| Short Communication | ||
Open Vet. J.. 2026; 16(3): 1932-1940 Open Veterinary Journal, (2026), Vol. 16(3): 1932-1940 Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014)Silvia Penelo1,2, Isabel Simarro3, Manuel Fuertes-Recuero1,4*, Tania Ayllón1,3 and Gustavo Ortiz-Díez1,21Complutense Veterinary Hospital, Veterinary Teaching School, Complutense University of Madrid, Madrid, Spain 2Department of Animal Medicine and Surgery, Veterinary Teaching School, Complutense University of Madrid, Madrid, Spain 3Department of Animal Health, Veterinary Teaching School–VISAVET Health Surveillance Center, Complutense University of Madrid, Madrid, Spain 4Department of Physiology, Veterinary Teaching School, Complutense University of Madrid, Madrid, Spain *Corresponding Author: Manuel Fuertes Recuero. Department of Animal Medicine and Surgery, Veterinary Teaching School, Complutense University of Madrid, Madrid, Spain. Email: manufuer [at] ucm.es Submitted: 17/11/2025 Revised: 25/01/2026 Accepted: 05/02/2026 Published: 31/03/2026 © 2026 Open Veterinary Journal
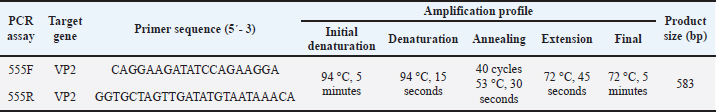
AbstractBackground: Canine parvovirus type 2 (CPV-2) remains a major cause of morbidity and mortality in young dogs. Clinical and molecular data obtained between 2003 and 2014 from a veterinary teaching hospital in central Spain were analyzed to characterize circulating CPV-2 antigenic variants and associated in-hospital outcomes. As the samples were collected more than a decade ago, the reported frequencies represent historical CPV-2 data rather than the current circulation. Aim: This study aimed to describe the historical distribution of CPV-2 antigenic variants (2a, 2b, and 2c) detected in hospital-presenting clinical samples collected between 2003 and 2014, document rare detections of feline panleukopenia virus (FPV) and the Cornell vaccine strain, and analyze clinical variables associated with in-hospital mortality. Methods: A retrospective series of 110 clinical samples were analysed using rapid antigen tests and polymerase chain reaction. Adequate amplification yielded 583-bp VP2 fragments, of which 50 were selected for sequencing (42 produced high-quality reads). Species identification (CPV-2 vs. FPV) was based on full-length amplicon alignment and phylogenetic clustering. Amino acid residue 426 was used as a supporting marker to distinguish the CPV-2a, CPV-2b, and CPV-2c antigenic variants. Logistic regression models estimated odds ratios (OR) with 95% confidence intervals (CI). Results: In the sequenced archival cases, CPV-2c was the predominant variant (42.9%, 18/42), followed by CPV-2a (31.0%, 13/42), the Cornell vaccine strain (11.9%, 5/42), CPV-2b (9.5%, 4/42), and FPV (4.8%, 2/42). Small-breed dogs (<15 kg) exhibited higher odds of in-hospital death (OR=2.74; 95% CI=1.18–6.37). Dogs presenting with gastrointestinal signs in combination with neurological and/or respiratory signs had markedly increased odds of mortality (OR=9.14; 95% CI=2.29–36.40). No statistically significant associations were observed between mortality and antigenic variant or vaccination status. Conclusion: This archival cohort provides a historical reference for CPV epidemiology in central Spain. CPV-2c was the most common variant. Breed size and multisystemic involvement were significantly associated with in-hospital mortality. However, contemporary sequencing would be necessary to monitor the evolution of CPV-2 beyond the study period due to the historical nature of the study. Keywords: Canine parvovirus, Clinical epidemiology, Hospital mortality, Antigenic variation, VP2. IntroductionCanine parvovirus type 2 (CPV-2) is a highly contagious enteric virus that induces considerable morbidity and mortality, particularly among unvaccinated puppies and juvenile dogs worldwide (Mazzaferro, 2020). CPV-2 is classified within the family Parvoviridae, subfamily Parvovirinae, and genus Protoparvovirus, under the species Carnivore protoparvovirus 1, which also includes feline panleukopenia virus (FPV), in accordance with ICTV taxonomy. Both CPV-2 and FPV exhibit a broad host range, with documented natural infections in domestic cats (Felis catus), raccoons (Procyon lotor), raccoon dogs (Nyctereutes procyonoides), minks (Neogale vison), and foxes (Vulpes vulpes) (Allison et al., 2016; Shackelton et al., 2005). Clinical manifestations of CPV-2 infection primarily affect puppies and typically include severe gastroenteritis, characterized by vomiting and diarrhea, as well as myocarditis and myocardial fibrosis (Ford et al., 2017; Mazzaferro, 2020). The CPV-2 genome comprises approximately 5.2 kilobases (kb) of single-stranded DNA, encoding two nonstructural proteins (NS1 and NS2) and two structural proteins (VP1 and VP2) (Reed et al., 1988). VP2, the major capsid protein, influences the host range and contains major antigenic epitopes targeted by neutralizing antibodies (Parrish, 1999). The virus is noted for rapid evolution (approximately 10⁻⁴ substitutions per site per year), particularly in VP2, supporting the continued emergence of antigenic variants (Cotmore et al., 2014). CPV-2 was first identified in canines in the late 1970s, causing acute and frequently fatal disease (Appel et al., 1979). Thereafter, several antigenic variants replaced the original virus. CPV-2a, which can infect both dogs and cats, emerged following mutations in VP2 (L87M, I101T, A300G, D305Y) (Buonavoglia et al., 2001; Allison et al., 2016). CPV-2b, first detected in the United States in 1984 and distinguished by N426D, co-circulates with CPV-2a with region-specific prevalence (Parrish, 1999; Zhou et al., 2017; Kwan et al., 2021). CPV-2c, characterized by D426E, was initially reported in Italy in 2000 (Buonavoglia et al., 2001) and subsequently disseminated globally (Decaro et al., 2007). Although initially considered less pathogenic (Decaro et al., 2005), CPV-2c causes severe, frequently fatal hemorrhagic enteritis, including in vaccinated adult dogs (Decaro et al., 2008). Subsequently, CPV-2a and CPV-2b variants emerged, differing from earlier forms by a single VP2 amino acid substitution (S297A) (Ohshima et al., 2008; Mukhopadhyay et al., 2014; Chen et al., 2019). Most studies indicate that current vaccines provide broad cross-protection against circulating variants despite the emergence of CPV-2 antigenic variants (Wilson et al., 2014; Packianathan et al., 2022). The emergence and distribution of CPV-2 have been documented in a small number of studies in Spain (Calatayud et al., 2020; de Ybañez et al., 1995; Decaro et al., 2006). The earliest study, conducted before the appearance of CPV-2c, reported CPV-2b as the predominant variant (De Ybanez et al., 1995). A subsequent study in Tarragona documented the CPV-2c distribution in the region (Decaro et al., 2006). More recently, canine and feline parvovirus sequences have been detected in wild carnivores in Spain (Calatayud et al., 2020; Canuti et al., 2025), suggesting the presence of potential environmental reservoirs or interspecies transmission between domestic and wild populations. However, no recent variant dynamics in domestic dogs and cats in Spain have been reported. CPV-2 remains a major cause of severe gastroenteritis in young dogs and can be associated with in-hospital mortality. Therefore, identifying host and clinical factors associated with mortality is important for risk stratification and case management. From an epidemiological standpoint, the limited number of studies conducted in Spain describing the distribution of CPV-2 variants in domestic hosts underscores the value of locally grounded datasets. In this context, a hospital-based series combining characterization of antigenic variants (CPV-2a/2b/2c) with clinically meaningful outcomes provides a robust baseline reference for future surveillance and comparative analyses. This study aimed to analyze retrospective (2003–2008) and prospective hospital-presenting cases (2009–2014) at a veterinary teaching hospital in central Spain to characterize the distribution of CPV-2 antigenic-variant (2a/2b/2c), report rare detections (FPV and the Cornell vaccine strain), and assess associations with in-hospital mortality. Materials and MethodsStudy setting and sample collectionA total of 110 samples (105 from dogs and five from cats) were collected between 2003 and 2014. Data collection was partly retrospective (2003–2008) and partly prospective (2009–2014) within the same hospital setting. Canine samples were obtained at the Veterinary Clinical Hospital of the Complutense University of Madrid and comprised 101 rectal swabs from symptomatic dogs or tissue macerates from deceased dogs that had undergone necropsy due to suspected parvoviral infection. The remaining 4 rectal swab samples were collected from asymptomatic in-contact dogs that had been exposed to a confirmed parvovirus-positive patient. All samples were collected by veterinarians as part of routine clinical practice. Sampling was convenience-based and was not intended to estimate population prevalence; no year-by-year analyses were performed. Demographic and clinical data, including age, sex, breed (4 categories by average adult bodyweight), vaccination status, and clinical signs, were recorded. Clinical signs (recorded during hospitalization) were classified into digestive (vomiting, diarrhea, and dehydration), neurological (ataxia, muscle tremors, or seizures), and respiratory (coughing, nasal discharge, or dyspnea). For feline patients, 5 samples were collected following a positive rapid in-house test, with owner consent recorded as for canine cases. Deaths were attributed to parvoviral infection (CPV-2/FPV) when infection was confirmed by laboratory tests and no alternative cause was identified; fatal outcomes occurred in the context of typical CPV-related complications. Mortality was subsequently analyzed in association with demographic variables and other characteristics (antigenic variant, vaccination status, and concurrent diseases). Initial diagnostic testingThe initial screening of all samples was conducted using rapid in-house diagnostic tests, including immunochromatographic assays such as INgezim PARVO-CROM® (Ingenasa, Spain), FASTest® PARVO Card (Megacor Diagnostik, Austria), dual CPV/canine coronavirus (CCV) antigen assays (Minitest CPV/CCV Ag®; SensPERT CCV Ag/CPV Ag®), or an enzyme-linked immunosorbent assay-based SNAP® Parvovirus test (IDEXX, USA). The rapid tests were part of routine diagnostic-laboratory quality assurance (specificity/sensitivity; data not shown). Canine coronavirus (CCV) was assessed using dual CPV/CCV antigen assays in routine practice. Other concurrent infectious diseases (e.g., canine distemper virus) were investigated when clinically suspected, according to the judgment of the attending clinician. No standardized screening protocol was applied for concurrent infections across the entire cohort. Sample preparation and polymerase chain reactionBoth stool and tissue samples used in the retrospective part of the study (2003–2008) were stored at −80°C until further analysis. Fecal samples and rectal swabs (approximately 1 g) were emulsified in 1 ml of phosphate-buffered saline (pH 7.2). For necropsy, sections (1 cm³) from the liver, spleen, kidney, heart, brain, lymph nodes, small intestine, and large intestine were homogenized in 10 ml phosphate-buffered saline using a Stomacher® 3500 (Seward, United Kingdom) and subsequently centrifuged at 2,000 rpm for 10 minutes. Supernatants were processed for DNA extraction (QuickGene-mini80) according to the manufacturer’s instructions; pre-digestion included proteinase K and dithiothreitol as per standard operating procedures (SOPs). Conventional polymerase chain reaction (PCR) targeting a 583-bp fragment of the parvovirus VP2 gene was used to detect parvoviral DNA. The primers were used as previously described by Buonavoglia et al. (2001) (Table 1). PCR reactions were performed under the following thermocycling conditions: initial denaturation at 94 °C for 10 minutes; 40 cycles of 94 °C for 30 seconds, annealing at 50 °C for 60 seconds, and extension at 72 °C for 1 minute; followed by a final extension at 72 °C for 10 minutes. The amplicon spans the MboII site encoding residue 426 (useful for antigenic-variant calling); species were not assigned to residue 426 alone. Table 1. Conventional PCR protocol for parvovirus detection (VP2, 583 bp).
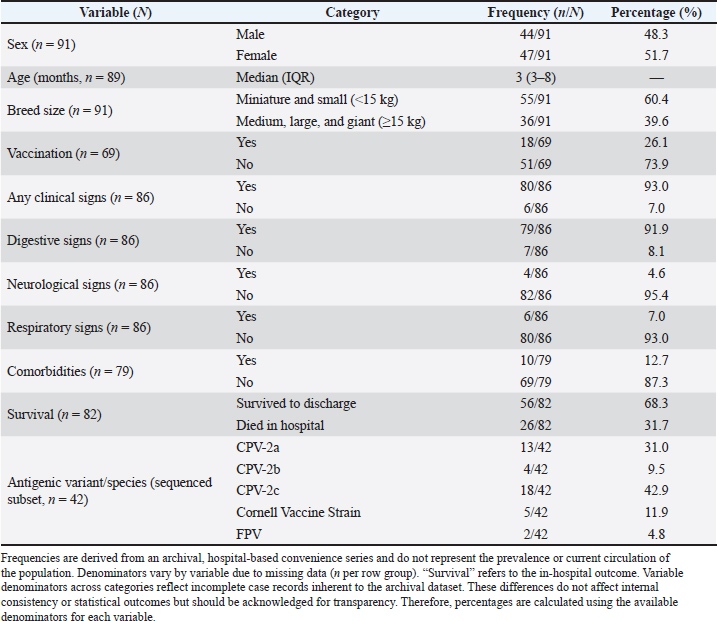
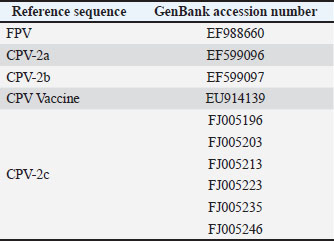
Sequencing and assignment of species/variantsA total of 50 PCR-positive samples were selected for sequencing using a convenience sampling approach, with the criterion of including a minimum of five PCR-positive samples per year (when available) to ensure temporal coverage across 2003–2014 within available sequencing resources. Of these, 42 produced VP2 amplicons of sufficient quality and quantity for downstream sequencing; the remaining samples were excluded due to degraded DNA or unreadable chromatograms in archived extracts. Amplicons were purified using the MinElute® PCR Purification Kit (Qiagen, Germany) and sequenced at the Madrid Technological Park (Faculty of Biological Sciences, Complutense University of Madrid). Sequence inspection was performed in Chromas Lite, and alignment with CPV and FPV reference sequences (NCBI) was performed using MEGA v6. The accession numbers for all reference sequences included in the alignment/phylogenetic reference panel are provided in Table S1. The species designation (CPV-2 versus FPV) was determined based on full-amplicon alignment (583 bp) and phylogenetic clustering with reference panels. The antigenic variants (2a/2b/2c) were assigned according to residue 426, which was not used for species differentiation (Allison et al., 2016). The original alignments and chromatogram files required for robust phylogenetic reconstruction were not retained because of the archival nature of the cohort and the period during which the sequences were generated. Moreover, systematic deposition of sequences in public repositories was not yet standard practice, precluding phylogenetic analysis and representing a limitation of the study. Statistical analysisStatistical analyses were conducted at the individual-patient level (n=99 dogs), using one record per animal. The sample-level dataset (110 samples) was only used for molecular and sequencing analyses. The dependent variable was in-hospital mortality (yes/no). Age was dichotomized at 3 months (≤3 vs. >3 months), corresponding to the sample median and clinical relevance for very young puppies. Descriptive statistics included mean ± SD for normally distributed variables and median with interquartile range for non-normally distributed variables. The Kolmogorov–Smirnov and Shapiro–Wilk tests were used to assess normality. Univariable logistic regression was initially performed to evaluate associations between mortality and independent variables (age, breed size, vaccination status, clinical signs, concurrent diseases, and antigenic variant). Quantitative variables were dichotomized according to the median or clinical plausibility of the dataset; for age, the cutoff was 3 months. Variables with p < 0.10 in univariable analyses, together with clinically relevant factors, were included in a multivariable logistic regression model using stepwise forward selection. For breed size, odds ratios (OR) were coded as small (<15 kg) medium/large/giant (reference). Effects are reported as ORs with 95% confidence intervals (CI). Model calibration was evaluated using the Hosmer–Lemeshow test, and discrimination was assessed using the area under the receiver operating characteristic curve (AUC). Internal validation was performed using bootstrap resampling (100 iterations with replacement) at the patient level, to estimate optimism-corrected model performance and account for within-patient clustering. All analyses were conducted in Stata 15.0, and statistical significance was set at p < 0.05. Analyses were not stratified by year, and temporal trends were not evaluated. Ethical approvalThe clinical veterinary procedures followed were non-experimental under national legislation; owners were informed of the study’s purpose, and informed consent was obtained for the collection of fecal samples from symptomatic dogs and organ samples from deceased individuals (Directive 2010/63/EU; Royal Decree 53/2013). Results and DiscussionThe CPV-2-positive rate obtained in the study was 97.3% (107/110). Frequencies refer to the entire archival period (2003–2014), unless otherwise specified. The positive samples included 101 symptomatic dogs, 4 asymptomatic in-contact dogs, and 2 cats. Sequencing was performed on a subset selected from available samples across years to ensure representation of the 2003–2014 period; the dataset is not population-representative. VP2 amplicons from 42 near full-length samples were successfully sequenced and analyzed. Sequence alignment against CPV/FPV reference panels supported classification as CPV-2a, CPV-2b, CPV-2c, FPV, or the Cornell vaccine strain. Residue 426 informed antigenic-variant assignment, whereas species determination relied on full-amplicon alignment against CPV/FPV reference sequences (Allison et al., 2016). Among the sequenced cases, the variant frequencies were as follows: CPV-2c, 42.9% (18/42); CPV-2a, 31.0% (13/42); Cornell vaccine strain, 11.9% (5/42); CPV-2b, 9.5% (4/42); and FPV, 4.8% (2/42). These frequencies represent archival, hospital-based cases and do not reflect the prevalence or contemporary distribution of the population. The median patient age was 3 months (interquartile range, 3-8 months). Males comprised 48.3% and females 51.6% of the infected animals. Breed distribution included 61% miniature or small breeds (<15 kg) and 39% medium, large, or giant breeds. Gastrointestinal signs (mostly vomiting and diarrhea, frequently hemorrhagic, with dehydration) were observed in 92% of the dogs. Neurological signs (ataxia, tremors, seizures) occurred in 4.6%, and respiratory signs (cough, nasal discharge, dyspnea) in 7.0%. Multisystemic signs likely reflect severe disease and potential co-infections, such as canine distemper virus, which were not systematically excluded. Approximately 7% of the dogs were asymptomatic at the time of testing. Comorbidities, including canine coronavirus or parasitic infestations, were present in 12.7% (Table 2). Table 2. Demographic variables, vaccination status, clinical signs, survival, comorbidities, and antigenic variant/species were described.
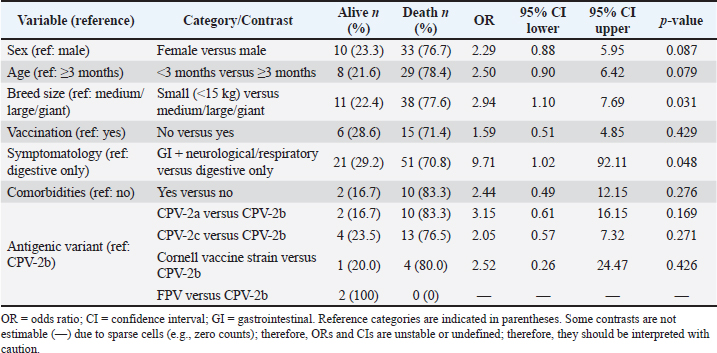
The univariable analysis identified associations between mortality and several variables, including breed size and symptomatology (Table 3). In multivariable logistic regression, small-breed size (<15 kg vs. medium/large/giant: OR 2.74; 95% CI 1.18–6.37) and multisystemic presentation (gastrointestinal plus neurological and/or respiratory signs versus mild: OR 9.14; 95% CI 2.29–36.40) remained independently associated with mortality (Table 4). These estimates indicate a moderate association for small-breed dogs and a strong association for multisystemic disease, albeit with wide confidence intervals reflecting limited precision. Vaccination status was not statistically significant for in-hospital mortality in this hospital-based series of PCR-confirmed CPV infections (unvaccinated vs vaccinated: OR 1.59, 95% CI 0.51–4.85; p=0.429), although the point estimate suggested higher odds of death among unvaccinated dogs (Table 3). Vaccination remains the cornerstone of CPV prevention, and unvaccinated puppies are widely reported to be at increased risk of severe disease. The absence of statistical significance for vaccination status in our analysis should be interpreted cautiously because vaccination histories were incomplete/heterogeneous and the analysis was restricted to infected dogs presenting to the hospital; therefore, these data do not estimate vaccine effectiveness against infection and provide limited power to detect a mortality association among cases. The antigenic variant and comorbidities were also not statistically significant in this dataset (Table 3). The model discrimination was moderate (AUC 0.662; 95% CI 0.527–0.796), indicating that the model supports etiologic interpretation of associations rather than individual-level prognostic accuracy. The calibration was acceptable (Hosmer–Lemeshow p=0.830). The association between small-breed status and mortality is consistent with previous observations in CPV-infected dogs (Ling et al., 2012). Table 3. Univariable logistic regression: mortality by candidate predictors.
Table 4. Multivariable logistic regression: risk factors associated with mortality in the hospital.
CPV-2 remains a major cause of morbidity and mortality in young dogs (Mazzaferro, 2020). Initial reports suggested that CPV-2c was less pathogenic than earlier variants (Decaro et al., 2005), but subsequent studies demonstrated its capacity to induce severe and frequently fatal hemorrhagic enteritis, including in vaccinated adult dogs (Decaro et al., 2008). High virulence has also been reported for circulating CPV-2a lineages in Asia (Chen et al., 2019). This archival series provides a historical baseline for comparison with contemporary circulation and clinical impact, without implying any variant’s current dominance. FPV detection in dogs was rare (2/42 sequenced cases). Detections classified as the Cornell vaccine strain likely reflect the presence of the vaccine virus, for example, post-vaccinal shedding (Decaro et al., 2007); however, the attribution of clinical disease to the vaccine strain cannot be determined from archival data. Consistent use of ICTV-compliant terminology (antigenic variants vs. strains) avoids conflation of antigenic categories with isolate-level designations, enhancing comparability across studies. This study is limited by its hospital-based, archival convenience sampling (2003–2014), and the frequencies reported may not reflect the population prevalence or current viral circulation. As the study covers a historical period and viral evolution and variant circulation have continued to change since then, these findings provide context for past patterns but should not be interpreted as representative of the contemporary epidemiological situation. Temporal trends were not assessed, and only 42 samples were sequenced due to constraints on DNA quality and chromatogram readability, potentially introducing selection bias. Species and variant assignment relied on a 583 bp VP2 fragment and phylogenetic clustering; although more robust than residue 426 alone, some misclassification risk remains (Allison et al., 2016). VP2 sequences could not be deposited in a public repository, as the original sequence files from this archival cohort were not available, limiting external reproducibility. Although clinically suspected concurrent infections were investigated, the absence of systematic screening across all cases may have resulted in residual bias, and multisystemic presentations may reflect severe parvoviral disease and/or undetected coinfections. ConclusionIn this archival, hospital-based cohort from central Spain (2003–2014), CPV-2c was the predominant variant among the sequenced cases, followed by CPV-2a, whereas CPV-2b and FPV were infrequently detected. Small-breed status and multisystemic clinical presentation were independently associated with in-hospital mortality. These data establish a documented historical basis, but conducting updated studies to establish the current circulation of the different viral strains is advisable. In addition, a systematic evaluation of coinfections is recommended. AcknowledgmentsNone. FundingThis study received no specific grant. Authors’ contributionsAll authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by SP, IS, TA, MFR, and GOD. The first draft of the manuscript was written by SP, and all authors commented on the previous versions of the manuscript. All authors have read and approved the final version of the manuscript. Conflict of interestThe authors declare no conflict of interest. Data availabilityAll data supporting this study’s findings are available within the manuscript; additional information is available from the corresponding author upon reasonable request. ReferencesAllison, A.B., Organtini, L.J., Zhang, S., Hafenstein, S.L., Holmes, E.C. and Parrish, C.R. 2016. Single Mutations in the VP2 300 Loop Region of the Three-Fold Spike of the Carnivore Parvovirus Capsid Can Determine Host Range. J. Virol. 90, 753–767; doi:10.1128/JVI.02636-15 Appel, M., Scott, F. and Carmichael, L. 1979. Isolation and immunisation studies of a canine parvo-like virus from dogs with haemorrhagic enteritis. Vet. Rec. 105, 156–159; doi:10.1136/vr.105.8.156 Buonavoglia, C., Martella, V., Pratelli, A., Tempesta, M., Cavalli, A., Buonavoglia, D., Bozzo, G., Elia, G., Decaro, N. and Carmichael, L. 2001. Evidence for evolution of canine parvovirus type 2 in Italy. J. Gen. Virol. 82(12), 3021–3025; doi:10.1099/0022-1317-82-12-3021 Calatayud, O., Esperón, F., Velarde, R., Oleaga, A., Llaneza, L., Ribas, A., Negre, N., Torre, A., Rodríguez, A. and Millán, J. 2020. Genetic characterization of Carnivore Parvoviruses in Spanish wildlife reveals domestic dog and cat-related sequences. Transbound. Emerg. Dis. 67, 626–634; doi:10.1111/tbed.13378 Canuti, M., Mira, F., Villanúa, D., Rodríguez-Pastor, R., Guercio, A., Urra, F. and Millán, J. 2025. Molecular ecology of novel amdoparvoviruses and old protoparvoviruses in Spanish wild carnivorans. Infect. Genet. Evol. 128, 105714; doi:10.1016/j.meegid.2025.105714 Chen, M.R., Guo, X.Y., Wang, Z.Y., Jiang, Y.T., Yuan, W.F., Xin, T., Hou, S.H., Song, T.Q., Lin, W.D., Zhu, H.F. and Jia, H. 2019. Isolation and sequence analysis of the complete VP2 gene of canine parvovirus from Chinese domestic pets and determination of the pathogenesis of these circulating strains in beagles. Pol. J. Vet. Sci. 22(2), 287–296. Cotmore, S.F., Agbandje-McKenna, M., Chiorini, J.A., Mukha, D.V., Pintel, D.J., Qiu, J., Soderlund-Venermo, M., Tattersall, P., Tijssen, P., Gatherer, D. and Davison, A.J. 2014. The family Parvoviridae. Arch. Virol. 159, 1239–1247; doi:10.1007/s00705-013-1914-1 De Ybanez, R., Vela, C., Cortes, E., Simarro, I. and Casal, J. 1995. Identification of types of canine parvovirus circulating in Spain. Vet. Rec. 136, 174–175; doi:10.1136/vr.136.7.174 Decaro, N., Desario, C., Addie, D.D., Martella, V., Vieira, M.J., Elia, G., Zicola, A., Davis, C., Thompson, G., Thiry, E., Truyen, U. and Buonavoglia, C. 2007. The Study Molecular Epidemiology of Canine Parvovirus, Europe. Emerg. Infect. Dis. 13, 1222–1224; doi:10.3201/eid1308.070505 Decaro, N., Desario, C., Campolo, M., Elia, G., Martella, V., Ricci, D., Lorusso, E. and Buonavoglia, C. 2005. Clinical and virological findings in pups naturally infected by canine parvovirus type 2 Glu-426 mutant. J. Vet. Diagn. Invest. 17, 133–138; doi:10.1177/104063870501700206 Decaro, N., Desario, C., Elia, G., Martella, V., Mari, V., Lavazza, A., Nardi, M. and Buonavoglia, C. 2008. Evidence for immunisation failure in vaccinated adult dogs infected with canine parvovirus type 2c. New Microbiol. 31, 125–130. Decaro, N., Martella, V., Desario, C., Bellacicco, A.L., Camero, M., Manna, L., D'Aloja, D. and Buonavoglia, C. 2006. First detection of canine parvovirus type 2c in pups with haemorrhagic enteritis in Spain. J. Vet. Med. B. Infect. Dis. Vet. Public Health 53, 468–472; doi:10.1111/j.1439-0450.2006.00974.x Ford, J., Mcendaffer, L., Renshaw, R., Molesan, A. and Kelly, K. 2017. Parvovirus Infection Is Associated With Myocarditis and Myocardial Fibrosis in Young Dogs. Vet. Pathol. 54, 964–971; doi:10.1177/0300985817725387 Kwan, E., Carrai, M., Lanave, G., Hill, J., Parry, K., Kelman, M., Meers, J., Decaro, N., Beatty, J.A., Martella, V. and Barrs, V.R. 2021. Analysis of canine parvoviruses circulating in Australia reveals predominance of variant 2b and identifies feline parvovirus-like mutations in the capsid proteins. Transbound. Emerg. Dis. 68, 656–666; doi:10.1111/tbed.13727 Ling, M., Norris, J.M., Kelman, M. and Ward, M.P. 2012. Risk factors for death from canine parvoviral-related disease in Australia. Vet. Microbiol. 158, 280–290; doi:10.1016/j.vetmic.2012.02.034 Mazzaferro, E.M. 2020. Update on Canine Parvoviral Enteritis. Vet. Clin. North. Am. Small. Anim. Pract. 50, 1307–1325; doi:10.1016/j.cvsm.2020.07.008 Mukhopadhyay, H.K., Matta, S.L., Amsaveni, S., Antony, P.X., Thanislass, J. and Pillai, R.M. 2014. Phylogenetic analysis of canine parvovirus partial VP2 gene in India. Virus. Genes. 48, 89–95; doi:10.1007/s11262-013-1000-5 Ohshima, T., Hisaka, M., Kawakami, K., Kishi, M., Tohya, Y. and Mochizuki, M. 2008. Chronological analysis of canine parvovirus type 2 isolates in Japan. J. Vet. Med. Sci. 70, 769–775; doi:10.1292/jvms.70.769 Packianathan, R., Hodge, A., Wright, J., Lavidis, L., Ameiss, K., Yip, H.Y.E., Akbarzadeh, M., Sharifian, M., Amanollahi, R., Khabiri, A. and Hemmatzadeh, F. 2022. Cross-Neutralization of Vanguard C4 Vaccine Against Australian Isolates of Canine Parvovirus Variants CPV-2a, CPV-2b, and CPV-2c. Viral Immunol. 35, 553–558; doi:10.1089/vim.2022.0027 Parrish, C.R. 1999. Host range relationships and the evolution of canine parvovirus. Vet. Microbiol. 69, 29–40; doi:10.1016/s0378-1135(99)00084-x Reed, A.P., Jones, E.V. and Miller, T.J. 1988. Nucleotide sequence and genome organization of canine parvovirus. J. Virol. 62(1), 266–276; doi:10.1128/JVI.62.1.266-276.1988 Shackelton, L.A., Parrish, C.R., Truyen, U. and Holmes, E.C. 2005. High rate of viral evolution associated with the emergence of carnivore parvovirus. Proc. Natl. Acad. Sci. U. S. A. 102, 379–384; doi:10.1073/pnas.0406765102 Wilson, S., Illambas, J., Siedek, E., Stirling, C., Thomas, A., Plevová, E., Sture, G. and Salt, J. 2014. Vaccination of dogs with canine parvovirus type 2b (CPV-2b) induces neutralising antibody responses to CPV-2a and CPV-2c. Vaccine 32, 5420–5424; doi:10.1016/j.vaccine.2014.07.102 Zhou, P., Zeng, W., Zhang, X. and Li, S. 2017. The genetic evolution of canine parvovirus - A new perspective. PLoS One 12, 175035; doi:10.1371/journal.pone.0175035 Supplementary MaterialTable S1. Reference sequences used for nucleotide alignment with selected clinical samples.
| ||
| How to Cite this Article |
| Pubmed Style Penelo S, Simarro I, Fuertes-recuero M, Ayllón T, Ortiz-díez G. Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 Web Style Penelo S, Simarro I, Fuertes-recuero M, Ayllón T, Ortiz-díez G. Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). https://www.openveterinaryjournal.com/?mno=297152 [Access: March 31, 2026]. doi:10.5455/OVJ.2026.v16.i3.48 AMA (American Medical Association) Style Penelo S, Simarro I, Fuertes-recuero M, Ayllón T, Ortiz-díez G. Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 Vancouver/ICMJE Style Penelo S, Simarro I, Fuertes-recuero M, Ayllón T, Ortiz-díez G. Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 Harvard Style Penelo, S., Simarro, . I., Fuertes-recuero, . M., Ayllón, . T. & Ortiz-díez, . G. (2026) Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 Turabian Style Penelo, Silvia, Isabel Simarro, Manuel Fuertes-recuero, Tania Ayllón, and Gustavo Ortiz-díez. 2026. Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 Chicago Style Penelo, Silvia, Isabel Simarro, Manuel Fuertes-recuero, Tania Ayllón, and Gustavo Ortiz-díez. "Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014)." doi:10.5455/OVJ.2026.v16.i3.48 MLA (The Modern Language Association) Style Penelo, Silvia, Isabel Simarro, Manuel Fuertes-recuero, Tania Ayllón, and Gustavo Ortiz-díez. "Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014)." doi:10.5455/OVJ.2026.v16.i3.48 APA (American Psychological Association) Style Penelo, S., Simarro, . I., Fuertes-recuero, . M., Ayllón, . T. & Ortiz-díez, . G. (2026) Canine parvovirus type 2 antigenic variants and in-hospital mortality in central Spain: Retrospective and prospective data (2003–2014). doi:10.5455/OVJ.2026.v16.i3.48 |