prevalence of flaA is consistent with the important role of motility in the pathogenesis of C. jejuni (Wysok et al., 2020). Flagella-mediated motility allows these bacteria to move toward and interact with the intestinal mucosa, thereby enhancing colonization and causing more effective infection. In a recent study, the post-transcriptional regulator CsrA was found to regulate the expression of the flagellin protein flaA by binding to flaA mRNA and repressing its translation, demonstrating the complexity of flaA gene expression regulation that contributes to C. jejuni virulence (Abbar et al., 2019). This regulation is important for modulating motility and bacterial adaptation to the host environment. Furthermore, flaA gene mutations have been shown to decrease the colonization and virulence of C. jejuni in the chicken intestine, confirming that flagellar motility is a major virulence factor for this bacterium (Abbar et al., 2019). The high prevalence of flaA in C. jejuni isolates supports the fundamental role of motility in pathogenesis and the effectiveness of bacterial infection in hosts at risk of infection, such as chickens and humans (Igwaran and Okoh, 2020). Compared with data from Southeast Asian countries such as Malaysia, Thailand, and Vietnam, the frequency of the flaA gene in chicken isolates is also high, with figures ranging from 90% to 98%. For example, a study in the Philippines reported a flaA gene frequency of 95% in C. jejuni isolates, indicating conservation and a crucial role for this gene in bacterial virulence in avian hosts, as well as being an indicator for assessing virulence potential and developing molecular-based therapies (Subejano and Penuliar, 2023). In South Korea, the frequency of flaA was also found to be very high in chicken isolates, supporting the prediction of this gene’s role in bacterial virulence and adaptation (Gahamanyi et al., 2021). Studies in several Southeast Asian countries have shown that the flaA gene has stable genetic conservation and plays a key role in bacterial motility and colonization, which are important in the disease mechanism in poultry (Melo et al., 2019; Gahamanyi et al., 2021). Other studies supporting the genetic conservation of flaA in Southeast Asia reported that this gene is frequently used as a molecular target in virulence assessment and vaccine development strategies due to its critical role in the pathogenesis of C. jejuni (Cayrou et al., 2021; Tawakol et al., 2023). These findings demonstrate that flaA detection is highly relevant as an indicator of virulence assessment and gene-based therapy. Thus, the high frequency of flaA gene presence in chicken isolates in Southeast Asia, ranging from 90% to 98%, indicates stable genetic conservation and its important role in bacterial adaptation and virulence in avian hosts, as revealed by various studies in the region. Overall, the virulence gene profiles of C. jejuni isolates from the intestines of broiler chickens in Indonesia indicate that these isolates have pathogenic potential, primarily due to the universal presence of the cdtA, cdtB, and cdtC genes, as well as the high prevalence of the flaA gene. These results underscore the importance of implementing effective control measures to reduce the prevalence of Campylobacter sp. in broiler chickens and minimize the risk of human infection (Hadiyan et al., 2022). Antimicrobial resistance in Campylobacter is also a growing concern, and studies in Indonesia have reported the presence of beta-lactam (blaOXA-61) (Yanestria et al., 2023), tetracycline (tetO), and fluoroquinolone (gyrA) resistance genes (Yanestria et al., 2024). Further research is needed to characterize other virulence factors, understand the pathogenic mechanisms of C. jejuni in Indonesia, and monitor trends in antimicrobial resistance. The virulence gene profile of C. jejuni isolates from the intestines of broilers in Indonesia has crucial implications for the national poultry industry, food safety regulations, and public health policies. In the poultry industry, the high prevalence of these virulence genes indicates an immediate need to strengthen biosecurity and sanitation practices in the production chain from farm to slaughter to prevent the spread of pathogenic strains with potential zoonotic potential (Li et al., 2023). Implementing microbiological surveillance that adopts molecular methods to detect virulence genes and antibiotic resistance is crucial for the early identification and management of pathogen risks. From a food safety regulatory perspective, these findings require revision of food microbial testing standards and protocols based not only on bacterial counts but also on the presence of virulence genes, so that risk assessments become more targeted and compatible with international standards such as Codex Alimentarius. Public health policies should also be directed at increasing consumer and industry awareness of the zoonotic risks of campylobacteriosis and the importance of rational antibiotic use in chicken farming to reduce AMR, which impacts human clinical therapy. Furthermore, the integration of One Health policies involving the agriculture, animal health, and human health sectors is essential for strengthening national preparedness in addressing zoonotic threats and improving epidemiological response systems. Overall, these findings underscore the urgency of developing integrated policies that combine strict surveillance, education, strengthened regulations, and further research on alternative virulence mechanisms and natural-based control measures, such as eugenol, to improve food safety and public health in Indonesia (Ramatla et al., 2022; Subajeno and Penuliar, 2023). ConclusionAnalysis of the virulence profiles of C. jejuni isolates from the intestines of broiler chickens in Indonesia showed that 100% of the isolates contained the cdtA, cdtB, and cdtC genes, and 95% of the isolates contained the flaA gene, indicating a public health risk. This is the first report on the virulence gene profiles of C. jejuni from the intestines of broilers in Indonesia, highlighting the urgent need for an effective surveillance system to detect highly pathogenic strains and the implementation of stricter biosecurity control measures in the poultry sector to reduce the risk of zoonoses to humans. An integrated approach of surveillance, biosecurity, antibiotic management, and research and development of natural therapies is a priority strategy to protect public health and the Indonesian poultry industry’s sustainability. Further research is needed to test the expression and functionality of virulence genes and conduct whole genome sequencing tests to determine the presence of virulence genes not detected in this study and other virulence genes in C. jejuni in more detail. AcknowledgmentsThe authors are thankful to the Faculty of Veterinary Medicine, Gadjah Mada University, for providing the necessary facilities for this study. Conflict of interestThe authors declare no conflict of interest. FundingThis study was supported in part by the Post-Doctoral Program of Gadjah Mada University 2025 Funding from Direktorat Penelitian Universitas Gadjah Mada, with grant number 4454/UNI.P2/Dit-Lit/PT.01.03/2025. Author’s contributionsSMY: Conceived, designed, and coordinated the study. TU: Designed data collection tools, supervised the field sample and data collection, and performed laboratory work and data entry. SA: Validation, supervision, and formal analysis. MHE: Contributed reagents, materials, and analysis tools. ARK and WT: Carried out the statistical analysis and interpretation and participated in the preparation of the manuscript. All authors have read, reviewed, and approved the final version of the manuscript. Data availabilityAll data are available in the revised manuscript. ReferencesAbbar, F.M.E., Li, J., Owen, H.C., Daugherty, C.L., Fulmer, C.A., Bogacz, M. and Thompson, S.A. 2019. RNA Binding by the Campylobacter jejuni Post-transcriptional Regulator CsrA. Front. Microbiol. 10(1), 1776. Alarjani, K.M., Elkhadragy, M.F., Al-Masoud, A.H. and Yehia, H.M. 2021. Detection of Campylobacter jejuni and Salmonella typhimurium in chicken using PCR for virulence factor hipO and invA genes (Saudi Arabia). Biosci. Rep. 41(9), BSR20211790. Aljasir, S.F. and Allam, S.A. 2025. Prevalence and antibiotic resistance of Salmonella spp. and Campylobacter spp. isolated from retail chickens in Saudi Arabia. Microbiol. Res. 16(1), 27. Amaliah, F., Sumiarto, B., Nugroho, W.S., Untari, T., Hartawan, D.H.W. and Agustia, A. 2023. Epidemiology Study of Brucellosis in Polewali Mandar District, Indonesia. J. Biodivers. Biotechnol. 3(1), 27–36. Amjad, M. and Zia, U.U.R. 2023. Poultry as a source and reservoir for Campylobacteriosis. Eur. J. Vet. Med. 3(1), 11–17. Awad, A., Yeh, H.Y., Ramadan, H. and Rothrock, M.J. 2023. Genotypic characterization, antimicrobial susceptibility and virulence determinants of Campylobacter jejuni and Campylobacter coli isolated from pastured poultry farms. Front. Microbiol. 14(1), 1271551. Baali, M., Lounis, M., Al Amir, H.L., Ayachi, A., Hakem, A. and Kassah-Laouar, A. 2020. Prevalence, seasonality, and antimicrobial resistance of thermotolerant Campylobacter isolated from broiler farms and slaughterhouses in East Algeria. Vet. World. 13(6), 1221–1228. Barakat, A.M.A., Abd El-razik, K.A., Elfadaly, H.A., Rabie, N.S., Sadek, S.A.S. and Almuzaini, A.M. 2020. Prevalence, molecular detection, and virulence gene profiles of Campylobacter species in humans and foods of animal origin. Vet. World. 13(7), 1430–1438. Cayrou, C., Barratt, N.A., Ketley, J.M. and Bayliss, C.D. 2021. Phase Variation During Host Colonization and Invasion by Campylobacter jejuni and Other Campylobacter Species. Front. Microbiol. 12(1), 705139. Datta, S., Niwa, H. and Itoh, K. 2003. Prevalence of 11 pathogenic genes of Campylobacter jejuni by PCR in strains isolated from humans, poultry meat and broiler and bovine faeces. J. Med. Microbiol. 52(Pt 4), 345–348. Divya, M., Rajakumar, S. and Ayyasamy, P.M. 2025. In vitro antimicrobial resistant pattern, plasmid profile and the survival of Campylobacter jejuni isolated from poultry sources. Br. Poult. Sci. 1(1), 1–11. Drehkopf, S., Scheibner, F. and Büttner, D. 2023. Functional characterization of virb/vird4 and icm/dot type iv secretion systems from the plant-pathogenic bacterium Xanthomonas euvesicatoria. Front. Cell. Infect. Microbiol. 13(1), 1203159. Elraheam Elsayed, M.S.A. 2019. Virulence repertoire and antimicrobial resistance of Campylobacter jejuni and Campylobacter coli isolated from some poultry farms in Menoufia governorate, Egypt. Pak. Vet. J. 39(2), 2074–7764. Faïs, T., Delmas, J., Serres, A., Bonnet, R. and Dalmasso, G. 2016. Impact of CDT Toxin on Human Diseases. Toxins (Basel). 8(7), 220. Gahamanyi, N., Song, D.G., Yoon, K.Y., Mboera, L.E.G., Matee, M.I., Mutangana, D., Amachawadi, R.G., Komba, E.V.G. and Pan, C.H. 2021. Antimicrobial resistance profiles, virulence genes, and genetic diversity of thermophilic campylobacter species isolated from a layer poultry Farm in Korea. Front. Microbiol. 12(1), 622275. Gharbi, M., Béjaoui, A., Ben Hamda, C., Ghedira, K., Ghram, A. and Maaroufi, A. 2022. Distribution of virulence and antibiotic resistance genes in Campylobacter jejuni and Campylobacter coli isolated from broiler chickens in Tunisia. J. Microbiol. Immunol. Infect. 55(6 Pt 2), 1273–1282. Gharst, G., Oyarzabal, O.A. and Hussain, S.K. 2013. Review of current methodologies to isolate and identify Campylobacter spp. From foods. J. Microbiological Methods 95(1), 84–92. Goualié, B.G., Akpa, E.E., Kakou-N'Gazoa, S.E., Ouattara, H.G., Niamke, S.L. and Dosso, M. 2019. Antimicrobial resistance and virulence associated genes in Campylobacter jejuni isolated from chicken in Côte d'Ivoire. J. Infect. Dev. Ctries. 13(8), 671–677. Hadiyan, M., Momtaz, H. and Shakerian, A. 2022. Prevalence, antimicrobial resistance, virulence gene profile and molecular typing of campylobacter species isolated from poultry meat samples. Vet. Med. Sci. 8(6), 2482–2493. Hickey, T.E., McVeigh, A.L., Scott, D.A., Michielutti, R.E., Bixby, A., Carroll, S.A., Bourgeois, A.L. and Guerry, P. 2000. Campylobacter jejuni cytolethal distending toxin mediates release of interleukin-8 from intestinal epithelial cells. Infect. Immun. 68(12), 6535–6541. Hızlısoy, H., Al, S., Onmaz, N.E., Yıldırım, Y., Gönülalan, Z., Barel, M., Güngör, C., Dışhan, A. and Dişli, H.B. 2020. Farklı Kesimhanelerden İzole Edilen Campylobacter Türlerinin Virülans Genleri, Antibiyotik Duyarlılık Profilleri ve Moleküler Karakterizasyonu Virulence Genes, Antibiotic Susceptibility Profiles and Molecular Characterization of Campylobacter Species Isolated from Different Slaughterhouses. Mikrobiyol. Bul. 54(1), 11–25. Igwaran, A. and Okoh, A. 2020. Campylobacteriosis agents in meat carcasses collected from two district municipalities in the Eastern Cape Province, South Africa. Foods 9(2), 203. Khairullah, A., Yanestria, S., Effendi, M., Moses, I., Kusala, M., Fauzia, K., Ayuti, S., Fauziah, I., Silaen, O., Riwu, K., Aryaloka, S., Dameanti, F., Raissa, R., Hasib, A. and Furqoni, A. 2024. Campylobacteriosis: a rising threat in foodborne illnesses. Open. Vet. J. 14(8), 1733–1750. Kim, H.J., Park, S.E. and Lee, J.H. 2019b. Prevalence and characterization of Campylobacter jejuni isolated from poultry in Korea. J. Microbiol. Biotechnol. 29(5), 771–779. Kim, J., Park, H., Kim, J., Kim, J.H., Jung, J.I., Cho, S., Ryu, S. and Jeon, B. 2019a. Comparative analysis of aerotolerance, antibiotic resistance, and virulence gene prevalence in Campylobacter jejuni isolates from retail raw chicken and duck meat in South Korea. Microorganisms 7(10), 433. Li, Y., Gu, Y., Lv, J., Liang, H., Zhang, J., Zhang, S., He, M., Wang, Y., Ma, H., French, N., Zhang, J. and Zhang, M. 2020b. Laboratory study on the gastroenteritis outbreak caused by a multidrug-resistant Campylobacter coli in China. Foodborne Pathog. Dis. 17(3), 187–193. Li, X., Xu, X., Chen, X., Li, Y., Guo, J., Gao, J., Jiao, X., Tang, Y. and Huang, J. 2023. Prevalence and genetic characterization of Campylobacter from clinical poultry cases in China. Microbiol. Spectr. 11(6), 79723. Li, Y., Zhou, G., Gao, P., Gu, Y., Wang, H., Zhang, S., Zhang, Y., Wang, Y., Jing, H., He, C., Zhen, G., Ma, H., Li, Y., Zhang, J. and Zhang, M. 2020. Gastroenteritis Outbreak Caused by Campylobacter jejuni—Beijing, China, August, 2019. China. CDC. Wkly. 2(23), 422. Lima, L.M., Perdoncini, G., Borges, K.A., Furian, T.Q., Salle, C.T.P., De Souza Moraes, H.L. and Do Nascimento, V.P. 2022. Prevalence and distribution of pathogenic genes in Campylobacter jejuni isolated from poultry and human sources. J. Infect. Dev. Ctries. 16(9), 1466–1472. Liu, Y., Xu, M., Wang, J., Cao, Y., Wang, T., Mu, L. and Niu, C. 2022. Detection of Campylobacter jejuni Based on a Real-Time Fluorescence Loop-Mediated Isothermal Amplification Method. Biomed. Res. Int. 1(1), 3613757. M.S.E., S. 2023. Characterization of Campylobacter jejuni and Campylobacter coli isolates from chicken offal in Metro Manila, Philippines: insights from virulence gene prevalence and multilocus sequence typing analysis. Trop. Biomed. 40(4), 422–429. Matys, J., Turska-Szewczuk, A. and Sroka-Bartnicka, A. 2020. Role of bacterial secretion systems and effector proteins - insight into the pathogenicity mechanisms in Aeromonas. Acta Biochim. Pol. 67(3), 283–293. Melo, R.T., Grazziotin, A.L., Júnior, E.C.V., Prado, R.R., Mendonça, E.P., Monteiro, G.P., Peres, P.A.B.M. and Rossi, D.A. 2019. Evolution of Campylobacter jejuni of poultry origin in Brazil. Food Microbiol. 82(1), 489–496. Montero, L., Medina-Santana, J.L., Ishida, M., Sauders, B., Trueba, G. and Vinueza-Burgos, C. 2024. Transmission of dominant strains of Campylobacter jejuni and Campylobacter coli between farms and retail stores in Ecuador: genetic diversity and antimicrobial resistance. PLos One 19(9), 308030. Nguyen, M.T., Tran, T.H. and Le, H.T. 2023. Detection of virulence genes in Campylobacter jejuni from poultry sources in Vietnam: implications for public health. Int. J. Food. Microbiol. 375(1), 109784. Pons, B., Vignard, J. and Mirey, G. 2019. Cytolethal distending toxin subunit B: a review of structure–function relationship. Toxins 11(10), 595. Prayudi, S.K.A., Effendi, M.H., Lukiswanto, B.S., Az Zahra, R.L., Benjamin, M.I., Kurniawan, S.C., Khairullah, A.R., Silaen, O.S.M., Lisnanti, E.F., Baihaqi, Z.A., Widodo, A. and Riwu, K.H.P. 2023. Detection of Genes on Escherichia coli producing extended spectrum β-lactamase isolated from the small intestine of ducks in traditional markets Surabaya City, Indonesia. J. Adv. Vet. Res. 13(8), 1600–1608. Ramandinianto, S.C., Khairullah, A.R., Effendi, M.H., Tyasningsih, W. and Rahmahani, J. 2020. Detection of Enterotoxin type B gene on Methicillin Resistant Staphylococcus aureus (MRSA) isolated from raw milk in East Java, Indonesia. Sys. Rev. Pharm. 11(7), 290–298. Ramatla, T., Mileng, K., Ndou, R., Tawana, M., Mofokeng, L., Syakalima, M., Lekota, K.E. and Thekisoe, O. 2022. Campylobacter jejuni from slaughter age broiler chickens: genetic characterization, virulence, and antimicrobial resistance genes. Int. J. Microbiol. 1(1), 1713213. Saito, M., Yamazaki, W. and Matsui, H. 2021. Genetic diversity of Campylobacter jejuni strains from poultry in Japan with focus on cytolethal distending toxin genes. Microb. Pathog. 151(1), 104726. Shaltout, S. and Shaltout, F. 2024. Foodborne bacterial disease due to consumption of meat, fish and poultry products. J. Thoracic Dis. Cardiothoracic Surg. 5(2), 2693–2156. Shams, S., Ghorbanalizadgan, M., Haj Mahmmodi, S. and Piccirillo, A. 2017. Evaluation of a Multiplex PCR assay for the identification of Campylobacter jejuni and Campylobacter coli. Infect. Epidemiol. Microbiol. 3(1), 6–8. Shenker, B.J., Korostoff, J., Walker, L.P., Zekavat, A., Dhingra, A., Kim, T.J. and Boesze-Battaglia, K. 2024. Aggregatibacter actinomycetemcomitans cytolethal distending toxin induces cellugyrin-(Synaptogyrin 2) dependent cellular senescence in oral keratinocytes. Pathogens 13(2), 155. Sierra-Arguello, Y.M., Perdoncini, G., Rodrigues, L.B., Ruschel Dos Santos, L., Apellanis Borges, K., Quedi Furian, T., Pippi Salle, C.T., De Souza Moraes, H.L., Pereira Gomes, M.J. and Pinheiro Do Nascimento, V. 2021. Identification of pathogenic genes in Campylobacter jejuni isolated from broiler carcasses and broiler slaughterhouses. Sci. Rep. 11(1), 4588. Subejano, M.S.E.P. and Penuliar, G.M. 2023. Multidrug resistance and high genotypic diversity in Campylobacter upsaliensis from household dogs in Metro Manila, Philippines. New Microbiol. 46(3), 303–307. Syarifah, I.K., Latif, H., Basri, C. and Rahayu, P. 2020. Identification and differentiation of Campylobacter isolated from chicken meat using real-time polymerase chain reaction and high resolution melting analysis of hipO and glyA genes. Vet. World 13(9), 1875–1883. Tang, Y., Jiang, Q., Tang, H., Wang, Z., Yin, Y., Ren, F., Kong, L., Jiao, X. and Huang, J. 2020. Characterization and prevalence of Campylobacter spp. from broiler chicken rearing period to the slaughtering process in Eastern China. Front. Vet. Sci. 7, 227. Tawakol, M.M., Nabil, N.M., Samir, A., M., H.H., Yonis, A.E., Shahein, M.A. and Elsayed, M.M. 2023. The potential role of migratory birds in the transmission of pathogenic Campylobacter species to broiler chickens in broiler poultry farms and live bird markets. BMC. Microbiol. 23(1), 66. Vizzini, P., Vidic, J. and Manzano, M. 2021. Enrichment free qPCR for rapid identification and quantification of Campylobacter jejuni, C. coli, C. lari, and C. upsaliensis in chicken meat samples by a new couple of primers. Foods 10(10), 2341. Wang, G., Clark, C.G., Taylor, T.M., Pucknell, C., Barton, C., Price, L., Woodward, D.L. and Rodgers, F.G. 2002. Colony multiplex PCR assay for identification and differentiation of Campylobacter jejuni, C. coli, C. lari, C. upsaliensis, and C. fetus subsp. fetus. J. Clin. Microbiol. 40(12), 4744–4747. Wanja, D.W., Mbindyo, C.M., Mbuthia, P.G., Bebora, L.C. and Aboge, G.O. 2024. Molecular detection of virulence-associated markers in Campylobacter coli and Campylobacter jejuni isolates from water, cattle, and chicken faecal samples from Kajiado County, Kenya. BioMed. Res. Int. 1(1), 4631351. Wysok, B., Wojtacka, J., Wiszniewska-Łaszczych, A. and Szteyn, J. 2020. Antimicrobial resistance and virulence properties of Campylobacter Spp. originating from domestic geese in Poland. Animals (Basel). 10(4), 742. Yanestria, S., Effendi, M., Tyasningsih, W., Moses, I., Khairullah, A., Kurniawan, S., Dameanti, F., Ikaratri, R., Pratama, J., Sigit, M., Hasib, A. and Silaen, O. 2024. Antimicrobial resistance patterns and genes of Campylobacter jejuni isolated from chickens in Pasuruan, Indonesia. Open Vet. J. 14(3), 759–768. Yanestria, S.M., Effendi, M.H., Tyasningsih, W., Mariyono, M. and Ugbo, E.N. 2023. First report of phenotypic and genotypic (blaOXA-61) beta-lactam resistance in Campylobacter jejuni from broilers in Indonesia. Vet. World. 16(11), 2210. Zhang, Y., Wang, J. and Liu, Z. 2022. Epidemiological surveillance of virulence-associated genes in Campylobacter jejuni from poultry farms in China. Poult. Sci. 101(3), 101741. | ||
| How to Cite this Article |
| Pubmed Style Yanestria SM, Untari T, Artanto S, Effendi MH, Tyasningsih W, Khairullah AR. Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Vet. J.. 2025; 15(12): 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 Web Style Yanestria SM, Untari T, Artanto S, Effendi MH, Tyasningsih W, Khairullah AR. Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. https://www.openveterinaryjournal.com/?mno=279134 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.12 AMA (American Medical Association) Style Yanestria SM, Untari T, Artanto S, Effendi MH, Tyasningsih W, Khairullah AR. Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Vet. J.. 2025; 15(12): 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 Vancouver/ICMJE Style Yanestria SM, Untari T, Artanto S, Effendi MH, Tyasningsih W, Khairullah AR. Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 Harvard Style Yanestria, S. M., Untari, . T., Artanto, . S., Effendi, . M. H., Tyasningsih, . W. & Khairullah, . A. R. (2025) Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Vet. J., 15 (12), 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 Turabian Style Yanestria, Sheila Marty, Tri Untari, Sidna Artanto, Mustofa Helmi Effendi, Wiwiek Tyasningsih, and Aswin Rafif Khairullah. 2025. Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Veterinary Journal, 15 (12), 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 Chicago Style Yanestria, Sheila Marty, Tri Untari, Sidna Artanto, Mustofa Helmi Effendi, Wiwiek Tyasningsih, and Aswin Rafif Khairullah. "Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia." Open Veterinary Journal 15 (2025), 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 MLA (The Modern Language Association) Style Yanestria, Sheila Marty, Tri Untari, Sidna Artanto, Mustofa Helmi Effendi, Wiwiek Tyasningsih, and Aswin Rafif Khairullah. "Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia." Open Veterinary Journal 15.12 (2025), 6287-6297. Print. doi:10.5455/OVJ.2025.v15.i12.12 APA (American Psychological Association) Style Yanestria, S. M., Untari, . T., Artanto, . S., Effendi, . M. H., Tyasningsih, . W. & Khairullah, . A. R. (2025) Virulence gene profile (cdtA, cdtB, cdtC, flaA, and virB) in Campylobacter jejuni isolated from the small intestinal tract of broiler chickens in Indonesia. Open Veterinary Journal, 15 (12), 6287-6297. doi:10.5455/OVJ.2025.v15.i12.12 |