| Research Article | ||
Open Vet. J.. 2026; 16(2): 1315-1328 Open Veterinary Journal, (2026), Vol. 16(2): 1315-1328 Research Article Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheepMarwa Saleem Hajeel and Monyer Abdulamier Abd Alfatlawi*Department of Veterinary Microbiology, College of Veterinary Medicine, University of Al-Qadisiyah, Al-Diwaniyah, Iraq *Corresponding Author: Monyer Abdulamier Abd Alfatlawi. Department of Veterinary Microbiology, College of Veterinary Medicine, University of Al-Qadisiyah, Al-Diwaniyah, Iraq. Email: monyerr.abd [at] qu.edu.iq Submitted: 17/11/2025 Revised: 14/01/2026 Accepted: 18/01/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
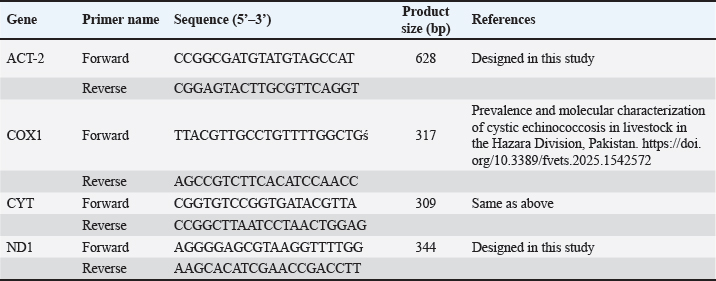
ABSTRACTBackground: Hydatid disease remains a major concern in sheep farming, particularly in endemic areas such as Iraq. Aim: This study aimed to explore the molecular characteristics of Echinococcus granulosus isolated from sheep using multilocus genetic analysis. Methods: Fertile cysts were collected from sheep slaughtered in Diwaniyah and processed for DNA extraction. Four genetic loci—ACT-2, COX1, CYT, and ND1—were amplified by polymerase chain reaction (PCR) and sequenced to evaluate the diversity of the parasite. Results: Each gene yielded fragments of expected lengths, and no contamination was observed in the negative controls. Sequences from sheep isolates were submitted to GenBank, with accession numbers PV959524–PV959533 (ACT-2), PV959270–PV959279 (COX1), PV975191–PV975200 (CYT), and PV975171–PV975180 (ND1). BLAST comparisons revealed identities ranging from 95.6% to 100%, with close matches to isolates from China, Brazil, Estonia, and Turkey. Phylogenetic trees showed clear clustering of local sheep strains with global references, particularly within the G1 lineage. Multiple sequence alignments confirmed a high degree of similarity with minor genetic variation. Conclusion: These results demonstrate the genetic stability of E. granulosus in sheep in southern Iraq and support its inclusion in regional parasite mapping efforts. The data also emphasize the importance of sheep as intermediate hosts contributing to the persistence and spread of this zoonotic parasite. The genetic analysis of E. granulosus from sheep showed close relationships with international strains, reflecting regional transmission and limited sequence divergence. Molecular typing remains essential for the epidemiological control and future vaccine development. Keywords: Alignment, Diagnostic, Liver, Sequencing, Zoonosis. IntroductionCystic echinococcosis (CE) is a disease caused by the larval form of E. granulosus sensu lato (s.l.) (Woolsey and Miller, 2021). This parasite belongs to the tapeworm group. It spreads widely around the world. CE affects both animals and humans. The disease is particularly common in areas where sheep are raised. Sheep often act as intermediate hosts. They get infected when they eat parasite eggs. These eggs hatch and form cysts, mainly in the liver or lungs of sheep. Dogs are the final hosts. They eat infected sheep organs and then pass the eggs in their feces (Manterola et al., 2022). This keeps the infection cycle going. The disease causes significant problems for farmers. Infected animals must be destroyed. This leads to losses in meat production. Farmers also face extra costs for control measures (García-Méndez et al., 2022). In many rural places, such as the Middle East, CE is still common. However, it is often not reported well. Studies using molecular tools have shown that E. granulosus s.l. has different genotypes. The most common species found in sheep is E. granulosus sensu stricto (G1–G3) (Ohiolei et al., 2022). Other types, such as G6 and G7, have also been found in sheep, but they are rare (Hua et al., 2022). Knowing the genotype is essential. It helps in understanding how a disease spreads in a region. Different types may behave differently. Some may cause more harm, affect different animals, or react differently to treatment (Wen et al., 2019; Khademvatan et al., 2019). Research from Europe, Asia, and Africa has shown that sheep strains are genetically similar to those found in other farm animals (Ali et al., 2020; Shams et al., 2022; Casulli et al., 2022a,b). This shows that the parasite easily moves between animals. Nevertheless, differences in parasite types have been found even in the same animal species from different places (Tamarozzi et al., 2020a,b; Borhani et al., 2024). Therefore, it is important to study the parasite locally. Iraq is a country in which such studies are still limited. A good way to study the parasite is through MLG. This method examines more than one gene. It helps to identify differences even within the same genotype. Scientists often use mitochondrial genes such as COX1 and ND1. They also use nuclear genes such as ACT-2 (Aregawi et al., 2024; Gharbi and Giraudoux, 2024). These genes help researchers understand how parasite strains are related. This is useful, especially in areas where the parasite infects many types of animals (Tamarozzi et al., 2020a,b). In Iraq, especially in the south, there are insufficient genetic data about E. granulosus in sheep. This makes it difficult to determine how the parasite is changing or spreading. Conducting more genetic studies on sheep can help. This will improve our understanding of the disease and how it moves between animals (Anvari et al., 2021a,b; Karshima et al., 2022). Sheep are essential in maintaining the parasite cycle. If we do not know which strains are present in sheep, we cannot effectively control the disease (Samei and Khedri, 2025). Good data are needed to make vaccines or deworming programs. Because sheep often live with other animals, they may pass the parasite to others. Studying their role carefully is important. Molecular tools can help us understand how parasites spread between animals and regions (Hua et al., 2022). This study was conducted to provide such information. The genetic types of E. granulosus found in sheep from southern Iraq were examined using PCR and gene sequencing. Materials and MethodsSample collectionFertile hydatid cysts were collected from the liver and lungs of infected sheep during routine postmortem inspection at a slaughter facility. Cysts were microscopically examined to confirm fertility and viability by identifying the presence of protoscoleces and live protoscoleces, respectively (Mokhtaria and Ammar, 2019). Frozen hydatid cysts can replace incineration and sterilize cysts. The germinal layer of each cyst was carefully separated using sterile tools and transferred into labeled Eppendorf tubes. The samples were transported in a cooled container and delivered to the laboratory within hours. Upon arrival, all samples were stored at 20°C until molecular analysis. Care was taken to avoid contamination, and each specimen was tracked with specific codes indicating the host species, tissue source, and collection date. This method ensured proper traceability and integrity throughout the study. Genomic DNA extractionDNA was extracted from the germinal layers using a silica membrane-based column protocol. Each sample was treated with 200 µl of lysis buffer and 20 µl of proteinase K solution and incubated at 56 °C for 1 hour. Enzymatic digestion allowed the release of genomic material from the parasite tissue. After incubation, the lysate was briefly centrifuged, and the clear supernatant was transferred to a fresh tube. Binding buffer and absolute ethanol were added to allow DNA to bind to the spin column membrane. After successive washing with two wash buffers and a final dry spin, DNA was eluted using 50 µl of elution buffer. The extracted DNA was stored at 20°C for downstream PCR. All steps were performed under sterile conditions, and the quality of the process was ensured by checking the clarity and absence of pellet debris in each spin. DNA quantificationTo confirm the quantity and quality of extracted DNA, measurements were performed using a fluorescence-based assay. A working solution of the DNA-binding dye was prepared by diluting the stock solution in 1X TE buffer. The blank, standard, and sample tubes were prepared according to the manufacturer’s instructions. Each reaction used 2 µl of DNA, which was mixed with 200 µl of dye solution, and the samples were read using a fluorometer. The results provided accurate concentrations for each DNA extract. This step was crucial to confirm whether the quantity of DNA was sufficient for PCR amplification and to ensure consistent loading across all gene targets. Primer selection and PCR amplificationFour different primer pairs were used to target the ACT-2, COX1, CYT, and ND1 regions. Each PCR reaction was 20 µl in volume and included master mix, forward and reverse primers (0.5 pmol each), nuclease-free water, and 2 µl of DNA template. To avoid contamination, reaction mixes were set up under sterile conditions (Table 1). Thermal cycling was performed under standardized conditions: initial denaturation at 95°C for 5 minutes; 39 cycles of denaturation at 95 °C (30 s), annealing at 60°C (35 s), and extension at 72 °C (35 s); followed by a final extension step at 72 °C for 5 minutes. Each PCR run included a no-template control to monitor potential contamination. The amplified products were stored at 4°C before analysis. Gel electrophoresisThe PCR products were resolved using 1.5% agarose gel in TBE buffer. The gel was stained with ethidium bromide and cast in wells in trays fitted with combs. After solidification, samples were loaded alongside a DNA ladder to estimate fragment size. Electrophoresis was performed at 100 V for 1 hour. Visualization was performed using UV illumination. Bands corresponding to expected sizes—628 bp (ACT-2), 317 bp (COX1), 309 bp (CYT), and 344 bp (ND1)—were confirmed. The clarity and position of the bands ensured proper amplification, and the absence of bands in negative controls validated the specificity of the reactions. DNA sequencing and phylogenetic analysisTen positive PCR products for each gene were selected for sequencing. Samples were purified and sent to Macrogen (Korea) using a commercial sequencing service. The returned chromatograms were manually trimmed to remove poor-quality regions and saved in the FASTA format. Clean sequences were submitted to GenBank and compared with international sequences using the BLASTn tool. Phylogenetic trees were constructed using the maximum likelihood method with MEGA11 software. To assess variations, multiple sequence alignments were performed using ClustalW. The analysis confirmed strong similarity between the local sheep isolates and global genotypes. Ethical approvalNot needed for this study. ResultsACT-2 genePCR amplification of the ACT-2 gene from sheep isolates resulted in clear bands at 628 bp in all 10 samples. The reactions were clean, and the expected fragment size was confirmed by comparison with the molecular weight marker (Fig. 1). Table 1. The primers used in the current study.
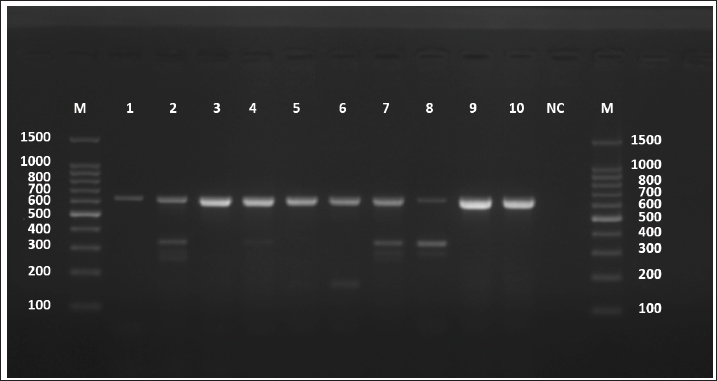
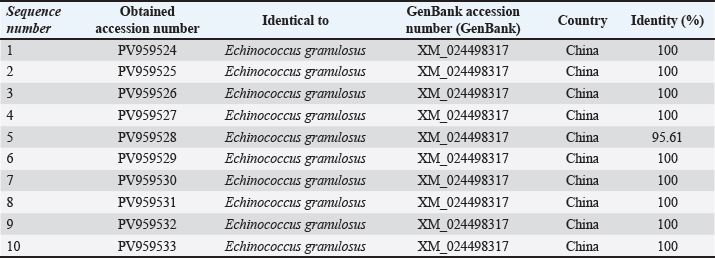
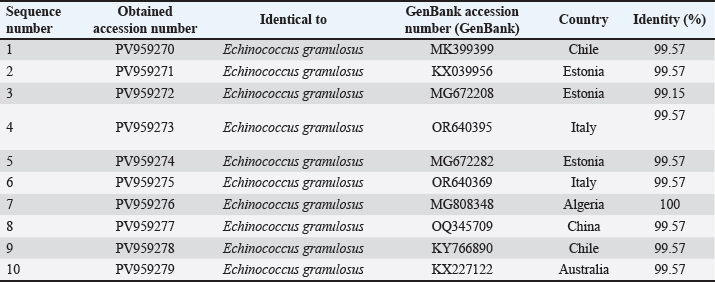
Fig. 1. Agarose gel electrophoresis image (1.5%) shows the amplicons of the ACT2 gene (size=628 bp) in sheep. NC is the negative control in which H2O was added instead of the template DNA. M is a molecular marker (100–1,500 bp) obtained from GeneDirex (South Korea). The negative control lane showed no amplification, confirming that the reactions were contamination-free. All positive samples exhibited distinct bands with no background smearing or nonspecific products, indicating high-quality DNA and primer specificity. The ten sheep-derived ACT-2 amplicons were sequenced following successful PCR. GenBank accession numbers PV959524 to PV959533 were assigned to these samples. Table 2 lists the BLASTn results. Nine of the 10 sequences showed 100% identity with the Chinese reference sequence XM_024498317. Only one sample (PV959528) showed a slightly lower identity at 95.61% (Fig. 2). The alignment results are shown in Figure 3. COX1 geneAll sheep samples amplified successfully at the COX1 locus, yielding a 317-bp band. The electrophoresis results are documented in Figure 4. The absence of amplification in the negative control confirmed that the reaction was not contaminated. Bands from sheep samples were sharp and aligned with the expected size, confirming the effectiveness of the primer. Table 2. The NCBI-BLAST homology sequence identity (%) between local E. granulosus targeting the ACT2 gene in sheep.
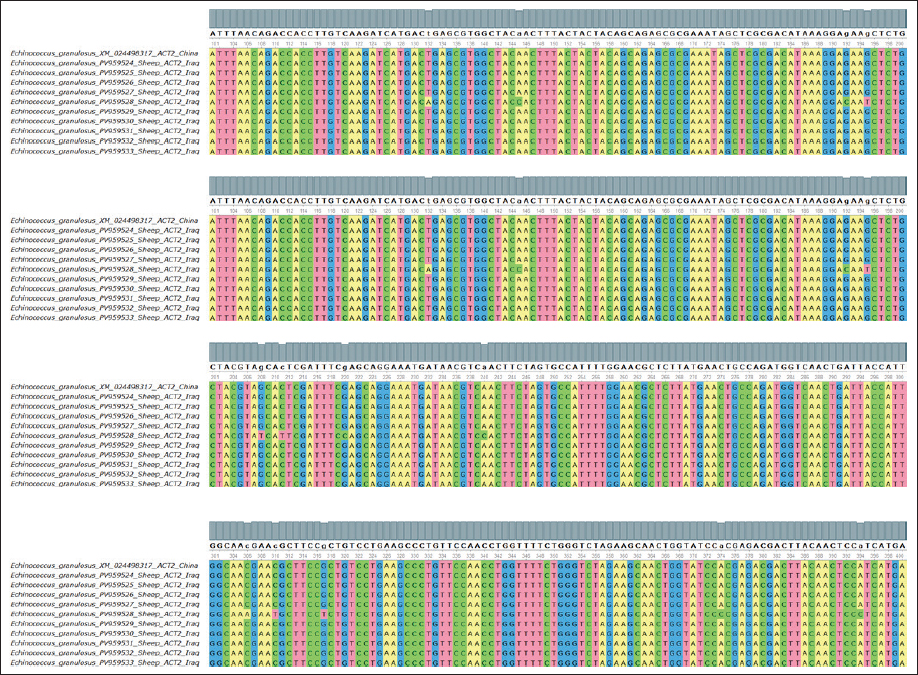
Fig. 2. Evolutionary analysis of E. granulosus using the maximum likelihood method targeting the ACT2 gene in sheep confirmed close clustering with Chinese isolates. The alignment results are shown in Figure 3.
Fig. 3. Multiple sequence alignment of the partial ACT2 gene in E. granulosus shows similarities and differences between the identified sequences in sheep, supporting the sequence identity findings and highlighting minimal variation among the sheep isolates.
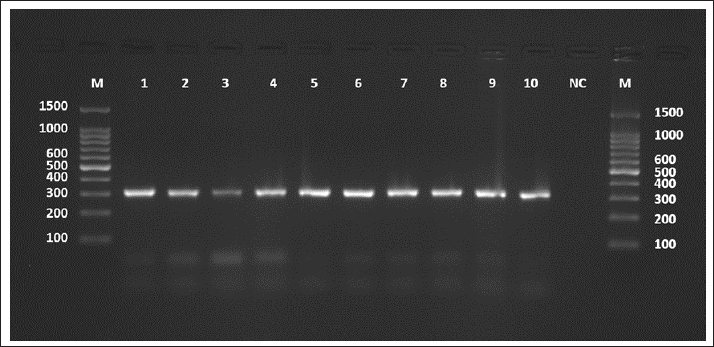
Fig. 4. Agarose gel electrophoresis image (1.5%) shows the amplicons of the COX1 gene (size=317 bp) in sheep. NC is the negative control in which H2O was added instead of the template DNA. M is a molecular marker (100–1,500 bp) obtained from GeneDirex (South Korea). Table 3. The NCBI-BLAST homology sequence identity (%) between local E. granulosus targeting the COX1 gene in sheep.
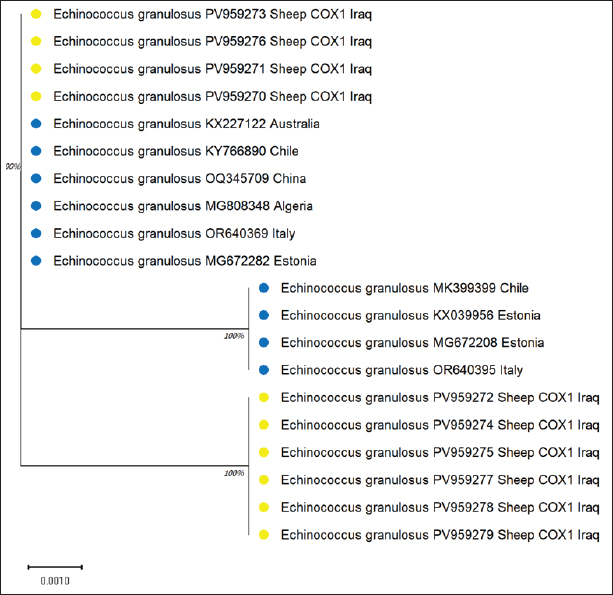
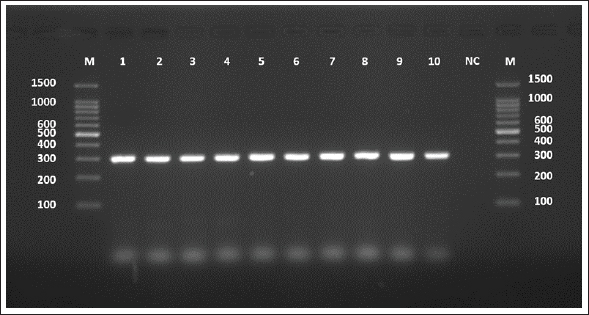
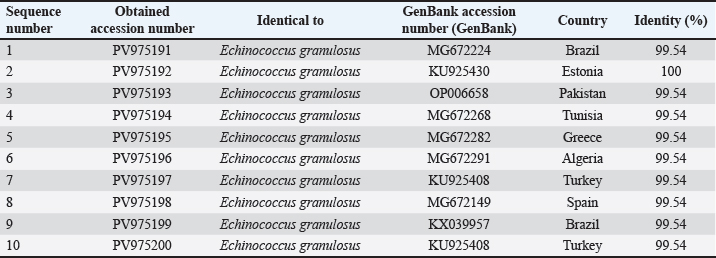
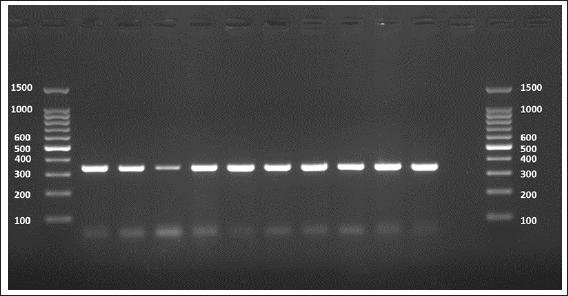
The sequences were submitted under GenBank accession numbers PV959270–PV959279. The identity scores are shown in Table 3. Most of the sequences had 99.57% identity with multiple isolates from Chile (MK399399), Estonia (KX039956), and Australia (KX227122), while one sample (PV959276) showed 100% identity with an Algerian isolate (MG808348). Tree analysis is shown in Figure 5. Alignment results are shown in Figure 6 Abd-Alhassen et al., (2021), Rojas-Pincheira et al., (2022), Casulli et al., (2022b) and Al-Taee et al., (2024ab). CYT geneThe CYT gene was successfully amplified in all 10 sheep isolates, producing distinct PCR bands at 309 bp (Fig. 7). Negative controls were clear, and each sheep sample showed single, specific bands. This consistency confirms that the DNA templates and primers were suitable for targeting cytochrome b in sheep E. granulosus. Sequencing results were deposited under GenBank accession numbers PV975191–PV975200. The identity data are detailed in Table 4.
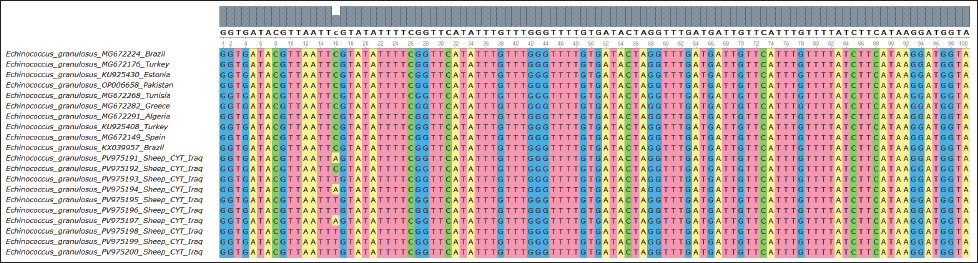
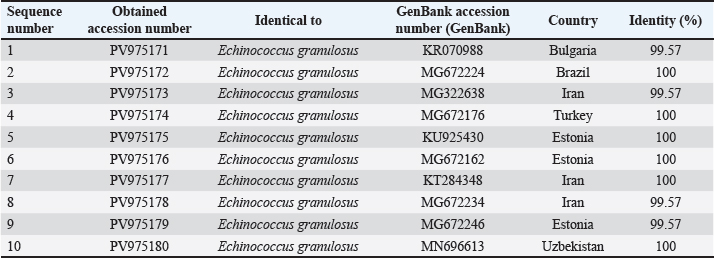
Fig. 5. Evolutionary analysis of E. granulosus using the maximum likelihood method targeting the COX gene in sheep placed within the same clade as these international strains. Alignment results are shown in Figure 6. All 10 samples showed high identity (99.54%–100%) with isolates from Brazil (MG672224), Estonia (KU925430), and Pakistan (OP006658). The phylogenetic tree is shown in Figure 8. ND1 geneThe ND1 gene was amplified in all sheep isolates, producing 344 bp fragments. The agarose gel output is shown in Figures 9 and 10. Sequencing data were submitted under PV975171 to PV975180. The results are shown in Table 5. Identity ranged from 99.57% to 100% compared with strains from Bulgaria (KR070988), Iran (MG322638), Turkey (MG672176), and Uzbekistan (MN696613). The phylogenetic tree is shown in Figure 11. Alignment results are shown in Figure 12. DiscussionThe molecular characterization of E. granulosus isolated from sheep in Diwaniyah revealed strong sequence identity with previously reported global isolates. Our ACT-2 sequences showed 100% similarity with a Chinese reference strain in nine out of ten samples, consistent with the genetic homogeneity observed in many endemic areas (García-Méndez et al., 2022). These findings support the view of García-Méndez et al. (2022) who indicated that the proteomic and genetic profiles of E. granulosus tend to be highly conserved in dominant hosts, such as sheep. Our COX1 gene data followed a similar pattern, closely aligning with isolates from Europe and South America. This resembles the camel-based genotyping trends documented by Anvari et al. (2021a,b) where genetic uniformity across continents was also evident. Despite the geographical distance, sheep strains from Iraq share a high genetic affinity with those from Brazil, Estonia, and China, demonstrating the global stability of the G1 genotype.
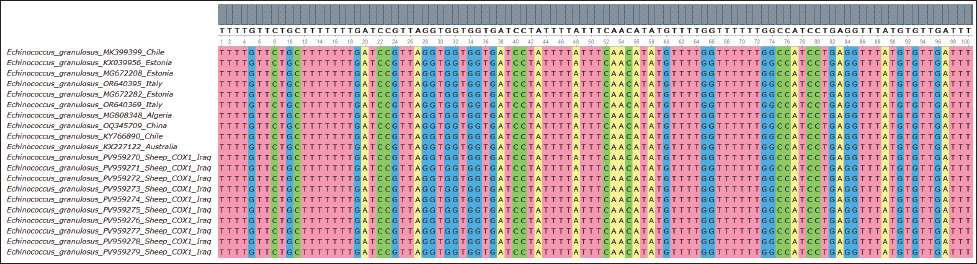
Fig. 6. Multiple sequence alignment of the partial COX1 gene in E. granulosus shows the similarity and differences between the identified sequences in sheep, further confirming the strong genetic consistency among the isolates.
Fig. 7. Agarose gel electrophoresis image (1.5%) shows the amplicons of the CYT gene (size=309 bp) in sheep. NC is the negative control in which H2O was added instead of the template DNA. M is a molecular marker (100–1,500 bp) obtained from GeneDirex (South Korea). Table 4. The NCBI-BLAST homology sequence identity (%) between local E. granulosus targeting the CYT gene in sheep.
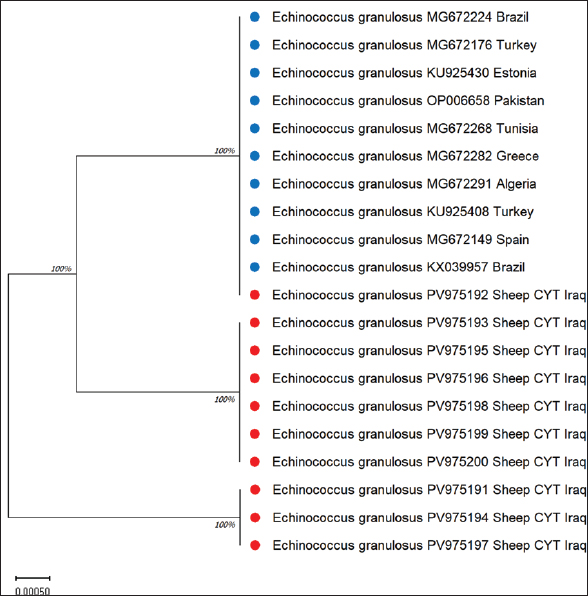
Fig. 8. Evolutionary analysis of E. granulosus using the maximum likelihood method targeting the CYT gene in sheep revealed that the sheep sequences were tightly grouped with reference strains from different continents. The alignment is visualized in Figure 9.
Fig. 9. Multiple sequence alignment of the partial CYT gene in E. granulosus reveals similarities and differences between the identified sequences in sheep Confirmed minimal variation, suggesting a high degree of conservation within this gene in the local sheep population.
Fig. 10. Agarose gel electrophoresis image (1.5%) shows the amplicons of the COX1 gene (size=344 bp) in sheep. NC is the negative control in which H2O was added instead of the template DNA. M is a molecular marker (100–1,500 bp) obtained from GeneDirex (South Korea). Each sheep sample produced a single, clean band, and no false positives were confirmed by the controls. Table 5. The NCBI-BLAST homology sequence identity (%) between local E. granulosus targeting the ND1 gene in sheep.
Fig. 11. Echinococcus granulosus evolutionary analysis using the maximum likelihood method targeting the ND1 gene in sheep showed clustering within a conserved lineage, confirming close global relationships.
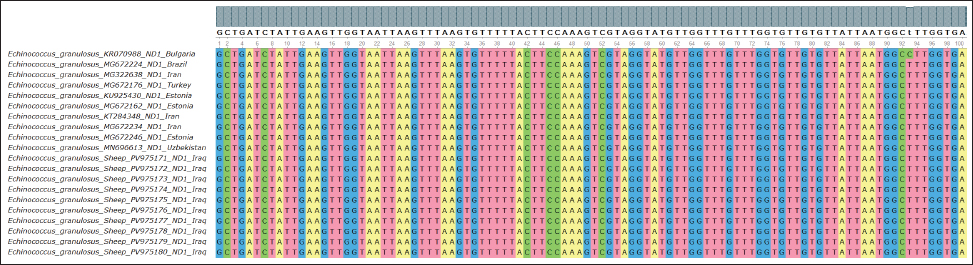
Fig. 12. Multiple sequence alignment of the partial ND1 gene in E. granulosus shows similarities and differences between the identified sequences in sheep, highlighting extremely limited genetic differences, further supporting the conserved nature of this mitochondrial gene in sheep isolates from the region. The phylogenetic trees from our gene sequences clearly grouped the sheep samples with known E. granulosus sensu stricto strains. This means that the parasites isolated from sheep in this study are closely related to standard reference types. Hamad et al. (2025) reported a similar result. They used different genes but found the same pattern in Iraqi sheep. These findings confirm that sheep continue to carry stable parasite strains. The close association between dogs and sheep explains why the parasite remains genetically stable. Larrieu et al. (2019) also highlighted this pathway. They called the dog-sheep link the main driver of the spread of CE. Koziol and Brehm (2015) noted that the parasite does not face strong evolutionary pressure in intermediate hosts. This allows the same strain to persist over time and across regions. Our results agree with these findings. The ND1 and CYT genes we studied showed almost no major changes compared with the global strains. Casulli et al. (2022a,b) also reported that parasite types often cross borders. Therefore, it is normal to find similar strains in different countries (Karim et al., 2018; Yaseen et al., 2020; Janabi, 2020; Janabi, 2021; Ghazi et al., 2024). This shows that molecular methods are strong and can be used in many countries and for different animals. The use of multilocus sequencing confirmed this. This helped us prove the identity and genetic closeness of the local sheep parasites. The ND1 and CYT genes in this study closely matched those in other samples from Eurasia. This means that the parasites in Iraqi sheep belong to the common G1 group. This group is stable and widespread. used similar tools to study Eimeria in sheep. They also found that molecular tests are effective for animal diseases (Sabeeh et al., 2018; Alfatlawi et al., 2021; Janabi, 2021; Klaif et al., 2022). In this study, we used four genes. All of them gave consistent results. This shows that multilocus testing is reliable. This method can also help future efforts to control CE in Iraq (Saied et al., 2022; Qui, 2023; Saied and Hameed, 2023; Maty, 2023; Al-Niaeemi and Dawood, 2023; Al-Taee et al., 2023; Mizory and Altaee, 2023; Hassan et al., 2024; Rahawi et al., 2024; Al-Taee, 2024; Al-Taee et al., 2024; Nofiu and Baharudin, 2024; Al-Kattan and Al-Bassam, 2024). ConclusionThis study showed that sheep in Diwaniyah, Iraq, are infected with E. granulosus strains belonging to the G1 genotype. This is the same type found in many countries. We studied four genes—ACT-2, COX1, CYT, and ND1. All the results confirmed that sheep carry a stable version of the parasite. Very few genetic differences were observed. This proves that sheep are important in the parasite life cycle. The strong correlation with global strains also shows that CE in Iraq is linked to wider international patterns. Molecular tools such as those we used are helpful in tracking and understanding these parasites. They are useful for disease control and future surveillance. Acknowledgment The authors thank the College of Veterinary Medicine at the University of Al-Qadisiyah for their continuous support. Their assistance with laboratory work and guidance throughout this research was greatly appreciated. Funding The authors have self-funded the study. No external funding source is available. Authors’ contributions All authors have participated in the study. Conflict of interest The authors have no conflicts of interest to declare. Data availability Data are available when requested by the corresponding author. ReferencesAbd-Alhassen, J.K., Janabi, A.H.D. and Aboktifa, M.A. 2021. Antioxidant and antimicrobial evaluation of lycopene isolated from watermelon. Biochem. Cell. Arch. 21, 2905–2910. Alfatlawi, M.A., Jasim, A.A., Jarad, N.E. and Khlaif, S.F. 2021. Clinical and molecular identification of Theileria annulata strains in calves in Al-Diwaniyah. Iraqi J. Vet. Sci. 35(1), 115–119; doi:10.33899/ijvs.2020.126429 Ali, R., Khan, S., Khan, M., Adnan, M., Ali, I., Khan, T.A., Haleem, S., Rooman, M., Norin, S. and Khan, S.N. 2020. Medicinal plants used against Echinococcus granulosus: a systematic review. PLos One. 15(10), 240456;doi:10.1371/journal.pone.0240456 Al-Kattan, H.H. and Al-Bassam, N.S. 2024. Sweet bean and roselle seeds improve growth of common carp. Mesopotamia J. Agric. 52(2), 58–67;doi:10.33899/mja.2024.146824.1374 Al-Niaeemi, B.H. and Dawood, M.H. 2023. Bioaccumulation of heavy metals in Bothriocephalus acheilognathi and Cyprinus carpio. Iraqi J. Vet. Sci. 37(4), 899–905;doi:10.33899/ijvs.2023.137615.2715 Al-Taee, N.T., Mohammad, M.A. and Abdul-Majeed, A.F. 2024. Effect of replacing animal protein with duckweed on carp growth performance. Mesopotamia. J. Agric. 52(3), 22–36; doi:10.33899/mja.2024.145298.1316 Al-Taee, S.K. 2024. Immunohistochemical and semiquantitative analysis of carp gills exposed to sodium thiosulfate. Iraqi. J. Vet. Sci. 38(1), 191–198; doi:10.33899/ijvs.2023.140746.3086 Al-Taee, S.K., Al-Zubaidy, M.H., Mustafa, K.A. and Ismail, H.K. 2023. Role of Saccharomyces cerevisiae in alleviating copper sulfate toxicity in Cyprinus carpio. Iraqi J. Vet. Sci. 37(3), 573–580;doi:10.33899/ijvs.2023.135148.2449 Anvari, D., Pourmalek, N., Rezaei, S., Fotovati, A., Hosseini, S.A., Daryani, A., Spotin, A., Sarvi, S., Hosseini, M., Narouei, M.R., Kalkali, M., Pendar, F. and Gholami, S. 2021. Hydatidosis in camels: global status and genetic data. Parasitology 148(3), 259–273; doi:10.1017/S0031182020001705 Anvari, D., Rezaei, F., Ashouri, A., Rezaei, S., Majidiani, H., Pagheh, A.S., Rezaei, F., Shariatzadeh, S.A., Fotovati, A., Siyadatpanah, A., Gholami, S. and Ahmadpour, E. 2021. Future prospects of E. granulosus vaccines: a systematic review. Transbound. Emerg. Dis. 68(3), 1080–1096;doi:10.1111/tbed.13772 Aregawi, W.G., Levecke, B., Ashenafi, H., Byaruhanga, C., Kebede, N., Mulinge, E., Wassermann, M., Romig, T., Dorny, P. and Dermauw, V. 2024. Epidemiology of Echinococcus granulosus in the Greater Horn of Africa. PLos Negl. Trop. Dis. 18(1), 11894; doi:10.1371/journal.pntd.0011894 Borhani, M., Fathi, S., Harandi, M.F., Casulli, A., Ding, J., Liu, M., Zhang, W. and Wen, H. 2024. Control measures with focus on vaccines. Parasitol. Vectors 17(1), 533; doi:10.1186/s13071-024-06581-2 Casulli, A., Massolo, A., Saarma, U., Umhang, G., Santolamazza, F. and Santoro, A. 2022. Species and genotypes causing human cystic echinococcosis in Europe (2000–2021). Parasitol. Vectors 15, 109; doi:10.1186/s13071-022-05197-8 García-Méndez, N., Manterola, C., Totomoch-Serra, A., Riffo-Campos, A.L. and Brito-Carreón, C.A. 2022. Proteomic profile of Echinococcus granulosus: a systematic review. J. Parasitol. 108(1), 64–69; doi:10.1645/20-86 Gharbi, M. and Giraudoux, P. 2024. Cystic echinococcosis in Tunisia: a One Health perspective. Parasite 31, 30;doi:10.1051/parasite/2024029 Ghazi, A.M., Al-Bayati, M.A. and Janabi, A.H.D. 2024. Metabolomics alterations from phytosomal propolis and lycopene in rats with induced BPH. Iraqi. J. Vet. Sci. 38(Suppl I–IV), 7–15;doi:10.33899/ijvs.2024.147764.3531 Hamad, W.A., Jawad, T.I. and Atiya, W.H. 2025. Phylogenetic evaluation of antigen B gene of E. granulosus. Open Vet. J. 15(2), 1043–1049;doi:10.5455/OVJ.2025.v15.i2.52 Hassan, W.S., Abdulrazzaq, K.M., Al-Obaidi, Q.T. and Al-Azow, K.A. 2024. Molecular detection of Anaplasma platys in dogs in Nineveh. Iraqi. J. Vet. Sci. 38(3), 677–682; doi:10.33899/ijvs.2024.148465 Hua, R.Q., Du, X.D., He, X., Gu, X.B., Xie, Y., He, R., Xu, J., Peng, X.R. and Yang, G.Y. 2022. Genetic diversity of Echinococcus granulosus sensu lato in China. Transbound. Emerg. Dis. 69(5), e1382–e1392;doi:10.1111/tbed.14469 Janabi, A.H.D. 2020. Anti-SARS-CoV-2 RdRp drugs based on docking: milbemycin, Ivermectin, Baloxavir. Avicenna J. Med. Biotechnol. 12(4), 246–250. Janabi, A.H.D. 2021. Molecular docking analysis of anti-SARS-CoV-2 ligands targeting spike glycoprotein and 3CLpro. J. Med. Signals Sens. 11(1), 31–36. Karim, S.M., Mansour, K.A., Janabi, A.H.D. and Al-Nakeeb, N.K.M. 2018. First phylogenetic characterization of pseudocowpox virus from cattle in Iraq. Iraqi. J. Vet. Sci. 33(1), 123–126; doi:10.33899/ijvs.2019.125525 Karshima, S.N., Ahmed, M.I., Adamu, N.B., Magaji, A.A., Zakariah, M. and Mohammed, K. 2022. Africa-wide meta-analysis of echinococcosis. Parasitol. Vectors 15, 357; doi:10.1186/s13071-022-05474-6 Khademvatan, S., Majidiani, H., Foroutan, M., Hazrati Tappeh, K., Aryamand, S. and Khalkhali, H.R. 2019. Echinococcus granulosus genotypes in Iran: a systematic review. J. Helminthol. 93(2), 131–138;doi:10.1017/S0022149X18000275 Klaif, S.F., Jassim, A., Alfatlawi, M.A. and Ali, M.J. 2022. MSP-4 gene-based detection of Anaplasma marginale from sheep in Al-Diwaniyah. Iraqi. J. Vet. Sci. 36(1), 85–88. Koziol, U. and Brehm, K. 2015. Recent advances in Echinococcus genomics and stem cell research. Vet. Parasitol. 213(3–4), 92–102;doi:10.1016/j.vetpar.2015.07.031 Larrieu, E., Gavidia, C.M. and Lightowlers, M.W. 2019. Control of cystic echinococcosis: background and prospects. Zoonoses Public Health 66(8), 889–899; doi:10.1111/zph.12649 Manterola, C., Totomoch-Serra, A., Rojas, C., Riffo-Campos, A.L. and García-Méndez, N. 2022. Echinococcus granulosus sensu lato genotypes in different hosts worldwide: a systematic review. Acta Parasitol. 67(1), 161–185;doi:10.1007/s11686-021-00439-8 Maty, H.N. 2023. Effect of sorbitol and L-carnitine on thyroid hormones and ATP in broilers. Iraqi. J. Vet. Sci. 37(3), 589–590; doi:10.33899/ijvs.2022.135305 Mizory, F.A. and Altaee, N.T. 2023. Growth performance of Cyprinus carpio fed Moringa oleifera floating leaves. Mesopotamia J. Agric. 51(1), 66–78; doi:10.33899/magrj.2023.137303.1210 Mokhtaria, K. and Ammar, S.S.M. 2019. Frozen hydatid cysts can replace incineration and sterilize cysts. Open. Vet. J. 9(1), 1–4; doi:10.4314/ovj.v9i1.1 Nofiu, N.B. and Baharudin, S.A. 2024. Vulnerability of smallholder farmers to flooding and poverty: a systematic review. Mesopotamia. J. Agric. 52(2), 1–13;doi:10.33899/mja.2024.149253.011424 Ohiolei, J.A., Yan, H.B., Odeniran, P.O., Li, L., Shumuye, N.A., Qurishi, S.A., Isaac, C., Fu, B.Q. and Jia, W.Z. 2022. Echinococcus granulosus sensu lato in animal intermediate hosts: organ location. Vet. Parasitol. 304, 109695; doi:10.1016/j.vetpar.2022.109695 Qui, N.H. 2023. Baker’s yeast (Saccharomyces cerevisiae) and its applications in poultry production: a review. Iraqi. J. Vet. Sci. 37(1), 213–221;doi:10.33899/ijvs.2022.132912.2146 Rahawi, A.M., Al-Taee, S.K., Ali, F.F., Altaey, O.Y. and Abdullah, D.A. 2024. Protective role of biosynthetic silver nanoparticles in broilers with aflatoxicosis: spleen histopathology. Iraqi J. Vet. Sci. 38(3), 565–572;doi:10.33899/ijvs.2024.146024.3414 Rojas-Pincheira, C., Manterola, C., García-Méndez, N. and Totomoch-Serra, A. 2022. Spatial distribution of E. granulosus in America. Rev. Med. Chil. 150(7), 944–957;doi:10.4067/s0034-98872022000700944 Sabeeh, S.A., Esraa, T.M., Alelaah, F.A., Al-Baghdadi, R.J.T. and Janabi, A.H.D. 2018. Metronidazole as feed additive targeting archaea to reduce frothy bloat in cows. Biochem. Cell. Arch. 18(2) , 2023–2027. Saied, A.F., Al-Taee, S.K. and Al-Taee, N.T. 2022. Morphohistopathological alterations in gills and CNS of carp exposed to lethal copper sulfate concentration. Iraqi. J. Vet. Sci. 36(4), 981–989;doi:10.33899/ijvs.2022.132781.2131 Saied, M.Q. and Hameed, H.M. 2023. Bioceutical role of nano and organic selenium in reproductive traits of laying hens. Iraqi J. Vet. Sci. 37(2), 325–331; doi:10.33899/ijvs.2022.134401.2364 Samei, A. and Khedri, M. 2025. Immunotherapeutic potential of hydatid cyst antigens. Iran. J. Allergy. Asthma Immunol. 24(3), 259–267. Shams, M., Khazaei, S., Naserifar, R., Shariatzadeh, S.A., Anvari, D., Montazeri, F., Pirestani, M. and Majidiani, H. 2022. Global distribution of Echinococcus granulosus genotypes in canids: a systematic review. Parasitology 149(9), 1147–1159; doi:10.1017/S0031182022000658 Tamarozzi, F., Deplazes, P. and Casulli, A. 2020. Reinventing the wheel of Echinococcus granulosus sensu lato transmission to humans. Trends. Parasitol. 36(5), 427–434; doi:10.1016/j.pt.2020.02.004 Tamarozzi, F., Legnardi, M., Fittipaldo, A., Drigo, M. and Cassini, R. 2020. Echinococcus granulosus in Mediterranean and Balkan countries. PLos Negl. Trop. Dis. 14(8), 8519;doi:10.1371/journal.pntd.0008519 Wen, H., Vuitton, L., Tuxun, T., Li, J., Vuitton, D.A., Zhang, W. and Mcmanus, D.P. 2019. Echinococcosis: advances in the 21st century. Clin. Microbiol. Rev. 32(2), e00075–e00018;doi:10.1128/CMR.00075-18 Woolsey, I.D. and Miller, A.L. 2021. Echinococcus granulosus sensu lato and Echinococcus multilocularis: a review. Res. Vet. Sci. 135, 517–522;doi:10.1016/j.rvsc.2020.11.010 Yaseen, M.M., Karawan, A.C., Alfatlawi, M.A.A. and Janabi, A.H.D. 2020. Role of gut bacterial cytochrome-P450 in degradation of temephos insecticide in mosquito larvae. Ann. Trop. Med. Public Health 23(1), S412;doi:10.36295/ASRO.2020.23126 | ||
| How to Cite this Article |
| Pubmed Style Hajeel MS, Alfatlawi MAA. Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Vet. J.. 2026; 16(2): 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 Web Style Hajeel MS, Alfatlawi MAA. Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. https://www.openveterinaryjournal.com/?mno=297313 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.48 AMA (American Medical Association) Style Hajeel MS, Alfatlawi MAA. Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Vet. J.. 2026; 16(2): 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 Vancouver/ICMJE Style Hajeel MS, Alfatlawi MAA. Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 Harvard Style Hajeel, M. S. & Alfatlawi, . M. A. A. (2026) Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Vet. J., 16 (2), 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 Turabian Style Hajeel, Marwa Saleem, and Monyer Abdulamier Abd Alfatlawi. 2026. Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Veterinary Journal, 16 (2), 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 Chicago Style Hajeel, Marwa Saleem, and Monyer Abdulamier Abd Alfatlawi. "Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep." Open Veterinary Journal 16 (2026), 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 MLA (The Modern Language Association) Style Hajeel, Marwa Saleem, and Monyer Abdulamier Abd Alfatlawi. "Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep." Open Veterinary Journal 16.2 (2026), 1315-1328. Print. doi:10.5455/OVJ.2026.v16.i2.48 APA (American Psychological Association) Style Hajeel, M. S. & Alfatlawi, . M. A. A. (2026) Genetic detection and phylogenetic assessment of multiple molecular markers of Echinococcus granulosus from sheep. Open Veterinary Journal, 16 (2), 1315-1328. doi:10.5455/OVJ.2026.v16.i2.48 |