| Research Article | ||
Open Vet. J.. 2026; 16(2): 1338-1350
Open Veterinary Journal, (2026), Vol. 16(2): 1338-1350 Review Article Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in EgyptEman AbdelMenamm Shosha1*, Ali Mahmoud Zanaty2, Ahmed Fotouh Abdallah3 and Ibrahim Eldaghayes41Virology Department, Faculty of Veterinary Medicine, New Valley University, El-Khargia, Egypt 2Reference Laboratory for Quality Control on Poultry, Animal Health Institute, Agriculture Research Center (ARC), Giza, Egypt 3Pathology and Clinical Pathology Department, Faculty of Veterinary Medicine, New Valley University, El-Khargia, Egypt 4Department of Microbiology and Parasitology, Faculty of Veterinary Medicine, University of Tripoli, Tripoli, Libya *Corresponding Author: Eman AbdelMenamm Shosha. Virology Department, Faculty of Veterinary Medicine, New Valley University, El-Khargia, Egypt. Email: Emanshosha25 [at] gmail.com Submitted: 12/11/2025 Revised: 15/01/2026 Accepted: 29/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
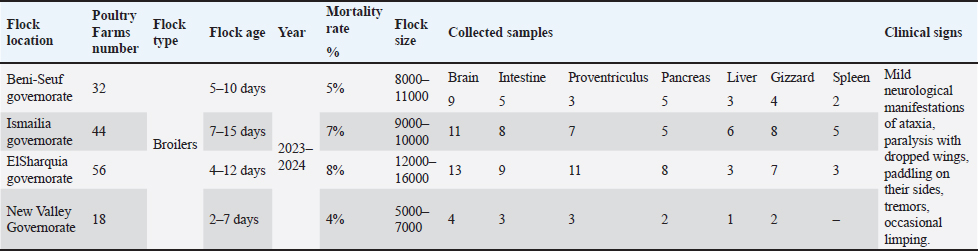
ABSTRACTBackground: Avian encephalomyelitis (AE) is an imperative viral disease of avian species worldwide caused by the AE virus (AEV). Aim: This retrospective study aimed to genetically characterize field AEV strains circulating in the local governorates of Egypt and assess their genetic divergence in comparison with commercially available vaccines. To the best of our knowledge, this is the first described study of AEV-VP1 gene sequencing and phylogeny in broilers in Egypt. Methods: A total of 150 tissue specimens from broilers of four Egyptian governorates were used for isolation and genetic characterization. The samples were tested for other pathogens, including egg drop syndrome virus, Newcastle disease virus, avian reovirus, avian reticuloendotheliosis virus, infectious bronchitis virus, Chicken anemia virus, and Marek’s disease virus. Results: Young chicks showed neurological symptoms and a mortality rate of 4%–8%. Only crop impaction and distended proventriculus were observed in sick chicks. Histopathological findings revealed typical lesions of lymphocytic encephalomyelitis in the cerebrum with mononuclear cuffing of perivascular spaces in the cortex and medulla. The proventriculus revealed mild lymphocytic infiltration within the submucosa. Typical pathological changes in leg rigidity and encephalomalacia were observed in specific pathogen-free embryonated chicken eggs (SPF-ECE). Regarding reverse transcriptase quantitative polymerase chain reaction (RT-qPCR), 8 (8/150) samples were positive with a prevalence rate of 5.3%, and the screening results were negative for other viral infections. The 4 selected positive samples were amplified using a one-step conventional RT-PCR assay for partial sequencing of the AEV-VP1 gene. Interestingly, the phylogenetic analysis exhibited that AEV isolates belong to cluster II within the same clade of Chinese strains and have a genetic similarity to Chinese, Iranian, American vaccinal, and British isolates, with 94%–98%a nucleotide similarity, and 96%–99% amino acid identity. Conclusion: Continuous monitoring and potentially updating the vaccination strategies in a timely manner are necessary to ensure vaccine effectiveness as an efficient control strategy in poultry flocks. Keywords: Avian encephalomyelitis virus, Young chicks, VP1 gene, Sequencing, Broilers. IntroductionAvian encephalomyelitis (AE) is a highly contagious viral disease that affects several avian species, including chickens, turkeys, pigeons, and quails. The disease is caused by the AE virus (AEV), which spreads readily among susceptible birds (Swayne et al., 2017; Hauck et al., 2017). AE typically manifests at an early age in broilers, most commonly between 1 and 3 weeks of age. Clinical disease may appear as early as 3–7 days after hatching, particularly in cases of vertical transmission from infected breeder hens. AEV is clinically characterized by typical neurological symptoms, such as ataxia, paresis, paralysis, muscular dystrophy, abnormal gait, and rapid head and neck tremors, with high morbidity and variable mortality in young infected chicks (Ali et al., 2021; Zhang et al., 2023). In addition, tremovirus A comprises two distinct strain categories: a natural enterotropic strain that usually infects the offspring of chicks and a neurotropic embryo-adapted strain. AEV transmission can occur horizontally via the fecal-oral route and vertically by infected embryonated eggs. In particular, vertical transmission leads to the development of pronounced neurological signs in newly hatched chicks and poses a serious risk to the poultry sector. The disease prevalence in affected birds ranged from 40% to 60%, with an average mortality rate of 25%. In contrast, AVE infection in adult birds is typically subclinical and is mainly associated with reduced egg production and decreased hatchability of fertile eggs, without neurological manifestations (Torre et al., 2018; Zhang et al., 2023). AEV is structurally classified within the genus Tremovirus of the family Picornaviridae (Ali et al., 2021). Its genome consists of a single-stranded, positive-sense RNA approximately 7.5 kb in length and contains a single large open reading frame (ORF) that encodes a polyprotein precursor (Ali et al., 2021). Similar to other picornaviruses, this polyprotein precursor is subsequently cleaved by intracellular proteases into three primary precursor proteins—P1, P2, and P3—yielding 11 structural and non-structural proteins. The P1 region encodes the four major structural proteins VP4, VP2, VP3, and VP1, whereas the P2 and P3 regions encode the nonstructural proteins 2A, 2B, 2C, 3A, 3B, 3C, and 3D (Lin et al., 2018). Notably, the VP1 capsid protein within the P1 region serves as a genetic diagnostic marker for AEV and constitutes the principal immunogenic protein responsible for inducing protective host immune responses (Wei et al., 2008; Hauck et al., 2017). Since the AEV outbreak was identified in the United States of America in 1930, it has achieved a worldwide endemic distribution, with sustained circulation reported across North America, Europe, Asia, Africa, the Middle East, and South America, particularly in regions with intensive poultry production (Braune and Gentry, 1971; Wang et al., 2023). Generally, AEV infection is confirmed either by detecting the virus through molecular diagnostic methods, including polymerase chain reaction, or by isolating the virus (Haryanto et al., 2016; Senties-Cue et al., 2016; Goto et al., 2019). Furthermore, serological assays, such as agar gel diffusion, virus neutralization, and chicken embryo susceptibility tests, can support diagnosis when interpreted alongside clinical findings (Pohjola et al., 2017; Taunde et al., 2017; Ali et al., 2021; Gethoffer et al., 2021). At present, no specific antiviral treatment is available; therefore, vaccination regimes remain the most effective strategy for AEV control and are significant in minimizing losses (Lin et al., 2018). Commercial vaccines used in breeder flocks are effective and widely recommended because they prevent viral infection during egg production and provide adequate maternal immunity to protect the progeny (Sarma et al., 2019). This study reports various cases of leg paralysis and head tremors with variable mortality. Taken together, this retrospective study was conducted to molecularly characterize the circulating field AEV strains in the local governorates of Egypt and assess their genetic divergence compared to commercially available vaccines. To the best of our knowledge, this is the first report of AEV-VP1 gene sequencing, providing informative data about the genetic diversity and phylogenetic characteristics of AEV-VP1 within respective chicken flocks. Materials and MethodsHistory of the investigated flocks and sample processingBetween 2023 and 2024, a total of 150 specimens were collected from an equal number of broiler flocks in four Egyptian governorates (New Valley, Beni-Suef, Ismailia, and El-Sharkia). The affected flocks exhibited mild neurological signs, including ataxia, paralysis-associated wing drooping, lateral recumbency with paddling movements, tremors, intermittent lameness, and fluctuating levels of morbidity and mortality. The sample size was determined based on the availability of clinically suspected broiler flocks exhibiting neurological manifestations across four governorates in Egypt. These flocks were included to ensure adequate geographic representation and enhance the likelihood of detecting circulating AEV field strains. No specific necropsy pictures were observed in the infected or freshly dead birds. Mostly, the breeders’ flocks have previously been vaccinated with the combined live vaccine containing AEV and Fowlpox virus (POXIMUNE® AE, Ceva Animal Health company, containing the Calnek strain of AEV, route of administration for POXIMUNE® AE vaccine is the wing-web method at 8–12 weeks of age). Table 1 lists the characteristic epidemiological characteristics of the examined broilers. The visceral tissue specimens were collected from the brain, intestine, liver, spleen, proventriculus, gizzard, and pancreas of each flock and then preserved at 80◦C until further use for virus isolation and molecular testing. The Furthermore, the collected specimens were processed as pooled samples and then diluted using sterile phosphate-buffered saline (PBS) at pH 7.4 to generate 10% (w/v) homogenates, including 10% antibiotic solution. The homogenates were then cooled and centrifuged at 3500 × g for 10 minutes after overnight incubation at 4°C. Finally, the supernatants were collected for further RNA extraction (Goto et al., 2019). Table 1. Epidemiological information of broilers assessed for AEV in the four local governorates.
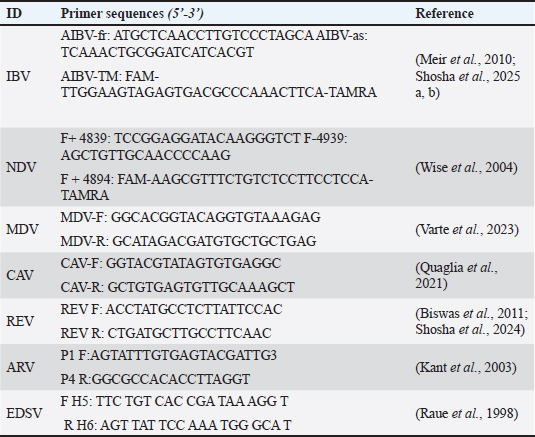
Histopathological examinationTissue specimens (brain, proventriculus, and gizzard) were collected microscopically immediately after necropsy and fixed in neutral-buffered formalin (10%) for 24–48 hours to ensure adequate tissue preservation. Tissues were trimmed, washed, dehydrated in a graded ethanol series (70%, 80%, 90%, 95%, and 100% ethanol), cleared, and embedded in paraffin wax using standard histological procedures. After sectioning the paraffin blocks at a thickness of 4–5 μm using a microtome. The obtained sections were mounted on glass slides, deparaffinized, rehydrated through descending grades of ethanol, and stained with hematoxylin and eosin (H&E) according to routine protocols. Finally, the stained sections were examined microscopically for histopathological alterations using a light microscope (Fotouh et al., 2024). AEV isolationThe supernatants of tissue homogenate (0.2 ml) were inoculated via the yolk sacs (5 ECE/sample) into 5–7-day-old specific pathogen-free embryonated chicken eggs (SPF-ECE), purchased from SPF farms located in Koom Oshiem region, Fayoum governorate, Egypt (Hoekstra, 1964), and then incubated at 37°C. The chicks were monitored daily for any AEV-associated signs during the first 10 days post-inoculation (PI). Consequently, once clinical signs were monitored, the six brain homogenates collected from six infected chicks were further centrifuged at 12,000 rpm for 10 minutes. Finally, the AEV isolate remained in the subsequent supernatants. A previously confirmed AEV reference strain (molecularly verified by RT-PCR) was used as a positive control to validate the virus isolation procedure. Ten SPF chicken embryos were used as the negative control with 0.1 M PBS (pH=7.4). RNA extraction and conventional reverse transcriptase polymerase chain reactionParticularly, the viral RNA was extracted promptly from positive samples using the QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations. A subset of confirmed positive isolates (n=8) was subjected to partial amplification of the VP1 gene using a conventional reverse transcription-polymerase chain reaction assay kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Amplification was performed using a specific primer pair (AEV-Forward: 5′-GAATTAGCTCCTGGTAAACCTCG-3′ and AEV-Reverse: 5′-CTCTATCGCAACACCCTCAGG-3′), targeting a 288-bp fragment of the VP1 genomic region (Zhang et al., 2024). Differential of AVE from other pathogens using RT-qPCRMoreover, all PCR-positive samples were screened for avian viral pathogens, including Egg drop syndrome virus (EDSV), Avian reovirus (ARV), Newcastle disease virus (NDV), avian reticuloendotheliosis virus (REV), Infectious bronchitis virus (IBV), Chicken anemia virus (CAV), and Marek’s disease virus (MDV), were also examined by RT-qPCR protocols, to differentially confirm the presence of AEV (Table 2) (Raue et al., 1998; Kant et al., 2003; Wise et al., 2004; Meir et al., 2010; Biswas et al., 2011; Mosad et al., 2021; El-Khabaz et al., 2021; Quaglia et al., 2021; Varte et al., 2023; Zhang et al., 2024; Shosha et al., 2024; Shosha et al., 2025). Table 2. Consensus oligonucleotide primers were subjected to RT-PCR amplification of other related pathogens.
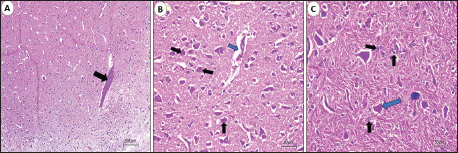
VP1 gene sequencing, phylogenetic analysis, and recombination eventsThe size-specific PCR amplicons were visualized by gel electrophoresis and subsequently purified using the QIAquick Gel Extraction kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol for further confirmation. In addition, the current purified amplicons were subsequently subjected to direct sequencing with the BigDye™ Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Waltham, MA) and an automated ABI 3500 × L Genetic Analyzer (Life Technologies, California). The resulting nucleotide and amino acid sequences were assembled, annotated, and aligned with other representative published strains of AEV vaccines, as well as available reference strains representing all AEV genotypes, using the BioEdit software (Clustal W alignment) (Kumar et al., 2016). Notably, the current nucleotide sequences of the four newly characterized strains were identified as AEV and subsequently deposited in GenBank under their accession numbers. Finally, the phylogenetic analyses of AEV were generated based on the maximum likelihood methodology (Bootstrap in 1000 replicates) using the Kimura 2-parameter model (Tamura et al., 2013) using MEGA software version 7.0. The recombination events of each strain were investigated to check for any new AEV-recombinant strains (RDP-5 Program, version 4.97; Martin et al., 2009) using default detection parameters with a significance threshold of p < 0.05. For comparison, we used the following algorithms: RDP5, BootScan, MaxChi, GENECONV, SiScan, Chimera, LARD, Phyl-Pro, and 3Seq (Martin et al., 2005). The recombination events were used through 4 independent approaches or more with statistically significant p-values, which were considered proven as detectable positive occurrences. Ethical approvalAll sampling and animal experiments in this study were conducted in accordance with the ethical guidelines approved by the New Valley Research Ethics Committee, Faculty of Veterinary Medicine, New Valley University (approval no. 02-3-3-2024-11). ResultsClinical findings, necropsy and histopathology findingsRegarding the AEV prevalence survey, the virus was detected in 8 out of 150 positive flocks, with a prevalence rate of 5.3%. The examined infected chicks exhibited neurological symptoms of varying intensity, including depression, incoordination, ataxia, tremors, leg stretching, lying on their sides, and paralysis with dropped wings. The mortality rate ranged from 4% to 8%, whereas the morbidity rate was approximately 30%, indicating a measurable impact at the flock level. No remarkable macroscopic lesions were observed in the AEV-infected chicks. Mild brain congestion or edema was observed in a few broiler cases. A crop impaction or a distended proventriculus and gizzard were also observed. Microscopic examination of the brain tissue, particularly the cerebrum (Fig. 1), revealed lesions consistent with viral lymphocytic encephalomyelitis in the majority of infected broilers. Moderate to marked perivascular mononuclear cell cuffing was observed in the cerebral cortex and medulla, predominantly composed of lymphocytes with occasional macrophages, expanding the perivascular spaces and extending into the surrounding neuropil. Diffuse and perivascular lymphocytic infiltration were evident within the cortical gray matter, indicating active inflammatory involvement.
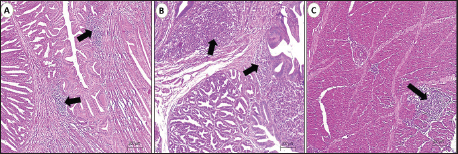
Fig. 1. Photomicrograph of the cerebrum of AEV-infected broiler chickens. (A) Lymphocytic encephalitis with perivascular cuffing (arrow). (B) Central chromatolysis of neurons (black arrows) and vasogenic edema (blue arrow). (c) Central chromatolysis of neurons (blue arrow) and astrocytosis (black arrows) (hematoxylin and eosin stain, Scale bar 200 μm). Neuronal degeneration and necrosis, accompanied by neuronophagia, were frequently observed, characterized by focal aggregation of activated microglial cells phagocytosing degenerated or necrotic neurons, particularly within the cortical layers. Additionally, affected areas demonstrated gliosis, which was manifested by the conspicuous proliferation and hypertrophy of microglial cells and astrocytes, reflecting a reactive glial response to neuronal injury. Prominent central chromatolysis was noted in multiple neurons, characterized by peripheral displacement of the neuronal nucleus and dissolution or loss of Nissl substance from the central cytoplasm, indicative of viral neurotropism-associated axonal injury and metabolic stress. Mild interstitial edema was present within the neuropil in some sections, further contributing to tissue disruption. Histopathological examination of the proventriculus (Fig. 2) revealed mild to moderate lymphocytic infiltration within the lamina propria and submucosa in approximately 50% of the examined samples. This infiltration was occasionally associated with submucosal edema and mild interglandular connective tissue expansion. In certain cases, the mucosa exhibited degenerative changes, including focal glandular epithelial atrophy and mild epithelial vacuolation, suggestive of chronic inflammatory insult. Histological sections of the gizzard revealed multifocal lymphocytic infiltration involving the submucosa and extending into the muscular layer. These inflammatory foci often accompanied by mild vascular congestion of mucosal and submucosal capillaries, without evidence of hemorrhage or necrosis. In most cases, the koilin layer remained largely intact, indicating a predominantly inflammatory rather than erosive process.
Fig. 2. Photomicrographs of the proventriculus and gizzard of AEV-infected broiler chickens. (A&B) Mild lymphocytic infiltrations (arrows) of the mucosa and submucosa of the proventriculus. (C) Focal lymphocytic infiltrations (arrows) in the muscular layers of the gizzard (hematoxylin and eosin stain, Scale bar 200μm). SPF-ECE isolation and genetic identificationAfter 10 days of SPF-ECE post incubation, AEV-inoculated embryos demonstrated typical pathological alterations in the form of leg rigidity and encephalomalacia. In contrast, only 8 out of 150 samples (5.3%) were considered positive using RT-PCR assay targeting the VP1 gene. Moreover, 4 (4/8) positive samples were successfully amplified using a one-step RT-PCR assay targeting AEV-VP1 gene partial sequencing and genetic identification (Fig. 3). Importantly, the RT-PCR screening results were negative for the other common viral infections mentioned above (NDV, EDSV, ARV, REV, CAV, MDV, and IBV).
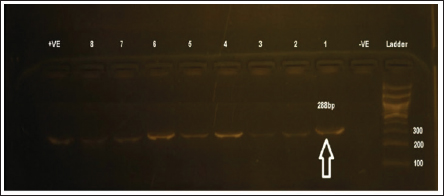
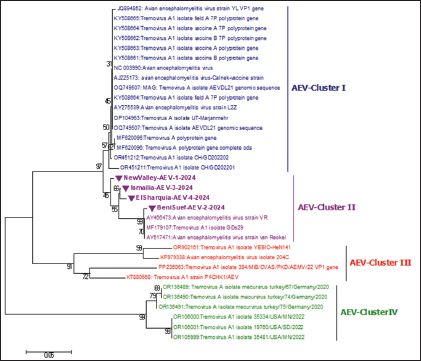
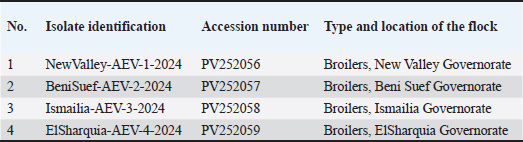
Fig. 3. Identification of the VP1 gene by RT-PCR assay of 288-bp PCR products for AEV-positive isolates. (A) Lanes 1- 8: AEV-positive field samples; the first lane is the negative control (PBS); the last lane is the positive control (Commercial vaccine). RT-PCR, reverse transcription polymerase chain reaction; PBS, phosphate-buffered saline. Lane M: represents a 100-bp ladder as a size standard. Sequencing, phylogenetic analysis of VP1 gene, and recombination eventsThe partial sequencing and phylogenetic evaluation targeted the VP1 gene of the four selected isolates (NewValley-AEV-1-2024, BeniSuef-AEV-2-2024, Ismailia-AEV-3-2024, and ElSharquia-AEV-4-2024), confirming that these Egyptian strains belonged to AEV-cluster II (Fig. 4). Besides, these four isolates were submitted to the GenBank database under accession numbers (PV252056–PV252059) (Table 3). Based on the phylogenetic analysis, the four current isolates belonged to a similar branch to the Chinese isolates: van Reokel, VR, and GDs29. The phylogenetic analysis was performed on various AEV strains available in GenBank to distinguish between field and vaccinal strains. Notably, the genetic findings exhibited that our NewValley-AEV-1-2024 strain was closely related to the AEV strain YL-VP1, AEV-CH/GD202202, AEV strain L2Z (Chinese isolates), AEV-UT-Marjanmehr (Iranian isolate), AEV-vaccine/B, AEV-isolate vaccine/A, NC/003990-AEV, AEV-vaccine/B/7P, AEV-vaccine/A/7P (American isolates), and AEV-Calnek-vaccine strain (British isolate, also used in combined vaccine in Egypt) with 98% nucleotide identity, and (97%–98%) amino acid similarity (Table 4).
Fig. 4. A phylogenetic tree based on partial sequencing was generated using a 288-bp fragment of the VP1 gene from multiple AEV strains, along with GenBank reference sequences. The VP1 gene phylogeny indicated that the four AEV isolates (marked with violet triangles) clustered within cluster II. The tree was constructed using the Maximum Likelihood method with 1,000 bootstrap replicates in MEGA 7.0 software, applying the Kimura 2-parameter model. Table 3. Data of recent isolates used for the partial sequencing analysis of the AEV-VP1 gene.
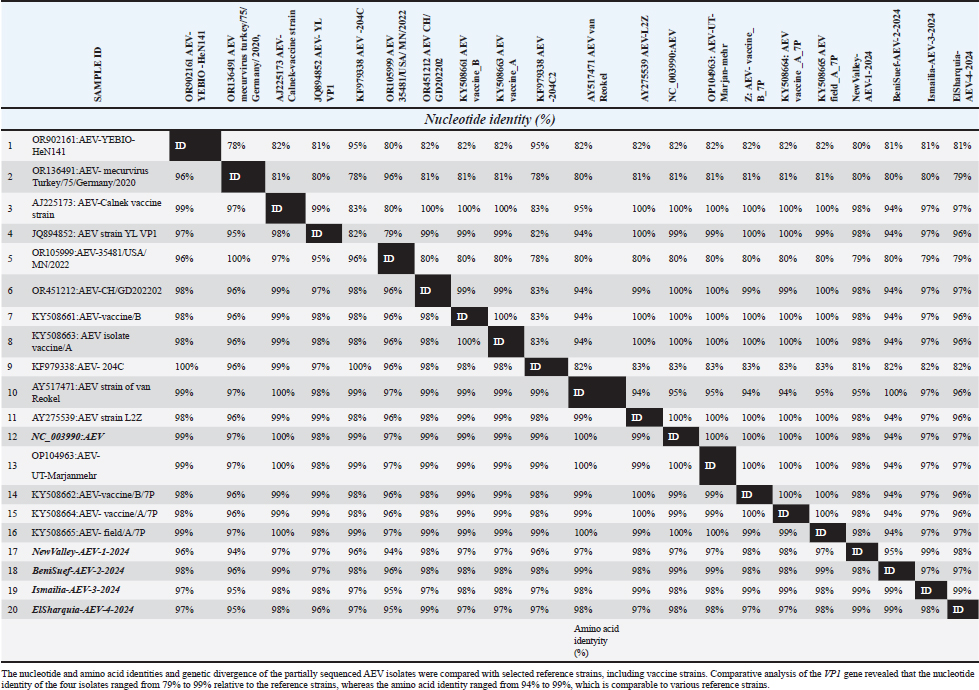
Table 4. Comparison of the nucleotide and amino acid identities of the partially sequenced AEV isolates with selected reference strains.
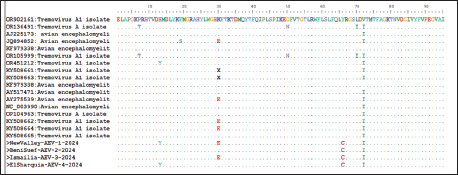
Likewise, our isolates, including Ismailia-AEV-3-2024 and ElSharquia-AEV-4-2024, were clustered phylogenetically closely related to strains detected in China (YL-VP1, AEV-CH/GD202202, AEV strain L2Z), Iran (AEV-UT-Marjanmehr), USA (AEV-vaccine/B, AEV-isolate vaccine/A, NC/003990-AEV, AEV-vaccine/B/7P, AEV-vaccine/A/7P), and UK (AEV-Calnek-vaccine strain) with (96%–97%) nucleotide identity, and (96%–99%) amino acid homology. Similarly, the BeniSuef-AEV-2-2024 strain showed a close phylogenetic relationship with the AEV strains circulating in Asia and North America, exhibiting 94% nucleotide identity and 97%–99% similarity at the amino acid level. Meanwhile, the current four isolates were distinctly apparent from AEV-35481/USA/MN/2022 (USA), AEV-YEBIO-HeN141 (China), AEV-mecurvirus turkey/75/Germany/2020 (Germany), and AEV isolate 204C (Hong Kang); shared (79%–80%), (80%–81%), (79%–80%), (80%–82%); respectively, nucleotide similarity and (96%–98%), (94%–96%), (94%–96%), (96%–98%); respectively, amino acid similarity (Table 4). Also, the amino acid identity analysis of the VP1 gene of NewValley-AEV-1-2024, BeniSuef-AEV-2-2024, Ismailia-AEV-3-2024, and ElSharquia-AEV-4-2024 isolates revealed 98% homology to each other and nucleotide identity of 99%. Regarding mutation analysis, multiple VP1-amino acids sequence comparison showed that the AEV four strains share amino acid mutations compared with other GenBank sequences (Fig. 5). At the amino-acid analysis level, NewValley-AEV-1-2024 harbored two substitutions, namely, glutamic acid (E) at position 30 and tyrosine (Y) at position 13, when compared with the reference strains. Ismailia-AEV-3-2024 retained a single substitution involving E at position 30, whereas ElSharquia-AEV-4-2024 showed a Y substitution at position 13 relative to the classical sequences. Notably, all four Egyptian isolates shared a cysteine (C) substitution, which was absent in the classical reference strains, indicating a common molecular signature. Similarly, the isoleucine residue (I) at position 72 was substituted across all four isolates. In particular, recombination analysis of the partial VP1 gene (288 bp) within our four AEV Egyptian isolates was performed using RDP5 as a preliminary screening approach. No putative recombination signals were detected within the analyzed fragment; however, the short sequence length limits the statistical power of recombination detection.
Fig. 5. Alignment of the VP1 amino acid sequences from the four Egyptian AEV isolates and the reference strains. The top row shows the amino acid positions in the YEBIO-HeN141 VP1 sequence. Letters denote amino acid substitutions, dots (.) represent identical amino acids, and dashes (−) indicate alignment gaps. DiscussionIn Egypt, there are many viral infections, causing multiple outbreaks with considerable impact on animal and poultry health (Abd El Hafez et al., 2021; Badr et al., 2022; Ramzy et al., 2024). These viral outbreaks have predominantly resulted from RNA viruses, causing remarkably devastating economic losses in the poultry sector (Amal et al., 2019; Abodalal et al., 2021; Ahmed et al., 2025). Since its first recorded outbreak in the USA in 1932, AEV has been a major concern and a globally prevalent disease causing substantial economic losses to the poultry sector in Africa, Asia, Australia, Europe, and the Americas (Xue et al., 2016; Zhang et al., 2023; Wang et al., 2023; Zhang et al., 2024). AEV has persisted on poultry farms for extended periods despite routine vaccination across different districts of Egypt, which is accomplished by vaccinating poultry flocks against AEV during the growing period. At present, AEV prevention is predominantly based on vaccine strategies. Thus, strengthening the farm’s daily feeding practices and biosecurity protocols, enhancing the breeding environment, and preventing secondary infection are essential (Lin et al., 2018; Sarma et al., 2019). This study aimed to isolate and genetically characterize all available AEV isolate sequences and other commercially available vaccines in four Egyptian governorates to guide AEV prevention and control. To the best of our knowledge, no report has been performed mainly on AEV in broilers in Egypt. The present study was conducted in broiler flocks due to the increasing occurrence of neurological manifestations and economic losses reported in young broilers in Egypt. Broilers represent a highly sensitive indicator of circulating field strains because they are typically not routinely vaccinated against AEV and therefore reflect exposure to natural viruses. Consequently, broilers were selected to better characterize the currently naturally circulating field AEV strains and to evaluate their genetic relationship with vaccine strains used in breeder flocks. As depicted in our findings, the AEV-positive flocks with a prevalence rate of 5.3% showed clinical mild neurological symptoms in young infected chicks, whereas in adult birds, they only showed transient decreases in egg productivity and hatchability, with a mortality rate ranging from 4% to 8% with non-specific necropsy in the investigated birds. These current findings came in accordance with previously reported clinical manifestations (Taunde et al., 2017; Gethoffer et al., 2021; Zhang et al., 2024; Al-Hammadi and Al-Rasheed 2024; Temeeyasen et al., 2024; Xu et al., 2025). Generally, AEV-gross lesions were mild, which is in agreement with previous evidence that the disease had a preponderant clinical presentation of functional impairment of the central nervous system over enormous macroscopic pathology (Temeeyasen et al., 2024; Xu et al., 2025). The most notable gross changes were impacted crops or gizzards, which are responsible for feeding difficulties due to tremors or bird paralysis. Additionally, these changes are considered to be secondary manifestations of neurogenic dysfunction and have been reported in AEV infections (Xu et al., 2025). The apparent lack of visible lesions within the brain despite extreme neurologic presentations reflects the requirement for microscopic examination in the diagnosis of AEV. This dilemma of AEV diagnosis was mainly histopathology and accurate molecular confirmation (Torre et al., 2018; Ghalyanchilangeroudi et al., 2024). Regarding the pathological dissection of the dead birds, changes in lymphocytic encephalomyelitis, particularly in the cerebrum, including perivascular lymphocytic cuffing, neuronal degeneration, neuronophagia, and gliosis, were observed. These lesions were the same as the current pathological situation previously described in the AEV experimental and natural infection (Hauck et al., 2017; Zhang et al., 2024). One of the characteristic lesions identified in this study was central chromatolysis in motor neurons of the cerebrum, a normal change indicating axonal damage or viral-mediated metabolic stress. The observations in this study also align with the findings of previous studies (Temeeyasen et al., 2024; Ghalyanchilangeroudi et al., 2024). This feature was observed in AEV as a response to viral replication in neurons and other neurotropic viral infections (Diab et al., 2019; Mohamed et al., 2025). Although the CNS remains the predominant target, slight to moderate lesions, including submucosal lymphocytic infiltration and glandular epithelial degeneration, were observed in the proventriculus and gizzard. They can point to viral effects on the enteric nervous system or ganglia, as AEV grows in the peripheral ganglia and autonomic plexuses (Shafren and Tannock, 1991). Concerning SPF-ECE inoculation, AEV-inoculated embryos exhibited typical pathological changes in the form of leg stiffness and encephalomalacia. These subsequent results were in line with those of previous studies (Shafren and Tannock, 1991; Taunde et al., 2017; Goto et al., 2019). More importantly, the RT-PCR results with AEV-specific primers revealed 8 positive samples. These current molecular results are closely similar to those of previous studies (Liu et al., 2014; Goto et al., 2019; Zhang et al., 2024; Ramzy et al., 2024). Subsequently, no co-infection with NDV, IBV, ILT, and IBDV was identified. These subsequent findings are not consistent with the reports of Zhang et al. (2024), who reported the prevalence rates of Mycoplasma synoviae and Gyrovirus galga1 as 5.71% and 2.86%, respectively. Particularly, to investigate the evolutionary relationships within AEV strains, the VP1 gene phylogenetic tree, the main structural protein targeted by the host immune response, was constructed in our AEV isolates. Based on the full genome analysis, AEV strains can be clustered into four main clades. According to the phylogenetic analysis, VP1 is considered the most immunogenic protein of AEV, playing a key role in inducing protective neutralizing antibodies, whereas VP2 (nonstructural proteins) is associated with strain differentiation and is implicated in distinguishing naturally occurring enterotropic field viruses from embryo-adapted neurotropic variants (Wei et al., 2008; Hauck et al., 2017; Zhang et al., 2024). The phylogenetic analysis based on partial gene sequencing revealed that four AEV isolates, including Chinese isolates, belonged to cluster II. Our findings differ from those of Al-Hammadi and Al-Rasheed (2024), who recorded that the KSA strain is clustered within a clade similar to that of a European strain. Moreover, these results are not in line with those of Temeeyasen et al. (2024), who indicated that the three novel AEV isolates were clustered into a unique cluster that was genetically distinct from the previously published AEVs. Specifically, the four current isolates showed a strong genetic similarity to Chinese isolates, Iranian isolate, American vaccinal isolates, and British isolate, with a nucleotide identity percentage of 94%–98%, and based on the amino acid identity, the percentages were 96%–99%, respectively, when compared with other referential AEV strains retrieved from GenBank. Similarly, Zhang et al. (2024) demonstrated that the AEV isolates shared 83.2%–98.0% at the nucleotide and 97.8%–99.6% at the amino level of the VP1 protein. The following results were in accordance with those of Al-Hammadi and Al-Rasheed (2024), who noted that the nucleotide similarities of the AEV/Al-Hassa/2010/KSA isolate to Chinese and European strains were between 94.6% and 99.6%. In the same line, Temeeyasen et al. (2024) reported that the nucleotide identities with the published sequences were 78.3%–79.7% and 93.3%–95.2%, respectively. Collectively, the high nucleotide (94%–98%) and amino acid (96%–99%) identities observed within the VP1 gene of four current isolates relative to Asian and Middle Eastern strains (Chinese and Iranian isolates) indicate a shared evolutionary ancestry. In addition, the close phylogenetic proximity of these Egyptian isolates to both field and vaccine-related strains from China and Iran suggests that the global dissemination of ancestral AEV lineages is potentially mediated through vertically infected breeder hens and long-established poultry production lines. Besides, the clustering of Egyptian AEV isolates within cluster II, which is closely related to the AEV-Calnek-vaccinal strain used in Egypt, suggests that the circulating field strains may originate from vertically transmitted vaccine-derived viruses from breeders or horizontally transmitted through environmental contamination. Particularly, the selection of the four isolates was justified by several considerations. First, all isolates showed robust AEV–specific RT-PCR amplification with low Ct values, indicating a substantial viral load. Second, an initial BLAST analysis of the partial VP1 gene demonstrated a high degree of genetic relatedness to recently reported AEV strains circulating in China and Iran. Third, these isolates were obtained from commercial broiler flocks experiencing evident clinical manifestations and production losses, underscoring their epidemiological significance. Finally, the isolates were chosen to ensure broad geographic coverage, representing El-Sharquia Governorate in northern Egypt (a densely populated poultry area), Ismailia Governorate in the Canal region, and New Valley and Beni Suef Governorates in Upper Egypt (southern Egypt). Interestingly, the analysis of the deduced amino acids demonstrated scattered amino acid mutations of the four mentioned strains compared with the published sequences. The number of observed amino acid substitutions was restricted. The recurrent E substitution at position 30 and Y substitution at position 13, detected among the Egyptian isolates, may represent a recent microevolutionary divergence. Furthermore, E substitution possibly alters epitope exposure and contributes to immune escape mechanisms, facilitating immune evasion. The Y substitution potentially alters the capsid topology. Particularly, the shared cysteine substitution among all four isolates may alter antigenicity and constitute a distinct molecular feature of Egyptian AEV strains, may suggest the emergence of a region-specific lineage. Similar inspections were reported by Temeeyasen et al. (2024), who stated that the majority of distinctive amino acids were found in the VP1 gene, indicating that the AEV can mutate through positive pressure. These genetic mutations were phenotypically expressed by altering the vaccine strain’s virulence and pathogenicity (Mattenberger et al., 2021). The recombination analysis was conducted on a partial VP1 gene fragment (288 bp), which may limit the statistical power to detect recombination events; thus, whole-genome of longer genomic fragment are recommended for more robust recombination detection. Live attenuated AEV vaccines are likely to remain effective; however, strict optimization of breeder flock vaccination is essential to prevent vertical transmission. Continuous molecular surveillance, particularly VP1-based monitoring, is recommended to detect changes that may affect antigenicity and long-term vaccine efficacy. ConclusionOur promising results provide the first preliminary molecular and clinical insights into AEV strains detected in broiler flocks from selected poultry-producing governorates in Egypt based on partial VP1 gene analysis, filling a critical gap in national molecular surveillance data that has been lacking in recent years. The current data demonstrate that AEV is a significant etiological agent of neurological disorders among young chicks, resulting in serious economic losses to the Egyptian poultry industry. Currently, the detection of AEV strains in four major poultry-producing governorates indicates ongoing virus circulation and geographic dissemination at the national level, underscoring the need for systematic surveillance programs. Moreover, the phylogenetic analysis of the AEV-VP1 gene of the Egyptian isolates (NewValley-AEV-1-2024, BeniSuef-AEV-2-2024, Ismailia-AEV-3-2024, and ElSharquia-AEV-4-2024, cluster II) displayed a genetic similarity to Chinese, Iranian, American, and British isolates with a nucleotide (94%–98%) and amino acid (96%–99%) identity percentages, respectively. Surprisingly, this study can serve as a guide for improving vaccination strategies as an efficient control and prevention method in the poultry industry. Although these data provide baseline molecular evidence essential for national AEV surveillance and risk assessment, the vaccination strategies remain speculative because this study did not include antigenic characterization, cross-protection assessments, or vaccine challenge experiments. Importantly, whole-genome sequencing, which has a higher discriminatory power, of recent AEV circulating isolates is strongly recommended to provide broad information on their pathogenicity, antigenicity, molecular epidemiology, and genetic relatedness among AEV and vaccinal strains. AcknowledgmentsThe authors would like to express their deep appreciation to the Virology and Pathology departments of the Faculty of Veterinary Medicine, New Valley University, Egypt, and the Gene Analysis Unit, Reference Laboratory for quality control on poultry, Animal Health Institute, Egypt, for their collaboration and support during all procedures of this experimental research. FundingThis study received no external funding. Authors’ contributionsA.F.A. and E.A-M.S.: performed the collection and processing of the samples and performed the laboratory pathological experiments. I.E.: reviewed and corrected the paper. E.A-M.S. and A.M.Z.: performed the computational analysis of the data. E.A-M.S. and I.E.: performed the laboratory virological experiments, wrote the paper, and reviewed the manuscript. Conflict of interestThe authors declare that they have no known competing financial and non-financial interests or even personal relationships that could have appeared to influence the work reported in this paper. Data availabilityData available upon request. ReferencesAbd El Hafez, M.S., Shosha, E.A.E.M. and Ibrahim, S.M. 2021. Isolation and molecular detection of pigeon pox virus in Assiut and New Valley governorates. J. Virol. Methods. 293, 114142. Abodalal, S.E.S.A., Hafez, M.S.H.A., Abd El-munem Shosha, E., Warda, F.F. and Hagag, N.M. 2021. Isolation and Molecular Characterization of Rabbit Haemorrhagic Disease Virus Strains Circulating in Rabbit Population Using Sequencing and Phylogenetic Analysis. J. World. Poultry. Res. 11(3), 302–311. Ahmed, D.A.A., Hassan, A.K., Ali, D.A.A., Shosha, E.A.E., Eldaghayes, I.M. and Khair, S.M.A.A. 2025. Molecular detection and sequencing of the NetB toxin Gene of clostridium perfringens and evaluation of its pathogenicity in broiler chicken. Poult. Sci. 105(2), 106289. Al-Hammadi, M.A. and Al-Rasheed, M. 2024. Occurrence of avian encephalomyelitis virus in naturally infected chicks in Saudi Arabia’s Eastern Province. Open Vet. J. 14, 335–340. Ali, M., Shaon, M., Moula, M., Bary, M., Sabuj, A., Khaled, S., Bhuiyan, Z. and Giasuddin, M. 2021. First report on the seroprevalence of avian encephalomyelitis virus antibody in Sonali (cross-bred) chickens in Bogura, Bangladesh. J. Adv. Vet. Anim. Res. 8, 78–83. Amal, A.M., Zaghloul, M., El-Munem, S.E.A., Darwish, D.M., Heba A Khafagy., Farouk, E.M. and Ibrahim, A.I. 2019. Cross neutralization between vaccinal strain of commercial bovine ephemeral fever virus vaccines and Egyptian isolate 2018 with serum samples of vaccinated cattle. Inter. J. Vet. Sci. 8(4), 329–334. Badr, H., Abdelmenamm Shosha, E., Roshdy, H., Abd El-halem Mohammed, A., Saad, N., Mostafa Aboelenin, S., Mohamed Soliman, M., El-Tahan, A.M., El-Saadony, M.T. and Yehia, N. 2022. Investigation of many bacterial and viral infections circulating in pigeons showing nervous symptoms. Saudi J. Biol. Sci. 29(4), 2911–2920. Biswas, S.K., Jana, C., Chand, K., Rehman, W. and Mondal, B. 2011. Detection of fowl poxvirus integrated with reticuloendotheliosis virus sequences from an outbreak in backyard chickens in India. Veterinaria. Italiana. 47, 147–153. Braune, M.O. and Gentry, R.F. 1971. Avian encephalomyelitis virus. I. Pathogenesis in chicken embryos. Avian Dis. 15, 638–647. Diab, M.S., Abd El Hafez, M.S., Ashry, M.A. and Elfeil, W.K. 2019. Occurrence of avian influenza H5N1 among chicken, duck, farms and human in Egypt. Am. J. Anim. Vet. Sci. 14, 26–32. El-khabaz, K., Shosha, E. and Abdel-ra’ouf, A. 2021. New epizootic of lumpy skin disease in Assiut-Egypt: molecular identification and characterization. Adv. Anim. Vet. Sci. 9(3), 446–452. Fotouh, A., Shosha, E.A.E.M., Zanaty, A.M. and Darwesh, M.M. 2024. Immunopathological investigation and genetic evolution of Avian leukosis virus Subgroup-J associated with myelocytomatosis in broiler flocks in Egypt. Virol. J. 21(1), 83. Gethoffer, F., Curland, N., Voigt, U., Woelfing, B., Ludwig, T., Heffels-Redmann, U., Hafez, H.M., Lierz, M. and Siebert, U. 2021. Seroprevalence of specific antibodies against avian pathogens in free-ranging ring-necked pheasants (Phasianus colchicus) in Northwestern Germany. PLos One 16, e255434. Ghalyanchilangeroudi, A., Madani, S.A., Najafi, H., Ziafati Kafi, Z., Sadri, N., Sarmadi, S., Eghbali, O.O., Jamiri, F., Bakhshi, A.A. and Hosseini, H. 2024. The full genome characterization of avian encephalomyelitis virus in Iran: a vertical transmission case. Virus Genes 60(4), 393–401. Goto, Y., Yaegashi, G., Kumagai, Y., Ogasawara, F., Goto, M. and Mase, M. 2019. Detection of avian encephalomyelitis virus in chickens in Japan using RT-PCR. J. Vet. Med. Sci. 81, 103–106. Haryanto, A., Ermawati, R., Wati, V., Irianingsih, S.H. and Wijayanti, N. 2016. Analysis of viral protein-2 encoding gene of avian encephalomyelitis virus from field specimens in Central Java region, Indonesia. Vet. World. 9, 25–31. Hauck, R., Sentíes-Cué, C.G., Wang, Y., Kern, C., Shivaprasad, H.L., Zhou, H. and Gallardo, R.A. 2017. Evolution of avian encephalomyelitis virus during embryo-adaptation. Vet. Microbiol. 204, 1–7. Hoekstra, J. 1964. Experiments with avian encephalomyelitis. Br. Vet. J. 120, 322–335. Kant, A., Balk, F., Born, L., Van Roozelaar, D., Heijmans, J., Gielkens, A. and Ter Huurne, A. 2003. Classification of Dutch and German avian reoviruses by sequencing the sigma C protein. Vet. Res. 34, 203–212. Kumar, S., Stecher, G. and Tamura, K. 2016. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. J. Mol. Genet. Evol. 33, 1870–1874. Lin, W., Lu, P., Li, A., Wu, Y., Li, H., Chen, F., Ma, J. and Xie, Q. 2018. Assessing the efficacy of a live vaccine against avian encephalomyelitis virus. Arch. Virol. 163, 2395–2404. Liu, Q., Yang, Z., Hao, H., Cheng, S., Fan, W., Du, E., Xiao, S., Wang, X. and Zhang, S. 2014. Development of a SYBR Green real-time RT-PCR assay for the detection of avian encephalomyelitis virus. J. Virol. Methods 206, 46–50. Martin, D.P. 2009. Recombination detection and analysis using RDP3. Methods. Mol. Biol. 537, 185–205. Martin, D.P., Posada, D., Crandall, K.A. and Williamson, C. 2005. A modified bootscan algorithm for automated identification of recombinant sequences and recombination breakpoints. AIDS Res. Hum. Retroviruses 21, 98–102; doi:10.1016/j.asdrhr.2005.09.010 Mattenberger, F., Latorre, V., Tirosh, O., Stern, A. and Geller, R. 2021. Globally defining the effects of mutations in the capsid of a picornavirus. Elife 10, e6425. Meir, R., Maharat, O., Farnushi, Y. and Simanov, L. 2010. Development of a real-time TaqMan® RT-PCR assay for the detection of infectious bronchitis virus in chickens, and comparison of RT-PCR and virus isolation. J. Virol. Methods. 163, 190–194. Mosad, S.M., Rasheed, N., Ali, H.S., El-Khabaz, K.A.S., Shosha, E.A.M. and El-Diasty, M. 2021. Incidence of lumpy skin disease virus with its characterization in vaccinated pregnant Holstein cows in Dakahlia governorate, Egypt. Ger. J. Vet. Res. 1(4), 23–33. Pan, J., Narayanan, B., Shah, S., Yoder, J.D., Cifuente, J.O., Hafenstein, S. and Bergelson, J.M. 2011. Single amino acid changes in the virus capsid permit coxsackievirus b3 to bind decay-accelerating factor. J. Virol. 85, 7436–7443. Pohjola, L., Tammiranta, N., Ek-Kommonen, C., Soveri, T., Hänninen, M.L., Fredriksson Ahomaa, M. and Huovilainen, A. 2017. A survey for selected avian viral pathogens in backyard chicken farms in Finland. Avian Pathol. 46, 166–172. Quaglia, G., Mescolini, G., Catelli, E., Berto, G., Muccioli, F. and Lupini, C. 2021. Genetic heterogeneity among chicken infectious anemia viruses detected in Italian fowl. Animals 11(4), 944. Ramzy, N.M., Ayatollah I Ibrahim., Mohamed S Abd., Eman, E.H. and Shosha. 2024. Isolation and genetic diversity of fowlpox virus circulating in chicken flocks in Egypt. J. Adv. Vet. Res. 14(5), 799–806. Raue, R. and Hess, M. 1998. Hexon based PCR’s combined with restriction enzyme analysis for rapid detection and differentiation of fowl adenoviruses and egg drop syndrome virus. J. Virol. Methods 73, 211–217. Sarma, G., Kersting, B.A. and Spina, G. 2019. Field safety and efficacy of a unique live virus vaccine for controlling avian encephalomyelitis and fowlpox in poultry. Vet. World 12, 1291–1298. Sentíes-Cué, C.G., Gallardo, R.A., Reimers, N., Bickford, A.A., Charlton, B.R. and Shivaprasad, H.L. 2016. Avian encephalomyelitis in Layer Pullets Associated with Vaccination. Avian Dis. 60, 511–515. Shafren, D.R. and Tannock, G.A. 1991. Pathogenesis of avian encephalomyelitis viruses. J. Gen. Virol. 72(11), 2713–2719. Shosha, E., Abdelnaser, S. and Zanaty, A.M. 2025a. A Full-length S1 gene sequencing of a novel emerged GI-19 and GI-23 lineages of Infectious bronchitis virus currently circulating in chicken flocks in Upper Egypt reveals marked genetic diversity and recombination events. Virol. J. 22, 135. Shosha, E.A.E., Abdelnaser, S., Zanaty, A.M., Elzanaty, A.E., Selim, K. and Eldaghayes, I. 2025b. Successive Efficacy Evaluation of Various Commercial Live-Attenuated Avian coronavirus Vaccination Schedules Against a Local GI-23.3 Challenge in SPF Broilers. Vaccines 13, 1132. Shosha, E.A.E., Mohamd, M.K., Shehata, M.A.E., Mohamed, M.H., Eldaghayes, I. and Abdelhafez, M.S. 2025. Genomic and evolutionary characterization of newly emerged highly pathogenic avian influenza H5N1 clade 2.3.4.4b viruses circulating in broiler flocks in Upper Egypt (2023–2025). Vet. World 18(12), 3745–3760; doi:10.1016/j.vw.2023 Shosha, E.A.E.M., Zanaty, A.M., Darwesh, M.M. and Fotouh, A. 2024. Molecular characterization and immunopathological investigation of Avian reticuloendotheliosis virus in breeder flocks in Egypt. Virol. J. 22(1), 259. Swayne, D. E., Glisson, J. R., McDougald, L. R., Nolan, L. K., Suarez, D. L., & Nair, V. L. (Eds.). Diseases of Poultry, 13th ed. Wiley-Blackwell; 2013. 1408 pp. Tamura K Stecher., Peterson, G., Filipski, D. and Kumar, S. 2013. Molecular biology and evolution. Mol. Biol. Evol. 30, 2725–2729. Taunde, P., Timbe, P., Lucas, A.F., Tchamo, C., Chilundo, A., Dos Anjos, F., Costa, R. and Bila, C.G. 2017. Serological evidence of avian encephalomyelitis virus and Pasteurella multocida infections in free-range indigenous chickens in Southern Mozambique: a case report. Trop. Anim. Health Prod. 49, 1047–1050. Temeeyasen, G., Sharafeldin, T., Gharaibeh, S., Sobhy, N.M., Porter, R.E. and Mor, S.K. 2024. A New Variant of Avian Encephalomyelitis Virus Associated with Neurologic Signs in Turkey Poults. Pathogens 13(758), 758. Torre, D.D.L., Nuñez, L., Parra, S., Astolfi-Ferreira, C. and Ferreira, A. 2018. Detection by rt-pcr and molecular characterization of tremovirus A obtained from clinical cases of avian encephalomyelitis (AE) outbreaks in brazil. Braz. J. Poultry Sci. 20(3), 527–536. Varte, L., Deka, D., Gupta, K. and Singh, A. 2023. Comparative sequence analysis of Meq oncogene of Marek’s disease virus field isolates detected in Marek’s disease affected birds from vaccinated poultry flocks. Indian J. Vet. Pathol. 47, 211–218. Wang, G., He, Y., Yan, X., Sun, Y., Yi, L., Tu, C. and He, B. 2023. Virome profiling of chickens with hepatomegaly rupture syndrome reveals coinfection of multiple viruses. Viruses 15, 1249. Wei, L., Chee, L.L., Wei, T., Kwang, J., Zhou, J., Wang, J., Shi, L. and Liu, J. 2008. The VP1 protein of avian encephalomyelitis virus is a major host-protective immunogen that serves as diagnostic potential. J. Virol. Methods 149, 56–62. Wise, M.G., Suarez, D.L., Seal, B.S., Pedersen, J.C., Senne, D.A., King, D.J., Kapczynski, D.R. and Spackman, E. 2004. Development of a real-time reverse-transcription PCR for detection of Newcastle disease virus RNA in clinical samples. J. Clin. Microbiol. 42, 329–338; doi:10.1016/j.jcm.2004.01.016 Xu, Z., Fan, P., Zhang, C., Guo, M., Bo, Z., Wu, Y. and Zhang, X. 2025. Identification of virulence-related amino acid mutations of avian encephalomyelitis virus associated with vaccination methods. Front. Vet. Sci. 12, 1548515. Xue, Q., Guo, H., Feng, Z. and Wang, J. 2016. Establishment of a real-time RT-PCR assay for avian encephalomyelitis virus. Med. Weter. 72, 418–422. Zhang, F., Luo, Y., Wei, Q., Xiong, L., Xie, Q., Tan, J., Wu, C., Li, N. and Kang, Z. 2024. Research Note: pathogenetic characteristics of avian encephalomyelitis virus in Guangdong and Jiangxi Provinces, China. Poult. Sci. 103(2), 103264. Zhang, G., Li, S., Shen, Z. and Wang, F. 2023. Progress in research on the molecular biological detection techniques of avian encephalomyelitis. Res. Vet. Sci. 159, 232–236. | ||
| How to Cite this Article |
| Pubmed Style Shosha EA, Zanaty AM, Abdallah AF, Eldaghayes I. Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Vet. J.. 2026; 16(2): 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 Web Style Shosha EA, Zanaty AM, Abdallah AF, Eldaghayes I. Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. https://www.openveterinaryjournal.com/?mno=298489 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.50 AMA (American Medical Association) Style Shosha EA, Zanaty AM, Abdallah AF, Eldaghayes I. Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Vet. J.. 2026; 16(2): 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 Vancouver/ICMJE Style Shosha EA, Zanaty AM, Abdallah AF, Eldaghayes I. Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 Harvard Style Shosha, E. A., Zanaty, . A. M., Abdallah, . A. F. & Eldaghayes, . I. (2026) Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Vet. J., 16 (2), 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 Turabian Style Shosha, Eman Abdelmenamm, Ali Mahmoud Zanaty, Ahmed Fotouh Abdallah, and Ibrahim Eldaghayes. 2026. Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Veterinary Journal, 16 (2), 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 Chicago Style Shosha, Eman Abdelmenamm, Ali Mahmoud Zanaty, Ahmed Fotouh Abdallah, and Ibrahim Eldaghayes. "Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt." Open Veterinary Journal 16 (2026), 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 MLA (The Modern Language Association) Style Shosha, Eman Abdelmenamm, Ali Mahmoud Zanaty, Ahmed Fotouh Abdallah, and Ibrahim Eldaghayes. "Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt." Open Veterinary Journal 16.2 (2026), 1338-1350. Print. doi:10.5455/OVJ.2026.v16.i2.50 APA (American Psychological Association) Style Shosha, E. A., Zanaty, . A. M., Abdallah, . A. F. & Eldaghayes, . I. (2026) Histopathological diagnosis and genetic diversity of avian encephalomyelitis virus circulating in broiler farms in Egypt. Open Veterinary Journal, 16 (2), 1338-1350. doi:10.5455/OVJ.2026.v16.i2.50 |