Open Veterinary Journal, (2026), Vol. 16(3): 1793-1807
Research Article
10.5455/OVJ.2026.v16.i3.36
Prevalence and characterization of Pseudomonas fluorescens and Pseudomonas aeruginosa isolates from breast chicken meat
Mat Sa’ad Syamimi-Hanim1, John Yew Huat Tang1,2*, Asmaliza Abd Ghani1 and
Nor Khaizura Mahmud Ab Rashid3,4
1School of Food Industry, Faculty of Bioresources and Food Industry, Universiti Sultan Zainal Abidin (Besut Campus), Besut, Malaysia
2Agriculture and Environment Research Cluster, Deputy Vice Chancellor (Research & Innovation) Office, Universiti Sultan Zainal Abidin (Gong Badak Campus), Kuala Nerus, Malaysia
3Department of Food Science, Faculty of Food Science and Technology, Universiti Putra Malaysia, Serdang, Malaysia
4Laboratory of Food Safety and Food Integrity, Institute of Tropical Agriculture and Food Security, Universiti Putra Malaysia, Selangor, Malaysia
*Corresponding Author: John Yew Huat Tang. School of Food Industry, Faculty of Bioresources and Food Industry, Universiti Sultan Zainal Abidin (Besut Campus), Besut, Malaysia. Email: jyhtang [at] unisza.edu.my
Submitted: 07/08/2025 Revised: 05/02/2026 Accepted: 21/02/2026 Published: 31/03/2026
© 2025 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Background: Although chicken meat is a major source of dietary protein globally, it is highly prone to microbial spoilage, especially under refrigeration, posing economic and public health risks. Psychrotrophic Pseudomonas spp., particularly Pseudomonas fluorescens and Pseudomonas aeruginosa, are key spoilage organisms due to their enzyme production, biofilm formation, and cold tolerance.
Aim: This study determined the prevalence, virulence gene profiles, enzymatic activity, and antibiotic resistance profiles of P. fluorescens and P. aeruginosa isolated from raw chicken meat sold in supermarkets.
Methods: Detection and isolation of Pseudomonas spp. in raw chicken samples were carried out using selective Pseudomonas agar. PCR assay used species-specific primers targeting adnA and toxA genes to confirm the presence of P. fluorescens and P. aeruginosa, respectively. The isolated P. fluorescens and P. aeruginosa were molecularly characterized using a polymerase chain reaction (PCR) assay for the presence of genes responsible for biofilm formation, proteolytic enzymes, and virulence. The proteolytic and lipolytic activities of the isolates at 30 °C and 4°C using skim milk agar and tributyrin agar. Antibiotic susceptibility was determined using the Kirby–Bauer disk diffusion method.
Results: Of 180 samples, 61 (33.9%) were positive for P. fluorescens and 34 (18.9%) were contaminated with P. aeruginosa, as confirmed by PCR assay. Molecular characterization of the P. fluorescens isolates revealed that 41.0% were positive for aprX and 8.2% had fliC, whereas the P. aeruginosa isolates showed the presence of oprL (58.8%), lasB (11.4%), and exoS (20.6%). Gene distribution varied across retail outlets, possibly reflecting differences in hygiene and storage conditions. At 4 °C, most P. fluorescens isolates (82%) displayed proteolytic activity, and all isolates showed lipolytic activity. P. aeruginosa showed low proteolytic (29.4%) but high lipolytic (88.2%) activity. These findings show their ability to degrade meat even under refrigeration conditions. Antibiotic susceptibility testing revealed high resistance to penicillin G, ampicillin, and piperacillin among all isolates. P. fluorescens showed notable resistance to ceftazidime (100%) but remained largely susceptible to gentamicin (80.3%) and levofloxacin (82.0%). P. aeruginosa exhibited similar trends, with high susceptibility to gentamicin (85.3%) and levofloxacin (91.2%). Moderate susceptibility to amikacin was observed in both of the aforementioned species. All isolates had MAR index scores above 0.2, and 19 P. fluorescens and 5 P. aeruginosa isolates were identified as extensively drug-resistant (XDR), resistant to at least five antibiotic classes.
Conclusion: These findings highlight the contamination of retail chicken meat by virulent, spoilage-causing, and antibiotic-resistant Pseudomonas spp., which requires improved control measures to minimize spoilage and safeguard public health.
Keywords: Pseudomonas fluorescens, Pseudomonas aeruginosa, Antibiotic resistance, Protease, Lipase.
Introduction
Chicken meat is widely consumed because of its high nutritional value, affordability, low fat content, and favorable taste. Chicken has become the primary source of dietary protein in Malaysia, with an average per capita consumption of approximately 52.0 kg (Syafuddin Tan et al., 2023). However, similar to other animal-derived foods, chicken meat is highly perishable and susceptible to microbial contamination, which can lead to spoilage and foodborne illnesses (Heredia and García, 2018).
Among the various microorganisms associated with meat spoilage, Pseudomonas spp. are considered predominant psychrotrophic bacteria that can adapt well to refrigeration conditions. Pseudomonas fluorescens and Pseudomonas aeruginosa are frequently isolated from chilled chicken meat and have been identified as key spoilage bacteria due to their ability to produce extracellular enzymes such as proteases and lipases (Rawat and S, 2018; Heir et al., 2021; Elbehiry et al., 2022). These enzymes degrade proteins and lipids in meat, resulting in off-odors, off-flavors, rancidity, and decreased shelf life (Raposo et al., 2017).
Contamination with Pseudomonas spp. can occur at any stage in chicken production, such as slaughtering, processing, transportation, and retail distribution. Other contributing factors include poor sanitation, improper handling, temperature abuse, and inadequate hygiene among food handlers (Karanth et al., 2023; Lianou et al., 2023). Moreover, the lack of awareness and implementation of hygiene practices at retail outlets increases the risk of contamination (Bantawa et al., 2018). These bacteria can multiply and cause spoilage, discoloration, and sensory degradation of meat products even under refrigeration (Carrizosa et al., 2017; Wickramasinghe et al., 2019). Contaminated meat is an acute source of gastrointestinal infections, and the microbial count can be used as an indicator of the hygienic condition and quality of meat (Fakhkhari et al., 2020).
In meat processing facilities, Pseudomonas spp. have been reported to form biofilms on stainless steel or plastic surfaces (Calhoun et al., 2025). When Pseudomonas spp. enter into a biofilm state, a higher antibiotic concentration of up to 40 times was needed to remove the biofilm compared with planktonic cells (Molina et al., 2014). Currently, antibiotic resistance is one of the most critical challenges to global health and food safety. Emerging bacterial resistance to several antibiotics poses a severe threat to public health worldwide. These resistant bacteria frequently prolong the treatment procedures and transfer genetic material to other bacteria, transforming them into drug-resistant strains (WHO and Organization, 2017). Pseudomonas spp. is one of the most critical Gram-negative bacteria that cause diseases and become resistant to multiple antibiotics. The predominance and persistence of Pseudomonas spp. in foods and on food equipment surfaces have also been related to the capability of these microorganisms to form biofilms, which enhances their tolerance to adverse conditions, including antimicrobial treatments (Moretro and Langsrud, 2017; Quintieri et al., 2019).
The European Committee on Antimicrobial Susceptibility Testing EUCAST (2022) has recently highlighted that non-fermenting Gram-negative Pseudomonas spp. are naturally resistant to certain antibiotic classes, including beta-lactams, macrolides, and glycopeptides, which cover antibiotics differentiated by the chemical structure. Heir et al. (2021) reported that P. fluorescens isolates from chicken meat exhibited significant resistance to aztreonam (72.6%), colistin (30.2%), imipenem (25.6%), and meropenem (12.6%). Meanwhile, P. aeruginosa strains isolated from chicken meat samples also showed higher levels of resistance to aztreonam and amoxicillin Jawher and Hassan (2023).
Previous studies have shown that the presence of P. fluorescens and P. aeruginosa in the poultry processing environment can significantly impact the quality, safety, and shelf life of the product. Therefore, the objectives of this study are (i) to detect P. fluorescens and P. aeruginosa contamination in raw chicken meat; (ii) to molecularly characterize P. fluorescens and P. aeruginosa isolates; (iii) to determine protease and lipase activities of P. fluorescens and P. aeruginosa isolates; and (iv) to evaluate antimicrobial resistance profiles of P. fluorescens and P. aeruginosa isolates. This study provides important insights into the spoilage potential and antibiotic resistance profiles of two important Pseudomonas spp. (P. fluorescens and P. aeruginosa) contamination in raw chicken meat.
Materials and methods
Samples collection
A total of 180 raw breast chicken meat samples displayed on ice flakes were purchased from five supermarkets in Kuala Terengganu, Terengganu, Malaysia, labeled 1SP, 2SP, 3SP, 4SP, and 5SP, with each supermarket comprising 36 samples. Samples were purchased and individually stored in a stomacher bag and transported immediately to the laboratory in an ice box.
Detection and isolation of P. fluorescens and P. aeruginosa
Approximately 10 g of the sample was added to 90 ml of 0.1% peptone water (Oxoid, United Kingdom) and homogenized using a stomacher (Interscience, France) (Bulut, 2014; Oh et al., 2014; Cetin et al., 2019). Aliquots of 0.1 ml from selected dilutions were spread plated on Pseudomonas agar base (Oxoid, United Kingdom) containing Pseudomonas CFC supplement (Oxoid, United Kingdom) that suppresses the growth of background microorganisms to facilitate the growth of Pseudomonas. Plates were incubated aerobically at 30°C for 3 days. Presumptive colonies were purified on tryptic soy agar (Merck, Germany). Gram staining and biochemical tests (oxidase, catalase, indole, nitrate, and gelatin) were performed. Glycerol stock cultures were prepared and kept at −20°C for further investigation.
Polymerase chain reaction identification of P. fluorescens and P. aeruginosa
DNA from the isolates was extracted using the conventional boiling method as described by Hsern Malcolm et al., (2015) with some modifications. PCR assay was performed to confirm the presence of P. fluorescens and P. aeruginosa DNA using adnA primers F:5′- ATGTGGCGTGAAACCAAAAT-3′ and R:5′-TCAATCATCCGCCTGTTCA-3′ (Xu
et al., 2017) and toxA primers F:5′- GACAACGCCCTCAGCATCACCAGC-3′ and R:5′- CGCTGGCCCATTCGCTCCAGCGCT-3′ (Algammal et al., 2020), respectively. The PCR amplification mixture consisted of 1x reaction buffer, 2 mM MgCl2, 0.2 mM dNTPs, 0.8 uM forward and reverse primers, 2 U Taq DNA polymerase, 1 µl DNA template and added up until 25 µl with sterile distilled water. The PCR assay was performed using a thermocycler (Applied Biosystems, USA). PCR cycling condition included the initial denaturation stage with temperature 95°C for 4 minutes (1 cycle), denaturation stage with temperature 95°C and 30 seconds, the annealing stage temperature 56°C and 30 seconds, the extension stage temperature 72°C and 1 minutes for total of 30 cycles, and final extension temperature 72°C for 5 minutes (1 cycle). PCR products were run through gel electrophoresis with 1% FloroSafe DNA Stain (1st BASE, Singapore), visualized, and recorded with a gel imaging system (Fujifilm, Japan).
Molecular characterization of P. fluorescens and P. aeruginosa
The PCR assay was performed to detect protease, lipase, and biofilm formation genes of P. fluorescens and P. aeruginosa. The primers used for P. fluorescens and P. aeruginosa are summarized in Table 1. The PCR amplification mixture consisted of 1× reaction buffer, 2 mM MgCl2, 0.2 mM dNTPs, forward and reverse primers (0.5 uM for aprX, fliC, lasB, and exoS, 0.1 uM for oprL), 2 U Taq DNA polymerase, 1 µl DNA template, and added up to 25 µl with sterile distilled water. The PCR assay was performed using a thermocycler (Applied Biosystems, USA). The PCR products were run through gel electrophoresis with 1% FloroSafe DNA Stain (1st BASE, Singapore), visualized, and recorded using a gel imaging system (Fujifilm, Japan).
Table 1. Description of primers for the characterization of P. fluorescens and P. aeruginosa.
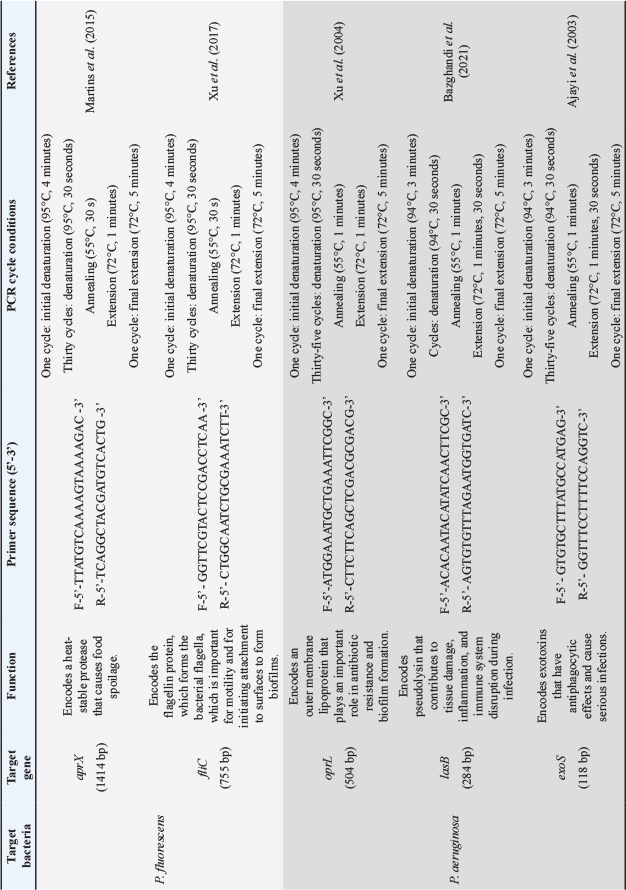
Proteolytic and lipolytic activities
The proteolytic and lipolytic activities of each P. fluorescens and P. aeruginosa isolate were determined using skim milk agar (Himedia, India) and agar base (Himedia, India) with a tibutyrin supplement (Himedia, India), respectively. Each isolate culture from tryptic soy broth (Merck, Germany) was aseptically stabbed onto agar plates using a needle loop. The stabbed plates were then incubated at 30°C for 3 days and 4°C for 10 days. The formation of a clear zone around each colony was measured using a ruler and classified as positive for proteolytic and lipolytic activities. The isolates were classified as low potential (proteolytic: halos ≤ 2 cm and lipolytic: halos < 1.5 cm) or high potential (proteolytic: halos >2 cm and lipolytic ≥1.5 cm), as described by Narvhus et al. (2021).
Antimicrobial susceptibility testing
All P. fluorescens and P. aeruginosa isolates were evaluated for their susceptibility to 16 antimicrobial agents (Oxoid, United Kingdom), including penicillin (P, 5 IU), ampicillin (AMP, 10 μg), piperacillin (PRL, 30 μg), amikacin (AK, 30 μg), gentamicin (CN, 10 μg), ciprofloxacin (CIP, 5 μg), ceftazidime (CAZ, 30 μg), cefepime (FEP, 30 μg), levofloxacin (LEV, 5 μg), norfloxacin (NOR, 5 μg), imipenem (IPM, 10 μg), meropenem (MEM, 10 μg), erythromycin (E, 15 μg), clindamycin (DA, 2 μg), tetracycline (TE, 30 μg), and rifampicin (RD, 5 μg). The standard Kirby–Bauer disc diffusion method (Hudzicki and J, 2009) was used. Bacterial isolate cultures equivalent to 0.5 McFarland standards were aseptically swabbed onto Mueller–Hinton agar plates (Merck, Germany) and allowed to dry at room temperature. The antibiotic discs were then placed on the surface of the agar plate using an antibiotic disc dispenser (Oxoid, Finland). Standards and interpretive criteria illustrated by the Clinical and Laboratory Standards Institute (CLSI, 2020) guidelines were used to identify whether the isolates were susceptible, intermediate, or resistant to the antibiotics tested. Each resistant isolate was calculated for the MAR index using the formula (Meng et al., 2020):
MAR=a/b
where
a: Number of antibiotics to which the isolate was resistant
b: Number of antibiotics tested
Extensive drug resistance (XDR) was determined based on antimicrobial resistance profiles. The isolates were classified as XDR when they exhibited resistance to all antimicrobial classes tested (n=7) or when they demonstrated resistance to nearly all antimicrobial classes, with remaining susceptible to two or fewer antimicrobial classes (Briega et al., 2025).
Ethical approval
Not needed for this study.
Results
Presumptive Pseudomonas spp. colonies measuring approximately 1–3 mm in size grew on selective Pseudomonas agar and were selected for further tests. Pseudomonas fluorescens formed a flat and greenish-yellow colony, whereas P. aeruginosa formed a straw-colored and blue-greenish colony. The isolated presumptive colonies were Gram-stained and subjected to biochemical tests before species identification with PCR using species-specific primers for the presence of adnA and toxA for P. fluorescens and P. aeruginosa, (Fig. 1) respectively. Pseudomonas fluorescens was detected in chicken meat samples obtained from all evaluated supermarkets (Table 2). Chicken samples from the 3SP showed the highest percentage at 50.0%, whereas chicken meat samples from the 1SP and 4SP had the lowest prevalence at 22.2%.
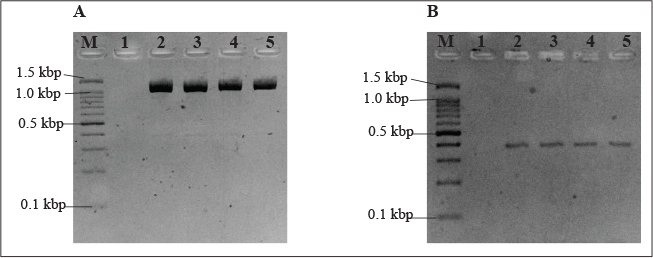
Fig. 1. PCR assay of P. fluorescens and P. aeruginosa detection in chicken meat samples. A. PCR using P. fluorescens species-specific primers adnA-F and adnA-R giving PCR product of 1.48kbp. (B) PCR using P. aeruginosa species-specific primers toxA-F and toxA-R giving PCR product of 396 bp. Lanes M, 100 bp markers; lane 1, negative control; lane 2, positive control; lanes 3-5, DNA extract from chicken meat samples.
Pseudomonas aeruginosa was also found in chicken samples from all five supermarkets (Table 2). The highest prevalence was found in 5SP, where 30.6% of the samples tested positive for P. aeruginosa. This was followed by 2SP, 3SP, and 4SP, which recorded contamination levels of 25.0%, 16.7%, and 13.9%, respectively. Supermarket 1SP had the lowest percentage of 8.33%.
Table 2. Prevalence of P. fluorescens and P. aeruginosa in chicken meat samples.
N=total isolates;
n=positive isolates; SP=Supermarket.
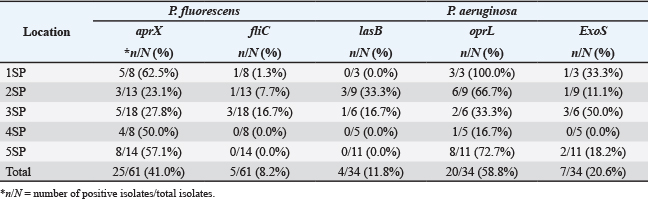
Table 3 shows distinct patterns in the presence of P. fluorescens and related genes. At 1SP, 5 isolates (62.5%) were positive for the aprX gene, (Fig. 2) the highest among all sampling locations, while only 1 isolate (1.3%) was positive for fliC (Fig. 2). At 2SP, 3 isolates (23.1%) carried aprX, and 1 isolate (7.7%) carried the fliC gene. At 3SP, 5 isolates (27.8%) were positive for aprX, while 3 isolates (16.7%) were positive for fliC. However, in both 4SP and 5SP no presence of fliC gene was detected in P. fluorescens isolates at 4SP and 5SP.
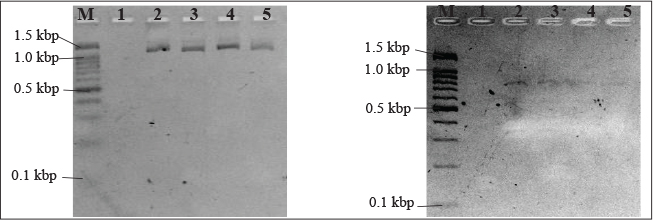

Fig. 2. PCR assay detection of P. fluorescens protease (aprX) and biofilm related (fliC) genes. A. PCR using P. fluorescens aprX-F and aprX-R primers giving PCR product of 1.43kbp. B. PCR using P. fluorescens fliC-F and fliC-R primers giving PCR product of 755 bp. Lanes M, 100 bp markers; lane 1, negative control; lane 2, positive control; lanes 3-5, DNA extract from chicken meat samples.
PCR assay was used to detect the presence of oprL, lasB and exoS genes in P. aeruginosa (Fig. 3). This study revealed variation in the distribution of virulence genes in P. aeruginosa isolates (Table 3).. The oprL gene was highest in isolates from 1SP (100%), followed by those from 5SP (72.7%) and 2SP (66.7%). In contrast, the lasB gene was found in only 33.3% of isolates from 2SP and 16.7% of isolates from 3SP, whereas it was completely absent in 1SP, 4SP, and 5SP. The exoS gene was detected in isolates from 3SP (50.0%), 1SP (33.3%), 2SP (11.1%), and 5SP (18.2%) but was absent in 4SP.
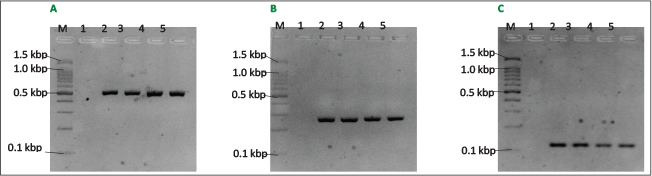
Fig. 3. PCR assay of P. aeruginosa virulence (oprL, lasB and exoS) genes. A. PCR using P. aeruginosa oprL-F and oprL-R primers giving PCR product of 504 bp. B. PCR using P. aeruginosa lasB-F and lasB-R primers giving PCR product of 284 bp. C. PCR using P. aeruginosa exoS-F and exoS-R primers giving PCR product of 118 bp. Lanes M, 100 bp markers; lane 1, negative control; lane 2, positive control; lanes 3-5, DNA extract from chicken meat samples.
Table 3. Characterization of P. fluorescens and P. aeruginosa isolates from chicken meat samples.

*
n/
N=number of positive isolates/total isolates.
The data in Table 4 illustrate that P. fluorescens isolates possessed significant proteolytic and lipolytic activity over a temperature range, indicating their potential role in refrigerated food deterioration. Higher proteolytic activity was found at 4°C (82%) than temperature of 30°C (63.9%). This enzymatic activity may contribute to the progressive degradation of protein-rich food matrices during refrigeration. At 4°C, 96.7% of the isolates exhibited lipolytic capability, and 85.2% at 30°C. Additionally, all isolates showed lipolytic activity, indicating a consistent ability to produce lipid hydrolysis.
Table 4. Proteolytic and lipolytic activities of P. fluorescens (N=61) and P. aeruginosa (N=34) isolates at 30 °C (day 3) and 4 °C (day 7).

N=total isolates;
n=positive isolates
Temperature affects the extracellular enzymatic activity of Pseudomonas spp. isolates. As shown in Table 4, the proportion of isolates exhibiting lipolytic activity was higher at the low-temperature condition, whereas most isolates maintained lipolytic activity at 30°C. Approximately 47.0% of P. aeruginosa isolates showed proteolytic activity at 30°C, with 17.6% showing high levels. At 4°C, only 29.5% of isolates showed protease activity, whereas the percentage of highly active protease isolates decreased to 3.0%. Lipolytic activity was observed in 88.2% of isolates at both 30°C and 4°C. Approximately 32.3% of P. aeruginosa isolates showed high lipolytic activity, while only one isolate (3.0%) showed lipase activity at 4°C. Most isolates (85.2%) exhibited low lipolytic activity at lower temperatures.
As shown in Table 5, all P. fluorescens isolates were highly resistant to the penicillin class, with penicillin G (100%), ampicillin (100%), and piperacillin (100%). Similarly, macrolides (erythromycin and clindamycin), as well as ceftazidime and rifampicin, also showed complete resistance (100%). In contrast, P. fluorescens isolates were highly susceptible to fluoroquinolones, with 90.2%, 82.0%, and 98.4% susceptibility to ciprofloxacin, levofloxacin, and norfloxacin, respectively. Aminoglycosides were also extremely effective, with amikacin and gentamicin showing bacterial susceptibility at 91.8% and 80.3%, respectively. Cefepime also showed high susceptibility at 89.0%.
Table 5. Antimicrobial profiles of P. fluorescens (N=61) and P. aeruginosa (N=34).
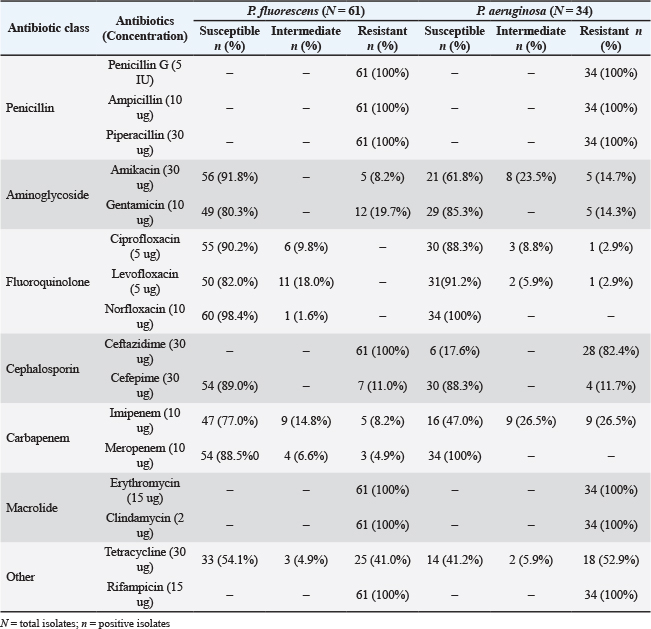
N=total isolates;
n=positive isolates
All P. aeruginosa isolates were highly resistant (100%) to penicillin G, ampicillin, and piperacillin. Norfloxacin and meropenem were found to be extremely effective against all isolates, with 100% susceptibility. Similarly, gentamicin exhibited a strong effect with 85.3% susceptibility, while cefepime, ciprofloxacin, and levofloxacin showed high susceptibility of 88.3%, 88.3%, and 91.2%, respectively. Amikacin, another aminoglycoside, also showed efficacy at 61.8%. However, P. aeruginosa isolates demonstrated high resistance to erythromycin (100%), clindamycin (100%), and rifampicin (100%), making them ineffective for treating P. aeruginosa infections. Compared with cefepime, ceftazidime revealed 82.4% resistance within the cephalosporin class, making it a poor choice. Meanwhile, carbapenem imipenem showed a mixed profile with 26.5% intermediate and 26.5% resistant isolates. Tetracycline showed about 52.9% resistance and 41.2% susceptibility in isolates. These findings emphasize the vital importance of susceptibility testing for ensuring optimum antibiotic selection and addressing the emerging issue of AMR in P. aeruginosa.
Table 6 shows that P. fluorescens isolates from chicken meat have a high level of MDR, with the majority of the isolates showing MAR indices of 0.5 or 0.6. The most common resistance profile, which included resistance to penicillin G, ampicillin, piperacillin, ceftazidime, tetracycline, clindamycin, rifampicin, and erythromycin, was observed in 14 isolates, giving the MAR score of 0.5. Another major group of 25 P. fluorescens isolates shared a similar profile but had no tetracycline resistance, resulting in a MAR value of 0.4. The prevalence of MAR indices at or above 0.4 across nearly all 61 isolates (35 isolates with a 0.5 or 0.6 MAR index) indicated widespread multidrug resistance in P. fluorescens isolates.
Table 6. Antibiotic resistance profiles and MAR index of P. fluorescens (N=61) and P. aeruginosa (N=34) isolates.
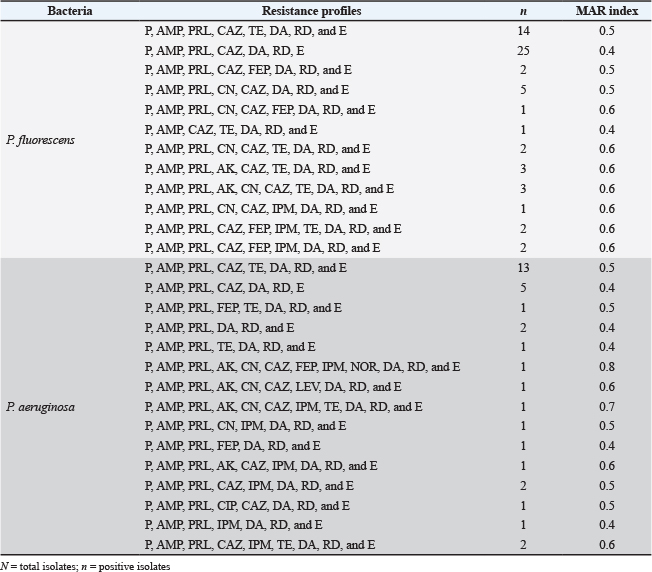
N=total isolates;
n=positive isolates
A total of 15 different resistance profiles among P. aeruginosa isolates were obtained (Table 6). Thirteen isolates (38.2%) showed resistance to penicillin G, ampicillin, piperacillin, ceftazidime, tetracycline, clindamycin, rifampicin, and erythromycin, resulting in a MAR score of 0.5. Crucially, every isolate exhibited a MAR index ranging from 0.4 to a hazardous 0.8.
Next, Table 7 shows the XDR profiles of P. fluorescens isolates, demonstrating a concern level of antibiotic resistance in this bacterial species. The majority of the isolates listed, including 3SP1801, 1SP2901, and 1SP2201, show resistance to five different antibiotic classes. The P. fluorescens isolate 2SP2201 has many broader resistances, extending to six antibiotic classes, including carbapenems. Resistance to imipenem from the carbapenem class has raised concerns because it is often considered the final option for treating Pseudomonas spp. infection (Wangchinda et al., 2025).
All five P. aeruginosa isolates, namely, 4SP1902, 2SP201, 3SP3502, 1SP102, and 2SP302, showed the same resistance pattern to penicillin G, ampicillin, piperacillin, ceftazidime, clindamycin, rifampicin, and erythromycin, indicating that these commonly used antibiotics are likely ineffective against these strains (Table 7). Among the isolates, 4SP1902 has the broadest resistance, being resistant to seven distinct antibiotic classes, including amikacin, gentamicin, cefepime, imipenem, and norfloxacin. The remaining four isolates (2SP201, 5SP3502, 1SP102, and 2SP302) are resistant to six antibiotic classes.
Table 7. XDR of P. fluorescens and P. aeruginosa isolates.
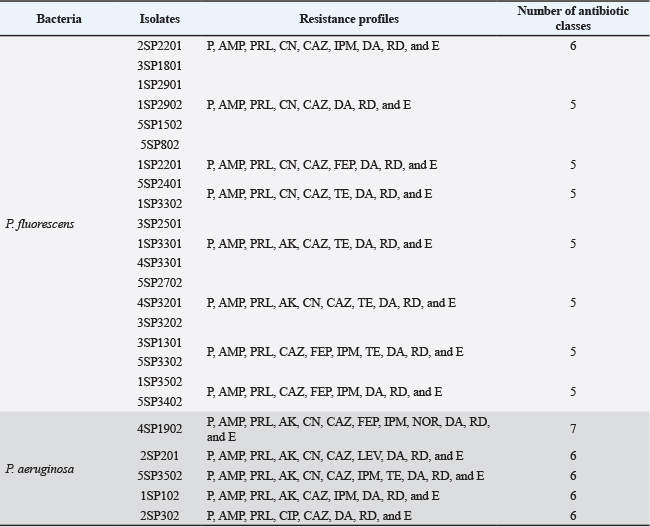
Discussion
The high prevalence of P. fluorescens corresponds with previous studies demonstrating its dominance among psychrotrophic bacteria in refrigerated chicken meat. Can (2022) found that approximately 78.7% of Pseudomonas isolates from refrigerated chicken drumsticks were P. fluorescens. Similarly, Hassan et al. (2020) found that P. fluorescens was the most common species in various chicken meat-derived products, including nuggets and burgers. Because of its psychrotrophic nature, the dominance of P. fluorescens in chicken meat allows it to proliferate during cold storage, contributing to spoilage and food safety concerns. Its prevalence across various chicken products underscores its adaptability and persistence during cold storage.
The variability in contamination rates among supermarkets likely reflects the differences in operational practices, including storage conditions, hygiene practices, and handling procedures (Meng et al., 2017; Ben Mhenni et al., 2023; Ahmed and Hassan, 2024). Therefore, the higher contamination rate of P. fluorescens at the 3SP may suggest less severe control procedures that lead to more favorable conditions for bacterial growth or a higher-risk environment for P. fluorescens cross-contamination.
The presence of P. aeruginosa in retail chicken meat is particularly concerning due to its pathogenicity and spoilage potential. A study conducted in Ethiopia discovered a prevalence rate of 27.6% in poultry carcasses, whereas another study in Egypt revealed a prevalence rate of 22.9% in healthy chickens (Tigabie et al., 2025). In contrast, research in Kenya found only approximately 0.3% P. aeruginosa in retail chicken meat samples (Muinde et al., 2023). Although its prevalence varies across regions, its ability to survive under diverse environmental conditions, including refrigeration (Elbehiry et al., 2022), makes it persistent in poultry products. Improper storage temperatures and poor hygiene during processing can promote the growth of P. aeruginosa, which contributes to higher contamination rates. The findings of this study highlight the importance of good hygiene practices to reduce the risk of P. aeruginosa contamination in chicken meat. Moreover, regular monitoring and compliance with food safety guidelines are crucial for ensuring the microbiological safety of poultry products and maintaining public health.
The presence of the aprX gene in P. fluorescens isolates is responsible for the production of thermostable alkaline metalloprotease, which is also known as the primary enzyme that causes spoilage in chilled food products (Glantz et al., 2020; Du et al., 2023). This enzyme is highly resistant to heat treatments such as high-temperature processing and will continue to break down protein during storage, resulting in undesirable changes such as bitterness, off-flavors, and gelation production (Andreani, 2016; Glantz et al., 2020).
The fliC gene in P. fluorescens encodes flagellin, which is responsible for bacterial motility in liquid environments and on food contact surfaces (Martínez-Granero et al., 2012; Redondo-Nieto et al., 2013). Motility is an important feature in bacterial colonization and adherence to surfaces, which is the first stage in biofilm formation (Chen et al., 2023). Biofilms can protect bacteria from adverse environmental conditions and antimicrobial treatments, resulting in persistent contamination and increased deterioration in food processing environments (Martínez-Granero et al., 2012). The motile capability of P. fluorescens plays a significant role in spoilage by facilitating bacterial transmission, colonization, and development in contaminated food products and processing equipment (Martínez-Granero et al., 2012; Redondo-Nieto et al., 2013).
The oprL gene of P. aeruginosa encodes the production of an outer membrane lipoprotein, which helps to maintain the strength of the bacterial outer membrane, and contributes to the inherent resistance of P. aeruginosa to antibiotics by affecting membrane permeability and efflux mechanisms (Shahat et al., 2019; Algammal et al., 2023). The lasB gene of P. aeruginosa plays a role in biofilm formation, tissue damage, and immune system stimulation. The pathogenesis of P. aeruginosa infections is associated with the degradation of host proteins, such as elastin, immunoglobulins, and complement factors (Galdino et al., 2017). The presence of lasB in a significant number of 2SP and 3SP isolates showed that P. aeruginosa strains from these supermarkets pose significant public health risks.
The P. aeruginosa exoS gene encodes an ADP-ribosyl transferase effector protein, which is another important virulence factor that destroys tissue, alters cellular structure, and increases bacterial transmission in host cells (Sato and Frank, 2004). The presence of these virulence genes in P. aeruginosa contaminating chicken meat shows that it can cause serious foodborne infection.
This study pattern of enzymatic reaction is consistent with previous studies in psychrotrophic Pseudomonas spp. cultures, which produce lipases optimally at low temperature (2–15 °C). However, the presence of enzymes is basically sustained at cold temperatures, and enzymatic effectiveness decreases when compared to warmer environments (Meng et al., 2017; Salwoom et al., 2019; Abdella et al., 2023). This finding is also complemented by a study conducted by Zhang and Lv (2014) in which proteases from Pseudomonas spp. were shown to function intensely at low temperatures. Hence, this study found that P. fluorescens remains capable of producing spoilage-associated protease and lipase enzymes even when stored under refrigeration, indicating that cold storage does not completely prevent spoiling potential and may contribute to meat degradation in the presence of rancid and off-flavor.
Our findings align with those of Elbehiry et al. (2022) who identified P. fluorescens isolates from chicken meat and showed high resistance to penicillin, especially ampicillin (71%) and moderate resistance to ciprofloxacin (49%). Similarly, Mohamed et al. (2022) discovered that P. fluorescens isolates from chicken meat were consistently resistant to piperacillin (63.6%) and moderately resistant to aminoglycosides, particularly gentamicin (27.2%), although aminoglycoside resistance in Pseudomonas spp. remained rare. In addition, a large-scale market survey of antibiotic resistance in Chiang Mai, Thailand, reported a P. fluorescens percentage of approximately 18% in fresh chicken samples (Dokuta et al., 2025). Resistance trends were comparable to those observed in the present study, especially for penicillin and older generation β-lactams. This study correlates with the recent findings of Abdullah et al. (2024) who found that P. aeruginosa isolates from poultry meat were highly susceptible to meropenem (84%) and amikacin (75%). Antibiotic susceptibility of Pseudomonas is important to ensure successful treatment if infection occurs from contaminated food consumption or handling of farm animals.
A MAR index greater than 0.2 implies exposure to substantial antibiotic selective pressure. The data in this study clearly suggest that the Pseudomonas isolates originated from a high-risk environment that commonly occurs in commercial poultry farms and utilizes antibiotics extensively (Mapipa et al., 2021). However, the findings of this study were not consistent with those of Guibert et al. (2024) who identified seven P. aeruginosa isolates from retail chicken meat as MAR, with all showing resistance to fluoroquinolones (including ciprofloxacin), highlighting the potential for high-level fluoroquinolone resistance in meat-borne isolates. The presence of carbapenem resistance in P. aeruginosa isolates highlights a potential risk of difficult-to-treat infections, particularly in immunocompromised individuals. The World Health Organization has classified carbapenem-resistant P. aeruginosa as one of three bacterial species that require new antibiotics to treat infections (Tacconelli et al., 2017). Hence, Pang et al. (2019) stated that new antibiotics and alternative therapy techniques are urgently needed to treat P. aeruginosa infections that are resistant to conventional antibiotics.
The discovery of P. aeruginosa XDR isolates from chicken meat highlights the risk of poultry emerging as a reservoir for high-level resistance. This study discovered five P. aeruginosa isolates identified as XDR strains, corroborating with an earlier report by Dong et al. (2022) who revealed an isolate (PA1662) from chicken in one of the poultry farms classified as XDR as its only susceptible to aztreonam and highly resistant to other tested antimicrobial classes including carbapenems (meropenem and imipenem), cephalosporin (ceftazidime), fluoroquinolone (ciprofloxacin), and amikacin. The carbapenem-resistant isolates from poultry isolates reflect selective pressure from the use of β-lactam or carbapenem analogs in veterinary practices, either for therapeutic or prophylactic purposes. In addition, Pseudomonas spp. can modify membrane structures, alter metabolic pathways, and activate stress responses in response to temperature changes, nutrient deprivation, and disinfectant exposure.
The discovery of these XDR isolates raises serious public health concerns that pose great challenges in clinical management, potentially leading to increased morbidity and mortality, especially when transmitted through contaminated food consumption. Proper handling of raw materials, preparation and processing of meat, and proper waste disposal may reduce the antibiotic resistance of Pseudomonas spp. in chicken and other poultry products (Jawher and Hassan, 2023). This emphasizes the critical need for continuous, comprehensive surveillance of AMR in both clinical and food settings, highlighting a crucial strategy for combating the worldwide challenge of drug-resistant bacteria.
Conclusion
The findings of this study highlight the high occurrence of P. fluorescens and P. aeruginosa in raw chicken meat, raising concerns over food quality degradation and public health risks. The presence of important virulence genes of P. fluorescens (aprX and fliC genes) and P. aeruginosa (oprL, lasB, and exoS) suggests that these bacteria cause substantial poultry spoilage and cross-contamination in the food processing environment. Importantly, this study revealed that both P. fluorescens and P. aeruginosa exhibited proteolytic and lipolytic activity at 4°C, demonstrating their ability to degrade chicken meat products during cold storage to extend shelf life. Nevertheless, the discovery of multidrug resistance among isolates emphasizes the importance of better surveillance, hygiene practices, and antimicrobial stewardship in poultry production and supply chain to reduce microbiological hazards.
Acknowledgment
The authors would like to thank the fund provider Ministry of Higher Education (MOHE) under Fundamental Research Grant Scheme (FRGS/1/2023/WAB04/UNISZA/02/5) and Universiti Sultan Zainal Abidin (UniSZA), Besut Campus, Terengganu, Malaysia for providing the laboratory facilities.
Funding
This study was supported by the Ministry of Higher Education (MOHE) under the Fundamental Research Grant Scheme (FRGS/1/2023/WAB04/UNISZA/02/5).
Authors' contributions
Mat Sa'ad Syamimi Hanim performed the lab work, analysis and draft the manuscript. John Yew Huat Tang designed the experiment and improved the manuscript. Asmaliza Abd Ghani help in the data analysis. Nor Khaizura Mahmud Ab Rashid help in designing the experiment.
Conflict of interest
The authors have no conflicts of interest to declare.
Data availability
All data were provided in the manuscript.
References
Abdella, B., Youssif, A.M., Sabry, S.A. and Ghozlan, H.A. 2023. Production, purification, and characterization of cold-active lipase from the psychrotroph Pseudomonas sp. A6. Braz. J. Microbiol. 54, 1623–1633.
Abdullah, R.M., Ali, S., Bilal, A. and Arshad, M.I. 2024. Molecular characterization and drug resistance pattern of Pseudomonas aeruginosa isolated from poultry meat and meat products. Pak. Vet. J. 44(3), 812–818.
Ahmed, A.H. and Hassan, M.G. 2024. Monitoring the prevalence of Pseudomonas fluorescens as a spoilage indicator in cow raw milk, teat surfaces, and milk tanks. Open. Vet. 14(8), 1983–1989.
Ajayi, T., Allmond, L.R., Sawa, T. and Wiener-Kronish, J.P. 2003. Single-Nucleotide-Polymorphism mapping of the Pseudomonas aeruginosa type III secretion toxins for development of a diagnostic multiplex PCR system. J. Clin. Microbiol. 41(8), 3526–3531.
Algammal, A.M., Eidaroos, N.H., Alfifi, K.J., Alatawy, M., Al-Harbi, A.I., Alanazi, Y.F., Ghobashy, M.O., Khafagy, A.R., Esawy, A.M., El-Sadda, S.S., Hetta, H.F. and El-Tarabili, R.M. 2023. OprL gene sequencing, resistance patterns, virulence genes, quorum sensing and antibiotic resistance genes of XDR Pseudomonas aeruginosa isolated from broiler chickens. Infect. Drug. Resist. 16, 853–867.
Algammal, A.M., Mabrok, M., Sivaramasamy, E., Youssef, F.M., Atwa, M.H., El-Kholy, A.W., Hetta, H.F. and Hozzein, W.N. 2020. Emerging MDR-Pseudomonas aeruginosa in fish commonly harbor oprL and toxA virulence genes and blaTEM, blaCTX-M, and tetA antibiotic-resistance genes. Sci. Rep. 10, 1–12.
Andreani, N. 2016. Into the blue: spoilage phenotypes of Pseudomonas fluorescens in food matrices (Doctoral dissertation, University of Padova) Available via https://paduaresearch.cab.unipd.it/9109/1/Andreani_ NadiaAndrea_tesi.pdf.pdf (Accessed 24 June 2025).
Bantawa, K., Rai, K., Subba Limbu, D. and Khanal, H. 2018. Food-borne bacterial pathogens in marketed raw meat of Dharan, eastern Nepal. BMC. Res. Notes 11(618), 1–5.
Bazghandi, S.A., Arzanlou, M., Peeridogaheh, H., Vaez, H., Sahebkar, A. and Khademi, F. 2021. Prevalence of virulence genes and Drug Resistance profiles of Pseudomonas aeruginosa isolated from clinical specimens. Jundishapur. J. Microbiol. 14(8), 1–8.
Ben Mhenni, N., Alberghini, G., Giaccone, V., Truant, A. and Catellani, P. 2023. Prevalence and antibiotic resistance phenotypes of Pseudomonas spp. in fresh fish fillets. Foods 12, 1–12.
Briega, I., Garde, S., Sánchez, C., Rodríguez-Mínguez, E., Picon, A. and Ávila, M. 2025. Evaluation of biofilm production and antibiotic resistance/susceptibility profiles of Pseudomonas spp. isolated from milk and dairy products. Foods 14, 1105.
Bulut, S. 2014. High pressure processing of chicken meat: change in total aerobic counts after pressure treatment and during chilled storage. Br. Microbiol. Res. J. 4(5), 540–549.
Calhoun, C., Geornaras, I. and Zhang, P. 2025. Pseudomonas in Meat Processing Environments. Foods 14, 1615.
Can, H.Y. 2022. Investigation of Pseudomonas species in chicken drumstick samples. Kocatepe. Vet. J. 15(2), 139–143.
Carrizosa, E., Benito, M.J., Ruiz-Moyano, S., Hernández, A., Villalobos, M.D.C., Martín, A. and Córdoba, M.D.G. 2017. Bacterial communities of fresh goat meat packaged in modified atmosphere. Food. Microbiol. 65, 57–63.
Cetin, B., Uran, H. and Konak, M. 2019. Effect of evaporated ethyl pyruvate on reducing Salmonella enteritidis in raw chicken meat. Braz. J. Poult. Sci.,
doi:10.1590/1806-9061-2018-0781
Chen, J., Sun, Z., Jin, J., Wang, F., Yang, Q., Yu, H., Yu, J. and Wang, Y. 2023. Role of siderophore in Pseudomonas fluorescens biofilm formation and spoilage potential function. Food Microbiol. 109, 104151; doi:10.1016/j.fm.2022.104151
Clinical and Laboratory Standard Institute (CLSI). 2020. Performance standards for antimicrobial susceptibility testing. Available via https://www.nih.org.pk/wp-content/uploads/2021/02/CLSI-2020.pdf (Accessed 20 May 2025).
Dokuta, S., Yadoung, S., Hongjaisee, S., Khamnoi, P., Manochomphu, S., Chuttong, B. and Hongsibsong, S. 2025. Seasonal determination of antibiotic-resistant microorganisms and ciprofloxacin residues in pork and chicken meats collected from fresh markets in Chiang Mai, Northern Thailand. Foods 14(2), 1–24.
Dong, N., Liu, C., Hu, Y., Lu, J., Zeng, Y., Chen, G., Chen, S. and Zhang, R. 2022. Emergence of an extensive drug-resistant Pseudomonas aeruginosa strain of chicken origin carrying blaIMP-45, tet(X6), and tmexCD3-toprJ3 on an IncP(R)BL16 plasmid. Microbiol. Spectr. 10(6), L1–X6.
Du, B., Lu, M., Liu, H., Wu, H., Zheng, N., Zhang, Y., Zhao, S., Zhao, Y., Gao, T. and Wang, J. 2023. Pseudomonas isolates from raw milk with high level proteolytic activity display reduced carbon substrate utilization and higher levels of antibiotic resistance. LWT. – Food Sci. Tech. 181, 1–8.
Elbehiry, A., Marzouk, E., Aldubaib, M., Moussa, I., Abalkhail, A., Ibrahem, M., Hamada, M., Sindi, W., Alzaben, F., Almuzaini, A.M., Algammal, A.M. and Rawway, M. 2022. Pseudomonas species prevalence, protein analysis, and antibiotic resistance: an evolving public health challenge. AMB. Express. 12(53), 1–14.
EUCAST. 2022. EUCAST Clinical Breakpoint Tables V.12.0. Available via https://www.eucast.org/clinical_breakpoints/ (Accessed 15 May 2025).
Fakhkhari, P., Tajeddin, E., Azimirad, M., Salmanzadeh-Ahrabi, S., Abdi-Ali, A., Nikmanesh, B., Eshrati, B., Gouya, M.M., Owlia, P., Zali, M.R. and Alebouyeh, M. 2020. Involvement of Pseudomonas aeruginosa in the occurrence of community and hospital acquired diarrhea, and its virulence diversity among the stool and the environmental samples. Int. J. Environ. Health Res. 32(1), 61–71.
Galdino, A.C.M., Branquinha, M.H., Santos, A.L.S. and Viganor, L. 2017. Pseudomonas aeruginosa and its arsenal of proteases: Weapons to battle the host. In Pathophysiological Aspects of Proteases. Eds., Chakraborti, S. and Dhalla, N. Cham, Switzerland: Springer, pp: 381–397. Available via https://link.springer.com/chapter/10.1007/978-981-10-6141-7_16 (Accessed 24 June 2025).
Glantz, M., Rosenlöw, M., Lindmark-Månsson, H., Buhelt Johansen, L., Hartmann, J., Höjer, A., Waak, E., Löfgren, R., Hallin Saedén, K., Svensson, C., Svensson, B., Lindau, J., Rauh, V. and Paulsson, M. 2020. Impact of protease and lipase activities on quality of Swedish raw milk. Int. Dairy. J. 107, 104724; doi:10.1016/j.idairyj.2020.104724
Guibert, F., Rojo-Bezares, B., Espinoza, K., Alonso, C.A., Oporto-Llerena, R., López, M., Taboada-Blanco, C., Sáenz, Y., Pons, M.J. and Ruiz, J. 2024. Antibiotic-resistant Pseudomonas aeruginosa from market meat in Peru. Foodborne Pathog. Dis. 22(11), 761-767; doi:10.1089/fpd.2024.0052
Hassan, W., Ibrahim, A., Shany, S. and Salam, H. 2020. Virulence and resistance determinants in Pseudomonas aeruginosa isolated from pericarditis in diseased broiler chickens in Egypt. J. Adv. Vet. Anim. Res. 7(3), 452–463.
Heir, E., Moen, B., Åsli, A.W., Sunde, M. and Langsrud, S. 2021. Antibiotic resistance and phylogeny of Pseudomonas spp. isolated over three decades from chicken meat in the Norwegian food chain. Microorganisms 9(2), 1–9.
Heredia, N. and García, S. 2018. Animals as sources of food-borne pathogens: a review. Anim. Nutr. 4(3), 250–255.
Hsern Malcolm, T.T., Cheah, Y.K., Radzi, C.W.J.W.M., Kasim, F.A., Kantilal, H.K., Huat John, T.Y., Martinez-Urtaza, J., Nakaguchi, Y., Nishibuchi, M. and Son, R. 2015. Detection and quantification of pathogenic Vibrio parahaemolyticus in shellfish by using multiplex PCR and loop-mediated isothermal amplification assay. Food Control 47, 664–671.
Hudzicki, J. 2009. Kirby-Bauer disk diffusion susceptibility test protocol. ASM 1–23.
Jawher, I.M. and Hassan, M.G. 2023. Antibiotics resistance patterns of Pseudomonas aeruginosa isolated from meat at Mosul city retails. Iraqi. J. Vet. Sci. 37(2), 363–367.
Karanth, S., Feng, S., Patra, D. and Pradhan, A. K. 2023. Linking microbial contamination to food spoilage and food waste: the role of smart packaging, spoilage risk assessments, and date labelling. Front. Microbiol. 14, 1–17.
Lianou, A., Panagou, E. Z. and Nychas, G-J. 2023. Meat safety – Foodborne pathogens and other biological issues. Lawrie’s Meat Science 9th ed, 549–590.
Mapipa, Q., Digban, T.O., Nnolim, N.E. and Nwodo, U.U. 2021. Antibiogram profile and virulence signatures of Pseudomonas aeruginosa isolates recovered from selected agrestic hospital effluents. Sci. Rep. 11, 1–11.
Martínez-Granero, F., Navazo, A., Barahona, E., Redondo-Nieto, M., Rivilla, R. and Martín, M. 2012. The Gac-Rsm and SadB signal transduction pathways converge on AlgU to downregulate motility in Pseudomonas fluorescens. PLos One 7(2), 1–9.
Martins, M.L., Pinto, U.M., Riedel, K. and Vanetti, M.C.D. 2015. Milk-deteriorating exoenzymes from Pseudomonas fluorescens 041 isolated from refrigerated raw milk. Braz. J. Microbiol. 46(1), 207–217.
Meng, L., Liu, H., Lan, T., Dong, L., Hu, H., Zhao, S., Zhang, Y., Zheng, N. and Wang, J. 2020. Antibiotic resistance patterns of Pseudomonas spp. isolated from raw milk revealed by whole genome sequencing. Front. Microbiol. 11, 1005.
Meng, L., Zhang, Y., Liu, H., Zhao, S., Wang, J. and Zheng, N. 2017. Characterization of Pseudomonas spp. and associated proteolytic properties in raw milk stored at low temperatures. Front. Microbiol. 8, 1–7.
Mohamed, H.M.A., Alnasser, S.M., Abd-Elhafeez, H.H., Alotaibi, M., Batiha, G.E.S. and Younis, W. 2022. Detection of β-lactamase resistance and biofilm genes in Pseudomonas species isolated from chickens. Microorganisms 10, 1–19.
Molina, L., Udaondo, Z., Duque, E., Fernández, M., Molina-Santiago, C., Roca, A., Porcel, M., De La Torre, J., Segura, A., Plesiat, P., Jeannot, K. and Ramos, J.L. 2014. Antibiotic resistance determinants in a Pseudomonas putida strain isolated from a hospital. PLos One. 9, e81604.
Moretro, T. and Langsrud, S. 2017. Residential bacteria on surfaces in the food industry and their implications for food safety and quality. Compr. Rev. Food Sci. Food Saf. 16, 1022–1041.
Muinde, P., Maina, J., Momanyi, K., Yamo, V., Mwaniki, J. and Kiiru, J. 2023. Antimicrobial resistant pathogens detected in raw pork and poultry meat in retailing outlets in Kenya. Antibiot 12(613), 1–11.
Narvhus, J.A., Baekkelund, O.N., Tidemann, E.M., Ostlie, H.M. and Abrahamsen, R.K. 2021. Isolates of Pseudomonas spp. from cold-stored raw milk show variation in proteolytic and lipolytic properties. Int. Dairy. 123, 105049.
Oh, S.R., Kang, I., Oh, M.H. and Ha, S.D. 2014. Inhibitory effect of chlorine and ultraviolet radiation on growth of Listeria monocytogenes in chicken breast and development of predictive growth models. Poult. Sci. 93(1), 200–207.
Pang, Z., Raudonis, R., Glick, B.R., Lin, T.J. and Cheng, Z. 2019. Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 37, 177–192.
Quintieri, L., Fanelli, F. and Caputo, L. 2019. Antibiotic Resistant Pseudomonas spp. spoilers in fresh dairy products: an underestimated risk and the control strategies. Foods 8(372), 1–32.
Raposo, A., Perez, E., De Faria, C.T., Ferrus, M.A. and Carrascosa, C. 2017. Food Spoilage by Pseudomonas spp.—An overview. In Foodborne Pathogens and Antibiotic Resistance. Singh, O.V. and Ed. Hoboken, NJ: John Wiley and Sons, Inc., pp: 41–58.
Rawat, S. 2018. Antibiotic resistance in Pseudomonas species isolated from poultry. J. Food. Saf. Qual. 9(2), 567–572.
Redondo-Nieto, M., Barret, M., Morrissey, J., Germaine, K., Martínez-Granero, F., Barahona, E., Navazo, A., Sánchez-Contreras, M., Moynihan, J.A., Muriel, C., Dowling, D., O’Gara, F., Martín, M. and Rivilla, R. 2013. Genome sequence reveals that Pseudomonas fluorescens F113 possesses a large and diverse array of systems for rhizosphere function and host interaction. BMC. Genomics. 14(54), F1–17.
Salwoom, L., Raja-Abd-Rahman, R.N.Z., Salleh, A.B., Mohd-Shariff, F., Convey, P., Pearce, D. and Mohamad-Ali, M.S. 2019. Isolation, characterisation, and lipase production of cold-adapted bacterial strain Pseudomonas sp. LSK25 isolated from Signy Island, Antarctica. Molecules 24(715), 1–14.
Sato, H. and Frank, D.W. 2004. ExoU is a potent intracellular phospholipase. Mol. Microbiol. 53(5), 1279–1290.
Shahat, H.S., Mohamed, H.M.A., Abd Al-azeem, M.W. and Nasef, S.A. 2019. Molecular detection of some virulence genes in Pseudomonas aeruginosa isolated from chicken embryos and broilers with regard to disinfectant resistance. Int. J. Vet. Sci. 2(2), 52–70.
Syafuddin Tan, M.Z., Bakar, A.N. and Ahmad, Y. 2023. Malaysia's chicken shortage, a solution proposal through consumerism advocacy. J. Agri. Mark. 10(1), 49–67.
Tacconelli, E., Magrini, N., Carmeli, Y., Harbarth, S., Kahlmeter, G., Kluytmans, J., Mendelson, M., Pulcini, C., Singh, N., and Theuretzbacher, U., 2017. Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics. World Health Organization 1–7. Available via http://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf (Accessed 13 July 2025).
Tigabie, M., Assefa, M., Gashaw, Y., Amare, A., Ambachew, A., Biset, S. and Moges, F. 2025. Prevalence and antibiotic resistance patterns of Pseudomonas aeruginosa and Acinetobacter baumannii strains isolated from chicken droppings on poultry farms in Gondar city, Northwest Ethiopia. Sci. One Health 4(1), 1–10.
Wangchinda, W., Wu, J.Y., Abbo, L.M., Ackley, R., Bartley, P., Gilboa, M., Harrington, J., Jaffa, R., Klatt, M.E., Kline, E.G., Kubat, R.C., Lepak, A.J., Mccreary, E.K., Miller, W.R., Pearson, J.C., Shah, S., Tran, T.T., Vega, A., Vendetti, M., Yucel, E., Pogue, J.M. and Shields, R.K. 2025. Effectiveness of imipenem-relebactam for multidrug-resistant Pseudomonas aeruginosa in pneumonia and bloodstream infections in the United States (MIRAGE). Antimicrob. Agents Chemother. 69(12), e0132525.
World Health Organization (WHO). 2017. Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics. Geneva, Switzerland. Available via https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed (Accessed 13 July 2025).
Wickramasinghe, N.N., Ravensdale, J., Coorey, R., Chandry, S.P. and Dykes, G.A. 2019. The predominance of psychrotrophic Pseudomonads on aerobically stored chilled red meat. Compr. Rev. Food. Sci. Food. Saf. 18, 1622–1635.
Xu, J., Moore, J.E., Murphy, P.G., Millar, B.C., Redmond, A.O.B. and Elborn, J.S. 2004. Molecular (PCR) detection of Pseudomonas spp. other than P. aeruginosa directly from the sputum of adults and children with cystic fibrosis. Br. J. Biomed. Sci. 61(3), 147–149.
Xu, Y., Chen, W., You, C. and Liu, Z. 2017. Development of a multiplex PCR Assay for detection of Pseudomonas fluorescens with biofilm formation ability. Food Sci. 82, 2337–2342.
Zhang, S. and Lv, J. 2014. Purification and properties of heat-stable extracellular protease from Pseudomonads fluorescens BJ-10 J. Food Sci. Technol. 51(6), 1185–1190.

















