| Review Article | ||
Open Vet. J.. 2026; 16(3): 1438-1450 Open Veterinary Journal, (2026), Vol. 16(3): 1438-1450 Review Article Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in PakistanAsfand Yar Khan1, Mustofa Helmi Effendi2*, Aftab Shaukat3 and Muhammad Muneeb41Faculty of Veterinary Sciences, University of Veterinary and Animal Sciences, Swat, Pakistan 2Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 3College of Veterinary Medicine, South China Agricultural University, Guangzhou, China 4Faculty of Veterinary and Animal Sciences, Gomal University, Dera Ismail Khan, Pakistan *Corresponding Author: Mustofa Helmi Effendi. Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia. Email: mustofa-h-e [at] fkh.unair.ac.id Submitted: 28/10/2025 Revised: 29/01/2026 Accepted: 09/02/2026 Published: 31/03/2026 © 2026 Open Veterinary Journal
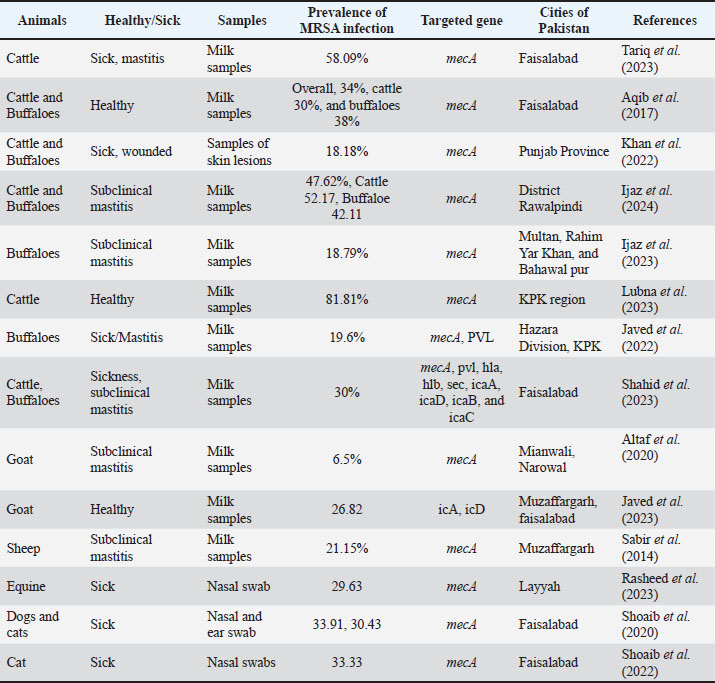
AbstractMethicillin-resistant Staphylococcus aureus (MRSA) is a significant public health concern because of its zoonotic potential and multidrug resistance. This review provides a comprehensive overview of the prevalence of MRSA in livestock and companion animals in Pakistan, emphasizing its molecular epidemiology and resistance mechanisms. MRSA has been detected in bovine mastitis, raw milk, meat, and companion animals, with alarming rates across species and regions. Risk factors include poor hygiene, inappropriate antibiotic use, and close human-animal interactions, all of which contribute to its spread at the animal-human-environment interface. Advanced diagnostic tools such as polymerase chain reaction and chromogenic media tests have facilitated rapid and accurate MRSA detection. The review also highlights alternative treatment strategies, including the use of medicinal plants, bacteriophage therapy, nanoparticles, and probiotics, as promising options to combat MRSA infections. Implementation of stringent hygiene measures, prudent antibiotic use, and further research into vaccines are critical for controlling MRSA in animals and safeguarding public health. Keywords: Methicillin-resistant Staphylococcus aureus, One Health, Antimicrobial resistance, Probiotic therapy, Public health. IntroductionStaphylococcus aureus (MRSA) is an opportunistic pathogen that inhabits the skin, mucosal membranes, and upper respiratory tracts of both animals and humans (Feng et al., 2008). In veterinary medicine, it is the leading causative agent of bovine mastitis in dairy cattle (Khairullah et al., 2022), resulting in huge economic losses worldwide (Li et al., 2018). Currently, most S. aureus strains classified as methicillin-resistant S. aureus (MRSA) exhibit resistance to penicillin derivatives (Grema et al., 2015). MRSA was first detected in the United Kingdom in 1961 (Khanal et al., 2022). The mecA gene, which encodes penicillin-binding protein 2a (PBP2a), is the gold standard for detecting MRSA (Javed, 2021). Since 1972, when MRSA was first isolated from bovine mastitis cases in dairy cattle in Belgium, multiple studies have verified the zoonotic transmission of MRSA from livestock to humans; after that, it was also recognized as livestock-associated MRSA (Khairullah et al., 2020). The World Health Organization (WHO) has classified methicillin-resistant MRSA as a "High Priority" pathogen on its Global Priority List of Antibiotic-Resistant Bacteria, underscoring the critical need for research and development of new therapeutics (Shrestha et al., 2021). MRSA has several virulence factors, including coa, spa, and Panton-Valentine leucocidin (pvl) genes (Sadiq et al., 2020). The pvl gene encodes the most potent staphylococcal leukotoxin, which can inhibit bovine neutrophils, destroy polymorphonuclear cells, and increase host resistance and pathogenicity (Shahid et al., 2023). The mecA gene in the chromosomal cassette of foreign DNA sections encodes the PBP2a protein, the primary mode of MRSA transmission (Ijaz et al., 2024). Enzymes such as proteases, lipases, elastases, and slime factors help host tissues grow and propagate to new sites (Gordon and Lowry, 2008). Until the mid-1940s, penicillin was referred to as the most effective antibiotic against many staphylococcal infections; not long after that, S. aureus strains started developing resistance to penicillin (Khan et al., 2014) and beta-lactamase enzyme production. The emergence of methicillin resistance, as depicted by the acquisition of the mecA gene, has led to a low affinity of bacteria for all β lactam drugs (Hamid et al., 2017). Furthermore, studies have detected MRSA in bovine mastitis (Imran et al., 2023), raw milk (Effendi and Harijani, 2017; Effendi et al., 2019), commercial raw meat, including chicken, beef, and mutton (Sadiq et al., 2020), and dairy farms (Tyasningsih et al., 2019; Khairullah et al., 2022; Imran et al., 2023). MRSA prevalence in animals is reported to be higher in many Asian countries compared to other continents, a trend often attributed to factors such as high-density livestock farming, variable antimicrobial stewardship practices, and the intensive use of antibiotics in animal production (Zaatout and Hezil, 2022), as well as their use in companion animals (Khairullah et al., 2023a). The polymerase chain reaction (PCR) is the "gold standard" for rapid and accurate screening tests for MRSA detection by identifying the mecA gene (Ijaz et al., 2023). The uncontrolled use of antibiotics in human and veterinary medicine is a significant contributor to the emergence and spread of MRSA infections, which limits therapeutic options (Khairullah et al., 2023b). The rise of AMR has increased the need for research into alternative antibiotics or treatment strategies (Jenkins et al., 2015). This review article aims to describe the molecular prevalence of MRSA in livestock and companion animals in Pakistan, developments in rapid diagnosis, and the exploration of various treatment options. Molecular prevalence of MRSA in animals in PakistanMRSA infection in large ruminants (cattle and buffaloes)There is still a paucity of information on studies in Pakistan that investigate the prevalence of MRSA in food-producing animals and their by-products. In fact, for some animals, data on MRSA investigation is nonexistent, especially with regard to pork meat from pigs, due to religious reasons, as it is considered a “taboo food” for Muslims. MRSA is considered as a contagious pathogen as it can cause subclinical and clinical mastitis in cattle and buffaloes. Interestingly, dairy farms have been reported to be important sources of MRSA spread, which have been linked to epidemics in both humans and animals (Tariq et al., 2023; Ijaz et al., 2023; Ijaz et al., 2024). A study was conducted in several Tehsils of Faisalabad District to determine the prevalence of MRSA in bovine milk. In this study, the prevalence of MRSA in cow and buffalo milk was 30% and 38%, respectively, whereas the overall prevalence of MRSA among analyzed samples was 34%. This study also indicated that MRSA was strongly associated with the spread of mastitis (Aqib et al., 2017). Another study from Islamabad reported 54% MRSA prevalence in cattle milk affected in subclinical mastitis (Khan, 2019). In a study conducted in the Punjab province of Pakistan, Haq et al. (2024) reported a prevalence of MRSA in raw milk from dairy cattle with subclinical mastitis of 23.40%. In different provinces of Pakistan, the prevalence of MRSA in cattle milk samples in Punjab and KPK was 58.09% and 81.81%, respectively (Lubna et al., 2023; Tariq et al., 2023). Milk samples from buffaloes in Punjab and KPK contained 18.79% and 19.6% MRSA prevalence, respectively (Javed et al., 2022; Javed et al., 2023). MRSA has consistently been noted to be a leading cause of wound infections (Khan et al., 2022); the carrier rate for MRSA in skin symptoms was 18.18%. The majority of reports from Pakistan detected MRSA through the targeting of mecA and PVL genes (Pérez-Roth et al., 2001). In Pakistan, ST22-T8934-MRSA-IVa has been isolated in cattle, and all identified MRSAs belong to sequence type ST22, which is related to the epidemic strain EMRSA-15, which is widely distributed in both animals and humans (Javed et al., 2022). MRSA infection in small ruminants (Sheep and Goat)In Pakistan, sheep and goats are valued for their milk and meat. In Pakistan, sheep and goats are culturally and economically significant, valued for both their milk and meat, which are widely consumed due to religious preferences and socioeconomic factors. This underscores their importance as a potential reservoir of MRSA within the local food chain (Yunita et al., 2020). In Pakistan, the prevalence of MRSA in goats with subclinical mastitis was 6.5%, and the isolates had 99% homogeneity with MRSA isolates from Turkey, India, and Japan (Altaf, 2020). Another study was conducted to investigate the prevalence of MRSA in goat milk samples, which revealed 35.92%. In addition, a low range of homology at the goat-human interface was observed, as well as a strong evolutionary relationship between the study isolate and the MRSA isolate from cattle and buffalo (Javed et al., 2023). A study on lactating ewes in the district of Muzaffargarh, Pakistan, found a 21.15% prevalence of MRSA by using a genotypic test, and isolates’ genomes showed very close resemblance to previously reported strains from Pakistan, China, and Myanmar. Isolates were also noted to be resistant to penicillin, ceftriaxone, and sulfamethoxazole-trimethoprim (Sabir et al., 2024). Currently, establishing the prevalence of MRSA in sheep and goats in Pakistan is challenging due to the paucity of data; however, existing evidence suggests a high prevalence of MRSA, as cited in Table 1. Table 1. Presence of MRSA in animals in different cities of Pakistan.
MRSA in the foodThe food industry in Pakistan is growing, but data on MRSA are insufficient. Only 1 study focusing on dairy products has been reported. In this study, the prevalence of MRSA in milk formula was 20%. Additionally, the molecularly typed MRSA strains belonged to the SCCmec type II group, a commonly isolated pathogenic strain in nosocomial settings (Mirani et al., 2017). Syed et al. (2018) investigated the prevalence of MRSA in table eggs in Haripur City, Pakistan, and reported that 86.8% of the egg samples were positive for MRSA. All the isolated strains in egg samples were detected by targeting the PVL gene, which empowers MRSA strains to cause skin and soft tissue infections and necrotizing pneumonia. The majority of MRSA isolates in the United States are from the 300 and Bengal Bay lineages, with 93.9% belonging to ST772, which was previously isolated from a human patient in Malakand and Rawalpindi (Madzgalla et al., 2016). Syed et al. (2021) observed a 6% prevalence of MRSA in eggs by spa typing of MRSA strains. Isolates were noted to belong to the Bengal Bay lineage, which is widespread in human infections all over the world and in eggs, suggesting that MRSA infection spread from humans to poultry via contamination. MRSA has been isolated from chicken, beef, and mutton meat (Table 2) in Rawalpindi, Islamabad, and Karachi, Pakistan. Another study on meat corporations in Rawalpindi and Islamabad, Pakistan, reported an overall MRSA prevalence of 79%. In this study, the highest MRSA prevalence was observed in chicken (77%), followed by beef (63%) and mutton (50%). The higher frequency of MRSA in meat samples in Pakistan necessitates the implementation of suitable hygienic measures and procedures during meat slaughtering, transportation, and marketing (Sadiq et al., 2020). Table 2. Prevalence and types of MRSA in Pakistani food products.
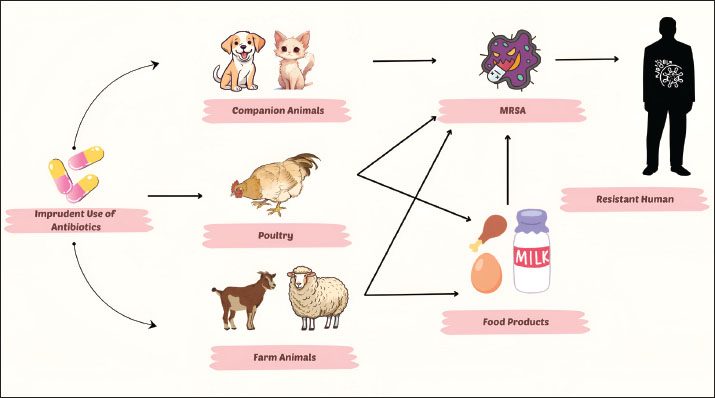
MRSA infection in companion animals (dogs and cats)In the current era, companion animals have been imported from various countries for various purposes (e.g., shows, sports, pets, and breeding). Public health awareness has led veterinary researchers in Pakistan to improve the diagnosis and treatment of animal diseases (Decline et al., 2020). Pakistan does not have enough data on MRSA colonization in companion animals. Few studies have reported that the prevalence of MRSA is 33.91% in dogs, 30.43% in cats, 25% in pet owners, and 50% in the environment (Shoaib et al., 2020a). Another study (Shoaib et al., 2022b) reported a 33.33% MRSA prevalence in cats, 46% in the environment, and 28% among cat owners. Sequencing of isolates further revealed that infection is transmitted between the environment and cats. The study concluded that MRSA spreads at the animal-human-environment interface in Pakistan. MRSA in the equineOnly one study from the Layyah district in Pakistan has proven the presence of MRSA in equines, with a prevalence of 29.63% (Rasheed et al., 2023). Ascertaining the exact prevalence of MRSA in horses in Pakistan is difficult due to differences in detection methods; only one study validated MRSA using PCR, whereas other studies evaluated the phenotypical frequency of MRSA in horses (Waqar et al., 2019). Risk factors for MRSA infectionThe misuse/abuse of antibiotics for treating dairy cattle is the primary cause of the high MRSA contamination rate at dairy farms. During the milking procedure, contamination can occur from the farmer’s hands as well as from the milk taken from the udder. The spread of these pathogenic bacteria in dairy farms can be minimized through good hygiene practices, especially during milking (Khairullah et al., 2022). Mastitis is a major source of MRSA contamination and transmission because it contaminates milk and milk products. MRSA might be transmitted through unhygienic milker hands (Javed, 2021), poor pre- and post-milking techniques, sharing udder cloths between animals, improper glove use during milking (Ijaz et al., 2023), environmental contamination, inappropriate antibiotic use, and manual hand milking rather than machine milking. The floor type and housing system in cattle and buffaloes are the primary factors that may contribute to the high incidence of MRSA (Muzammil et al., 2021; Ijaz et al., 2024). Interestingly, teat dipping has been reported to be very useful in reducing the transmission of MRSA (Zaatout and Hezil, 2022). MRSA-infected horses pose a risk to humans who work closely with them. In Pakistan, MRSA was discovered in nasal swabs of horses, indicating a possible reservoir for human infection (Zaheer et al., 2017). A study identified cats aged >1 year and in poor physical condition, as well as households with >4 dogs, as significant risk factors associated with MRSA carriage (Loeffler et al., 2011). Older humans have also been found to have a higher risk of contracting MRSA infections. The environment interface and non-sterilized tools have also been identified as potential sources of MRSA (Shoaib, 2020). This complex transmission cycle at the animal-human-environment interface is summarized in Figure 1.
Fig. 1. Transmission of MRSA between animals, their products, and humans. Pet owners should be instructed to maintain good hygienic practices with their pets and avoid the misuse/abuse of antibiotics to prevent the development of staphylococcal resistance. Veterinary professionals should be aware of MRSA infections as they are also at risk, especially with frequent contact with animals. Furthermore, more research should be conducted on the prevalence of MRSA in healthy dogs and cats. Rapid diagnostic approaches for MRSA infectionThe identification of MRSA strains using traditional cultural procedures and susceptibility testing methods is time-consuming, taking between 48 and 72 hours to reach a reasonable result. Advanced molecular and non-molecular testing methods have helped in reducing the time required to detect MRSA (Palavecino, 2020). Chromogenic media testsIt involves the use of chromogenic agar to identify bacteria of interest through their appearance. In chromogenic media, antibiotics are employed to allow only resistant bacteria to grow; MRSA can be discovered within 20–26 hours. A series of trials revealed a test specificity of 97% at 18–24 hours and fell to 94% after 48 hours. While the tests had greater sensitivity and specificity than the traditional culture techniques, a key limitation of chromogenic media is its inability to provide a viable isolate for downstream applications, such as antimicrobial susceptibility testing or whole-genome sequencing, which are achievable with traditional culture (Imran et al., 2023). It is simple to use, takes less time, and is cost effective, especially in low-resource settings (Ahmed et al., 2022). It offers a cost-effective alternative where molecular tests are unavailable, especially in identifying the preliminary characteristics of S. aureus (Shevchenko et al., 2023). Polymerase chain reactionAccurate and early diagnosis of MRSA using traditional methods is difficult; a specific identification procedure is required to control MRSA endemicity (Gorgun et al., 2021). The gold standard approach for detecting the mecA gene in MRSA is PCR (Madhavan et al., 2021). PCR is a common molecular test used to detect MRSA [48]. Tests can be performed directly on samples such as milk, nose swabs, or wound swabs suspected of possible MRSA contamination, with results available within 1–3 hours (Badua et al., 2020). PCR test is faster than other traditional tests, but requires time to transfer samples, run the test, and interpret the results (Sheet et al., 2021). A primary limitation of PCR is that it detects the presence of the mecA resistance gene but does not confirm the bacterium’s viability or provide an isolate for subsequent antimicrobial susceptibility testing. Furthermore, PCR requires specialized equipment and trained personnel and has higher per-test costs (Elbehiry et al., 2025). PCR takes much less time than the chromogenic media test. The sensitivity and specificity of PCR are 97.9% and 94.4%, respectively (Clay et al., 2021). PCR has significantly higher sensitivity than the chromogenic media test (Hiermandi et al., 2023). Immunochromatographic testThe latex agglutination test is an example of the immunochromatographic test. This test employs monoclonal antibodies against a protein PBP2a encoded by the mecA gene (Kolesnik-Goldmann et al., 2021). The antibodies first sensitize the latex particles, which then react with PBP2a and clump together, resulting in agglutination. The sensitivity and specificity of the test are 97% and 100%, respectively. The test takes 6 minutes to complete, and the results appear as colored lines on test strips (Tominaga and Ishii, 2020). A comparative summary of the diagnostic approachesAll diagnostic approaches have specific pros and cons that apply to unique situations. Traditional culture and AST cannot be substituted with viable isolates to obtain complete characterization with full susceptibility profiles and genomic studies, which are time-consuming (48–72 hours) (Palavecino, 2020). The chromogenic media provide a more considerable time-saving (20–26 hours) and can be afforded in low-resource environments, but do not provide isolates to perform any additional testing (Ahmed et al., 2022; Imran et al., 2023). PCR is the most specific and fastest to detect the mecA gene (1–3 hours) as well as the molecular gold standard, but it is also equipment intensive, more expensive, and does not provide a viable isolate to further confirm the phenotype (Clay et al., 2021; Gorgun et al., 2021). Immunochromatographic systems (e.g., latex agglutination of PBP2a) are fast (minutes), can be performed at the point of care, and are somewhat less sensitive than molecular systems (Tominaga and Ishii, 2020; Kolesnik-Goldmann et al., 2021). The test selection ultimately comes down to the requirements of speed, cost, isolate recovery, and available laboratory facilities, especially in resource-variable locations such as Pakistan. Treatment options for MRSA infectionNatural extracts have anti-MRSA potentialA study on medicinal plants found that they can be an effective source of antimicrobials for various diseases. The aqueous extracts of medicinal plants, Allium sativum and Allium cepa, have strong anti-MRSA properties (Hussain et al., 2023). According to Naeem et al. (2021), the chloroform extract of Azaderachta indica has been exhibited to be effective in skin treatments. Phytochemicals found in medicinal plants have antibacterial activity (Nandhini et al., 2022). The following are their multiple mechanisms. The multiple actions are:
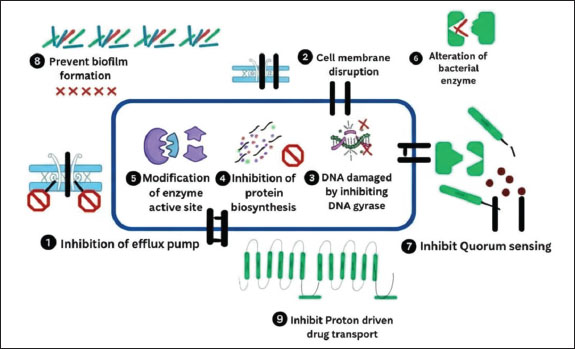
According to a study by Ali et al. (2022), EGEO may be an alternate therapeutic option for the treatment of MRSA infections. Rasheed et al. (2024) suggested that some fruit juices, Citrus reticulate and Punica granatum, have anti-MRSA properties. The antibacterial mechanisms of these phytochemicals are visually summarized in Figure 2.
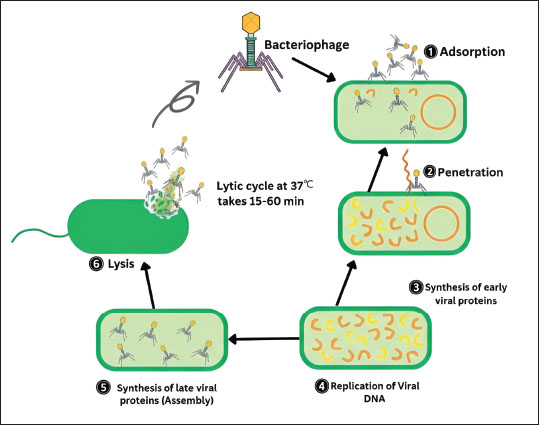
Fig. 2. Antibacterial mechanisms of various phytochemicals against methicillin-resistant Staphylococcus aureus stain. Natural drugsSeveral natural agents show powerful bactericidal properties against drug-resistant South aureus. The use of herbal medications to treat infections has increased globally. Curcuma longa (Turmeric) (Fatima et al., 2023) and Piper nigrum (black pepper) (Khan et al., 2024), Zingiber zerumbet (ginger) (Ramzan et al., 2022), and A. sativum (garlic) (Nayab et al., 2023) have potential antibacterial activity against MDR S. aureus. Apple cider vinegar, created from cider or apples, is known for its antimicrobial properties and is considered the mother of all vinegars. Acetic acid, a main element in vinegar, can kill bacteria or prevent them from multiplying (Saqib et al., 2021). Honey (predominantly from Apis mellifera) has been used as an Ayurvedic and Yunani medicine for hundreds of years (Tanuğur Samanci et al., 2024). Today, scientists recognize honey as a beneficial product for treating a variety of ailments. Its therapeutic properties are recognized due to its antioxidants, which include phenolic acid and flavonoids. Talha et al. (2022) also found the antibacterial activity of honey. Plant phytochemical compounds exhibit antibacterial activity because flavonoids form a complex with the bacterial cell wall, extracellular substrate, and soluble proteins, whereas tannins inactivate enclosed proteins and enzymes, causing bacterial lysis. Bacteriophage therapyIn 1915 and 1917, Twort and D'Herelle identified virus particles in 1915 and 1917, naming them bacteriophages (Duckworth, 1976). In 1915, Frederick Twort first described its antibacterial action against S. aureus in 1915. The National Institute of Bacteriophages in Georgia was founded in 1923, and biological control of infectious diseases has been ongoing since then. Some bacteriophages are very specific against specific bacterial species (narrow host range), whereas others have a broad host range and can infect several species across bacterial genera. Bacteriophages can be either lytic (virulent) or lysogenic (temperate). Lytic phages infect bacterial cells and hijack their machinery, causing their phage DNA and protein to rapidly multiply and cause cell lysis and death in a short time, releasing hundreds of infectious virus particles that further infect other host cells (productive infections). Lysogenic phages can integrate their genome into bacterial chromosomes after infection and remain latent (prophage) until they become reactivated into the lytic scycle, where they can transfer genes from one bacterial strain to another (Bianchessi et al., 2024). The general lytic and lysogenic cycles of the bacteriophages are illustrated in Figure 3.
Fig. 3. General antibacterial mechanism of bacteriophages against methicillin-resistant Staphylococcus aureus stain. Advantages of phage therapy
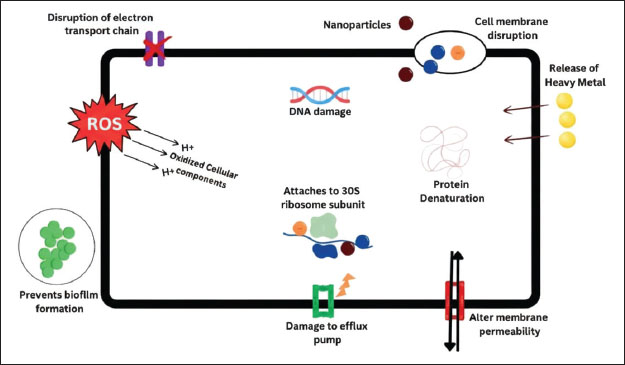
Nanoparticles for the treatment of MRSAThese are considered the wonders of modern medicine (Sungkaworn and Triampo, 2007). Its antibacterial activity is affected by its stability in infected cells and predictability of target site delivery (Azam et al., 2023). Nanoparticles are tiny, have a vast surface area, and can interact with specific receptors; their desirable qualities include biodegradability, biocompatibility, non-immunogenicity, and stability in physiological fluids. Nanoparticles can deliver high doses of antibiotics to the site of infection while reducing toxicity to nontarget cells (Andrade et al., 2022). Silver nitrate, zinc oxide, platinum, aluminum oxide, titanium dioxide, gold, magnesium oxide, iron oxide, and sodium alginate are the most commonly used metal oxide nanomaterials (Akhtar et al., 2019; Spirescu et al., 2021; kov et al., 2022; Mendes et al., 2022). Figure 4 depicts the multifaceted antibacterial mechanisms of nanoparticles.
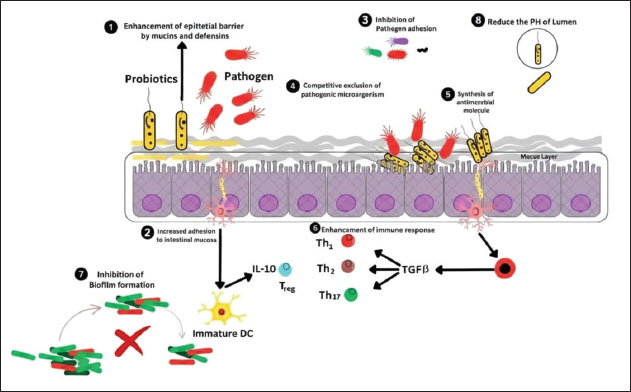
Fig. 4. Schematic representation of the proposed antibacterial mechanisms of various nanoparticles against methicillin-resistant Staphylococcus aureus. ProbioticsProbiotics are live organisms (e.g., yeast or bacteria) that colonize the host and exert positive effects. Lactobacillus, Lactococcus, Streptococcus, and Enterococcus are lactic acid bacteria species that are well-known as probiotics (Soccol et al., 2014). These bacteria possess properties, such as enzymatic activity and acidity, that enable them to withstand harsh conditions in the host organism. They can colonize the host and improve health by regulating the microbiota and performing biological functions (Pereira et al., 2019). In 2002, the Food and Agriculture Organization and the World Health Organization published the "Guidelines for the evaluation of probiotics in food" in 2002, which established probiotic safety and efficacy standards (Araya et al., 2002). The safety of a probiotic strain is determined by its origin, absence of interaction with pathogenic cultures, and antibiotic resistance profile (Markowiak and Śliżewska, 2017). Lactobacillus is the oldest known probiotic. It includes 183 recognized species commonly used in industrial food processing (Fröhlich et al., 2009). The main biological mechanisms of action of probiotics include increased epithelial barrier, increased adhesion to the intestinal mucosa, inhibition of microbial adhesion and competitive exclusion of pathogenic microorganisms, production of antimicrobial substances, and immune system modulation (Bermudez-Brito et al., 2012). According to evidence, probiotic strains can reduce side effects, improve antibiotic function, and enhance mucosal immunity (Jalalifar et al., 2022). Probiotics have anti-MRSA properties (antibacterial, antibiofilm, antivirulence, antidrug resistance, coaggregation, and antiquorum sensing) by producing numerous antagonistic compounds such as organic acid, hydrogen peroxide, biosurfactants, and bacteriocin-like inhibitory substances (Nataraj et al., 2021). These complex mechanisms are outlined in Figure 5.
Fig. 5. Antibacterial mechanism of various probiotics against methicillin-resistant Staphylococcus aureus stain. MRSA controlOveruse of antibiotics in animals leads to the establishment of MRSA (Ramandinianto et al., 2020). According to public health concerns, MRSA infections require the collaboration of microbiologists, epidemiologists, veterinarians, and medical experts (Algammal et al., 2020). To tackle MRSA infections in animals, MRSA strains need to be investigated using PCR to identify the mecA gene (Vestergaard et al., 2019). Furthermore, environmentally friendly therapy regimens such as herbal medication, probiotics, and nanoparticles should be employed (Nandhini et al., 2022). Antimicrobial susceptibility testing should be used to determine the optimal treatment for MRSA infection, rather than the frequent and blind use of broad-spectrum antibiotics (Shoaib et al., 2023). To reduce the spread of MRSA in the community, veterinarians must take standard precautions such as good hygiene, infection control regimen, and environment disinfection. To reduce MRSA contamination in human food, good Hazard Analysis and Critical Control Point guidelines should be followed when handling and processing meat, milk, and dairy products. MRSA-infected animals must be isolated and treated, and MRSA-infected wounds must be cleaned and protected (Algammal et al., 2020). ConclusionThis review highlights the significant prevalence and multidrug resistance of methicillin-resistant MRSA in Pakistani livestock and companion animals. The findings underscore the zoonotic potential of MRSA, emphasizing its presence in bovine mastitis, raw milk, meat products, and companion animals. Key factors contributing to the emergence and spread of MRSA include poor hygiene practices, uncontrolled antibiotic use, and close human-animal interactions. The review also explores advanced diagnostic methods, including PCR and chromogenic media tests, and innovative treatment strategies, such as phytochemicals, bacteriophage therapy, nanoparticles, and probiotics, which offer promising alternatives to conventional antibiotics. AcknowledgmentsThis study received no funding from public agencies. Conflict of interestThe authors declare no conflicts of interest. FundingNone. Authors' contributionsAYK wrote the whole manuscript. AS and MHE supervised the study. MM added the references, edited and formatted the manuscript. All authors have read and approved the final version of the manuscript. Data availabilityAll data were provided in the manuscript. ReferencesAhmed, A., Ijaz, M., Khan, J.A. and Anjum, A.A. 2022. Molecular characterization and therapeutic insights into biofilm positive Staphylococcus aureus isolated from bovine subclinical mastitis. Pak. Vet. J. 42, 1–8. Akhtar, S., Shahzad, K., Mushtaq, S., Ali, I., Rafe, M.H. and Fazal-Ul-Karim, S.M. 2019. Antibacterial and antiviral potential of colloidal titanium dioxide (TiO2) nanoparticles suitable for biological applications. Mater. Res. Express. 6, 105409. Algammal, A.M., Hetta, H.F., Elkelish, A., Alkhalifah, D.H.H., Hozzein, W.N., Batiha, G.E.S., El Nahhas, N. and Mabrok, M.A. 2020. Methicillin-resistant Staphylococcus aureus (MRSA): one health perspective approach to the bacterium’s epidemiology, virulence factors, antibiotic resistance, and zoonotic impact. Infect. Drug. Resist. 13, 3255–3265. Ali, W., Zaman, S., Subhan, Z., Razaq, A., Nabi, M., Khattak, M., Naeem, N., Bangash, S.A. and Ullah, I. 2022. Methicillin-resistant Staphylococcus aureus (MRSA) epidemiology and antibiotics susceptibility profile isolated from different clinical samples in a tertiary care hospital. Pak. Biomed. J. 5, 108–112. Altaf, M. 2020. Molecular characterization of methicillin-resistant Staphylococcus aureus (MRSA) and associated risk factors with the occurrence of goat mastitis. Pak. Vet. J. 40, 19–24. Andrade, R.G.D., Ferreira, D., Veloso, S.R.S., Santos-Pereira, C., Castanheira, E.M.S., Côrte-Real, M. and Rodrigues, L.R. 2022. Synthesis and cytotoxicity assessment of citrate-coated calcium and manganese ferrite nanoparticles for magnetic hyperthermia. Pharmaceutics 14, 2694. Aqib, A.I., Ijaz, M., Anjum, A.A., Malik, M.A.R., Mehmood, K., Farooqi, S.H. and Hussain, K. 2017. Antibiotic susceptibilities and prevalence of methicillin-resistant Staphylococcus aureus (MRSA) isolated from bovine milk in Pakistan. Acta Trop. 176, 168–172. Araya, E., Hofner, P., Churchwell, E. and Kurtz, S. 2002. Arecibo observations of formaldehyde and radio recombination lines toward ultracompact H II regions. Astrophys. J. Suppl. Ser. 138, 63–72. Azam, M., Qureshi, T.M., Hussain, S., Aqib, A.I., Khan, S.R., Akram, K., Ijaz, M., Murtaza, M., Muneer, A. and Mahmood, S. 2023. Resistance modulation of Staphylococcus aureus isolates of dairy cattle through metallic oxide nanoparticles. Pak. J. Zool. 55, 2415–2423. Badua, A., Boonyayatra, S., Awaiwanont, N., Gaban, P. and Mingala, C. 2020. Antibiotic resistance and genotyping of mecA-positive methicillin-resistant Staphylococcus aureus (MRSA) from milk and nasal carriage of dairy water buffaloes (Bubalus bubalis) in the Philippines. J. Adv. Vet. Anim. Res. 7, 397–407. Bermudez-Brito, M., Plaza-Díaz, J., Muñoz-Quezada, S., Gómez-Llorente, C. and Gil, A. 2012. Probiotic mechanisms of action. Ann. Nutr. Metab. 61, 160–174. Bianchessi, L., De Bernardi, G., Vigorelli, M., Dall’Ara, P. and Turin, L. 2024. Bacteriophage therapy in companion and farm animals. Antibiotics 13, 294. Clay, T.B., Orwig, K.W., Stevens, R.A., Davis, E.P., Jennings, T.M., Long, T.E., Riley, B.L. and Hambuchen, M.D. 2021. Correlation of MRSA polymerase chain reaction (PCR) wound swab testing and wound cultures in skin and soft tissue infections. Diagnostic Microbiol. Infect. Dis. 100, 115389. Decline, V., Effendi, M.H., Rahmaniar, R.P., Yanestria, S.M. and Harijani, N. 2020. Profile of antibiotic-resistant and presence of methicillin-resistant Staphylococcus aureus from nasal swab of dogs from several animal clinics in Surabaya, Indonesia. Int. J. One Health 6(1), 90–94. Duckworth, D.H. 1976. Who discovered bacteriophage?. Bacteriol. Rev. 40, 793–802. Effendi, M.H. and Harijani, N. 2017. Cases of methicillin-resistant Staphylococcus aureus (MRSA) from raw milk in East Java Indonesia. Glob. Vet. 19, 500–503. Effendi, M.H., Hisyam, M.A.M., Hastutiek, P. and Tyasningsih, W. 2019. Detection of coagulase gene in Staphylococcus aureus from several dairy farms in East Java, Indonesia, by polymerase chain reaction. Vet. World. 12(1), 68–71. Elbehiry, A., Marzouk, E., Abalkhail, A., Abdelsalam, M.H., Mostafa, M.E., Alasiri, M., Ibrahem, M., Ellethy, A.T., Almuzaini, A., Aljarallah, S.N. and Abu-Okail, A. 2025. Detection of antimicrobial resistance via state-of-the-art technologies versus conventional methods. Front. Microbiol. 16, 1549044. Fatima, H., Ibrahim, A., Hamdani, S.D.A., Rajput, T.A., Noor, A., Gul, A. and Babar, M.M. 2023. Natural product formulations to overcome poor ADMET properties. In Drug discovery and design using natural products. Cham, Switzerland: Springer, pp: 435–52; doi:10.1007/978-3-031-35205-8_15 Feng, Y., Chen, C.J., Su, L.H., Hu, S., Yu, J. and Chiu, C.H. 2008. Evolution and pathogenesis of Staphylococcus aureus: lessons learned from genotyping and comparative genomics. FEMS Microbiol. Rev. 32, 23–37. Fröhlich, J., König, H. and Claus, H. 2009. Rapid detection and identification with molecular methods. In Biology of microorganisms on grapes, in must and in wine. Berlin, Germany: Springer, pp: 429–449. Gordon, R.J. and Lowy, F.D. 2008. Pathogenesis of methicillin-resistant Staphylococcus aureus infection. Clin. Infect. Dis. 46, S350–S359. Gorgun, S., Isler, H. and Turgut, M.C. 2021. Comparison of rapid and conventional methods for investigating mecA presence in Staphylococcus species. Pak. J. Med. Sci. 37, 1467–1472. Grema, H.A., Geidam, Y.A., Gadzama, G.B., Ameh, J.A. and Suleiman, A. 2015. Methicillin resistant Staphylococcus aureus (MRSA): a review. Adv. Anim. Vet. Sci. 3, 79–98. Hamid, S., Bhat, M.A., Mir, I.A., Taku, A., Badroo, G.A., Nazki, S. and Malik, A. 2017. Phenotypic and genotypic characterization of methicillin-resistant Staphylococcus aureus from bovine mastitis. Vet. World. 10, 363–367. Haq, I.U., Kamal, M., Swelum, A.A., Khan, S., Ríos-Escalante, P.R.D.L. and Usman, T. 2024. Alarming multidrug resistance in Staphylococcus aureus isolated from raw milk of cows with subclinical mastitis: antibiotic resistance patterns and occurrence of selected resistance genes. PLos One. 19, 301200. Hiermandi, N., Foster, C., Purnell, K., Dunn, J., Campbell, J. and Marquez, L. 2023. MRSA PCR improves sensitivity of detection of colonization in neonates. Antimicrob. Steward. Healthc. Epidemiol. 3, s58–s59. Hussain, S.B., Rafi, H., Aslam, A., Noor, S. and Zubair, M. 2023. Genetic analysis of methicillin-resistant Staphylococcus aureus (MRSA) isolated from the population of Southern Punjab, Pakistan. Asian J. Biol. 19, 73–82. Ijaz, M., Ghumman, N.Z., Javed, M.U., Ahmed, A., Rasheed, H., Anwaar, F., Sabir, M.J. and Muzammil, I. 2024. Genotypic insights to Panton–Valentine leukocidin positive methicillin-resistant Staphylococcus aureus isolated from cattle mastitis. J. Dairy. Res. 91, 1–3. Ijaz, M., Javed, M.U., Ahmed, A., Rasheed, H., Shah, S.F.A. and Ali, M. 2023. Evidence-based identification and characterization of methicillin-resistant Staphylococcus aureus isolated from subclinical mastitis in dairy buffaloes of Pakistan. Iran. J. Vet. Res. 24, 165–171. Imran, H., Khan, Z., Saleem, F., Gull, S. and Tahir, A. 2023. The growing threat of antibiotic resistance in wound infections: evidence from tertiary care in Pakistan. Arch. Biol. Sci. 75, 21–31. Jalalifar, S., Mirzaei, R., Motallebirad, T., Razavi, S. and Talebi, M. 2022. The emerging role of probiotics and their derivatives against biofilm-producing MRSA: a scoping review. Biomed. Res. Int. 2022, 4959487. Javed, M.U. 2021. Frequency and antimicrobial susceptibility of methicillin and vancomycin-resistant Staphylococcus aureus from bovine milk. Pak. Vet. J. 41(1), 1–7. Javed, M.U., Ijaz, M., Durrani, A.Z. and Ali, M.M. 2023. On-farm epidemiology, virulence profiling, and molecular characterization of methicillin-resistant Staphylococcus aureus at goat farms. Microb. Pathog. 185, 106456. Javed, S., Mcclure, J., Syed, M.A., Obasuyi, O., Ali, S., Tabassum, S., Ejaz, M. and Zhang, K. 2022. Epidemiology and molecular characterization of Staphylococcus aureus causing bovine mastitis in water buffaloes from the Hazara division of Khyber Pakhtunkhwa Pakistan. PLos One 17, 268152. Jenkins, B.N., Vincent, N. and Fortier, M.A. 2015. Differences in referral and use of complementary and alternative medicine between pediatric providers and patients. Complement. Ther. Med. 23, 462–468. Khairullah, A., Sudjarwo, S., Effendi, M., Ramandinianto, S., Gelolodo, M., Widodo, A., Riwu, K. and Kurniawati, D. 2023a. Pet animals as reservoirs for spreading methicillin-resistant Staphylococcus aureus to human health. J. Adv. Vet. Anim. Res. 10(1), 1–13. Khairullah, A.R., Kurniawan, S.C., Silaen, O.S.M., Effendi, M.H., Sudjarwo, S.A., Ramandinianto, S.C., Gololodo, M.A., Widodo, A., Riwu, K.H.P., Kurniawati, D.A. and Rehman, S. 2023b. Methicillin-resistant Staphylococcus aureus (MRSA) isolation and mecA gene detection from milk and farmer hand swab in Tulungagung, Indonesia. Trop. Anim. Sci. J. 46, 231–238. Khairullah, A.R., Ramandinianto, S.C. and Effendi, M.H. 2020. A review of livestock-associated methicillin-resistant Staphylococcus aureus (LA-MRSA) on bovine mastitis. Syst. Rev. Pharm. 11, 172–183. Khan, A. 2019. Epidemiology and antimicrobial susceptibility of methicillin-resistant Staphylococcus aureus in cattle of Pothohar Region Pakistan. Pak. Vet. J. 39, 353–356. Khan, K.M., Nadeem, M.F., Mannan, A., Chohan, T.A., Islam, M., Ansari, S.A., Alkahtani, H.M., Ansari, I.A., Khurshid, U., Abidin, S.A.Z. and Shah, S.A.A. 2024. Biochemical, toxicological, and in silico aspects of Trillium govanianum Wall. ex D. Don (Trilliaceae): a rich source of natural bioactive compounds. Chem. Biodivers. 21, e202301375. Khan, M.A., Irfan, S., Zeeshan, M., Ahmed, I. and Zafar, A. 2022. Antimicrobial susceptibility pattern of daptomycin-sensitive MRSA (methicillin-resistant Staphylococcus aureus) from skin and soft tissue infections. Infect. Dis. J. Pak. 31, 67–71. Khan, R.A., Rahman, A.U., Ahmad, A., Jaseem, M., Jabbar, A., Khan, S.A., Iqbal, A., Ahmad, N., Wadood, A. and Rahman, T.U. 2014. Prevalence and antibiotic susceptibility profile of methicillin-resistant Staphylococcus aureus (MRSA) isolated from different clinical samples in district Peshawar. J. Appl. Environ. Biol. Sci. 4, 40–46. Khanal, S., Boonyayatra, S. and Awaiwanont, N. 2022. Prevalence of methicillin-resistant Staphylococcus aureus in dairy farms: a systematic review and meta-analysis. Front. Vet. Sci. 9, 947154. Kolesnik-Goldmann, N., Bodendoerfer, E., Röthlin, K., Herren, S., Imkamp, F., Marchesi, M. and Mancini, S. 2021. Rapid detection of PBP2a in Staphylococci from shortly incubated subcultures of positive blood cultures by an immunochromatographic assay. Microbiol. Spectr. 9, 550. Li, S.M., Zhou, Y.F., Li, L., Fang, L.X., Duan, J.H., Liu, F.R., Liang, H.Q., Wu, Y.T., Gu, W.Q., Liao, X.P. and Sun, J. 2018. Characterization of the multi-drug resistance gene cfr in methicillin-resistant Staphylococcus aureus (MRSA) strains isolated from animals and humans in China. Front. Microbiol. 9, 2925. Loeffler, A., Pfeiffer, D.U., Lindsay, J.A., Magalhães, R.S. and Lloyd, D.H. 2011. Prevalence of and risk factors for MRSA carriage in companion animals: a survey of dogs, cats and horses. Epidemiol. Infect. 139(7), 1019–1028. Lubna., Hussain, T., Shami, A., Rafiq, N., Khan, S., Kabir, M., Khan, N.U., Khattak, I., Kamal, M. and Usman, T. 2023. Antimicrobial usage and detection of multidrug-resistant Staphylococcus aureus: methicillin- and tetracycline-resistant strains in raw milk of lactating dairy cattle. Antibiotics 12, 673. Madhavan, A., Sachu, A., Balakrishnan, A., Vasudevan, A., Balakrishnan, S. and Vasudevapanicker, J. 2021. Comparison of PCR and phenotypic methods for the detection of methicillin-resistant Staphylococcus aureus. Iran. J. Microbiol. 13, 31–38. Madzgalla, S., Syed, M.A., Khan, M.A., Rehman, S.S., Müller, E., Reissig, A., Ehricht, R. and Monecke, S. 2016. Molecular characterization of Staphylococcus aureus isolates causing skin and soft tissue infections in patients from Malakand Pakistan. Eur. J. Clin. Microbiol. Infect. Dis. 35, 1541–1547. Markowiak, P. and Śliżewska, K. 2017. Effects of probiotics, prebiotics, and synbiotics on human health. Nutrients 9, 1021. Mendes, C., Thirupathi, A., Corrêa, M.E.A.B., Gu, Y. and Silveira, P.C.L. 2022. The use of metallic nanoparticles in wound healing: new perspectives. Int. J. Mol. Sci. 23, 15376. Mirani, Z.A., Aqeel, A. and Naz, S. 2017. Prevalence of Staphylococci in commercially processed food products in Karachi-Pakistan. J. Microbiol. Infect. Dis. 7, 83–87. Muzammil, I., Saleem, M.I., Aqib, A.I., Ashar, A., Mahfooz, S.A., Rahman, S.U., Shoaib, M., Naseer, M.A., Sohrani, I.K., Ahmad, J., Saqi, R., Lodhi, F.L. and Tanveer, Q. 2021. Emergence of pathogenic strains of Staphylococcus aureus in goat milk and their comparative response to antibiotics. Pak. J. Zool. 53, 1611–1619. Naeem, S., Siddique, A.B., Zahoor, M.K., Muzammil, S., Nawaz, Z., Waseem, M., Yasmin, A. and Asif Zahoor, M. 2021. In vitro efficacy of Azadirachta indica leaf extract against methicillin-resistant Staphylococci isolated from skin infection. Pak. J. Pharm. Sci. 34, 2303–2308. Nandhini, P., Kumar, P., Mickymaray, S., Alothaim, A.S., Somasundaram, J. and Rajan, M. 2022. Recent developments in methicillin-resistant Staphylococcus aureus (MRSA) treatment: a review. Antibiotics 11, 606. Nataraj, B.H., Ramesh, C. and Mallappa, R.H. 2021. Characterization of biosurfactants derived from probiotic lactic acid bacteria against methicillin-resistant and sensitive Staphylococcus aureus isolates. LWT-Food. Sci. Technol. 151, 112195. Nayab, A., Khan, F., Marrium, F., Waqas, M., Bashir, A., Abbas, A., Yousaf, N., Shaheen, S. and Perveen, F. 2023. Recent advances, innovative aspects, botanical description of therapeutic medicinal plants, and different applications. Haya Saudi J. Life Sci. 8, 44–50. Palavecino, E.L. 2020. Rapid methods for detection of MRSA in clinical specimens. In: Methicillin-resistant Staphylococcus aureus (MRSA) protocols: cutting-edge technologies and advancements. New York, NY: Springer, pp: 29–45; doi: 10.1007/978-1-62703-664-1_3. Pereira, R., Bortoluzzi, C., Durrer, A., Fagundes, N.S., Pedroso, A.A., Rafael, J.M., Perim, J.E.D.L., Zavarize, K.C., Napty, G.S., Andreote, F.D. and Costa, D.P. 2019. Performance and intestinal microbiota of chickens receiving probiotics in the feed and submitted to antibiotic therapy. J. Anim. Physiol. Anim. Nutr. 103, 72–86. Pérez-Roth, E., Claverie-Martı́n, F., Villar, J. and Méndez-Álvarez, S. 2001. Multiplex PCR for simultaneous identification of Staphylococcus aureus and detection of methicillin and mupirocin resistance. J. Clin. Microbiol. 39, 4037–4041. Khairullah, A.R., Rehman, S., Sudjarwo, S.A., Effendi, M.H., Ramandinianto, S.C., Gololodo, M.A., Widodo, A., Riwu, K.H.P. and Kurniawati, D.A. 2022. Detection of mecA gene and methicillin-resistant Staphylococcus aureus (MRSA) isolated from milk and risk factors from farms in Probolinggo Indonesia. F1000Research 11, 1234. Ramandinianto, S.C., Khairullah, A.R., Effendi, M.H. and Hestiana, E.P. 2020. Profile of multidrug resistance (MDR) and methicillin-resistant Staphylococcus aureus (MRSA) on dairy farms in East Java Province, Indonesia. Indian J. Forensic Med. Toxicol. 14, 3439–3445. Ramzan, M., Karobari, M.I., Heboyan, A., Mohamed, R.N., Mustafa, M., Basheer, S.N., Desai, V., Batool, S., Ahmed, N. and Zeshan, B. 2022. Synthesis of silver nanoparticles from extracts of wild ginger (Zingiber zerumbet) with antibacterial activity against selective multidrug-resistant oral bacteria. Molecules 27, 2007. Rasheed, H., Ijaz, M., Muzammil, I., Ahmed, A., Anwaar, F., Javed, M.U., Ghumman, N.Z. and Raza, A. 2023. Molecular evidence of β-lactam resistant Staphylococcus aureus in equids with respiratory tract infections: frequency and resistance modulation strategy. Acta. Trop. 245, 106967. Rasheed, H.I., Ali, A., Saeed, M.H., Aqsa, N., Sultan, S. and Riaz, S. 2024. Evaluation of antibacterial activity of fruit juices on biofilm-producing methicillin-resistant Staphylococcus aureus. Lahore Garrison Univ. J. Life Sci. 8, 49–64. Sabir, M.J., Ijaz, M., Ahmed, A., Rasheed, H., Javed, M.U. and Anwaar, F. 2024. First report on genotypic estimation of MRSA load in udder of nomadic sheep flocks affected with subclinical mastitis in Pakistan. Res. Vet. Sci. 166, 105107. Sabir, R., Alvi, S.F.D., Fawwad, A. and Basit, A. 2014. Antibiogram of Pseudomonas aeruginosa and methicillin-resistant Staphylococcus aureus in patients with diabetes. Pak. J. Med. Sci. 30, 814–818. Sadiq, M., Samad, M., Saddam, N., Basharat, B., Ali, S., Roohullah, Z., Saad, Z., Khan, A.N., Ahmad, Y. and Khan, A. 2020. Methicillin-resistant Staphylococcus aureus (MRSA) in slaughter houses and meat shops in capital territory of Pakistan during 2018–2019. Front. Microbiol. 11, 577707. Saqib, U., Khan, M.A., Alagumuthu, M., Parihar, S.P. and Baig, M.S. 2021. Natural compounds as antiatherogenic agents. Cell. Mol. Biol. 67, 177–188. Shahid, M., Hussain, R., Nawaz, Z., Aslam, B., Ahmad, M.Z., Siddique, A.B., Ahsan, H., Fatima, A., Khan, I., Mustafa, B., Iqbal, R., Al Syaad, K.M. and Shami, A. 2023. Occurrence of virulence genes among methicillin-resistant Staphylococcus aureus isolated from subclinical bovine mastitis. ACS Omega 8, 38111–38117. Sheet, O.H., Hussein, S.A. and Al-Chalaby, A.Y. 2021. Detection of methicillin-resistant Staphylococcus aureus from broiler carcasses in Mosul city. Iraqi J. Vet. Sci. 35, 489–493. Shevchenko, M., Andriichuk, A., Bilyk, S., Dovhal, O., Mazur, T. and Tsarenko, T. 2023. Biofilm forming ability of coagulase-positive Staphylococci isolated from animals in Ukraine. Regul. Mech. Biosyst. 14, 576–580. Shoaib, M. 2020. Diversified epidemiological pattern and antibiogram of mecA gene in Staphylococcus aureus isolates of pets, pet owners, and environment. Pak. Vet. J. 40, 331–336. Shoaib, M., Aqib, A.I., Ali, M.M., Ijaz, M., Sattar, H., Ghaffar, A., Sajid Hasni, M., Bhutta, Z.A., Ashfaq, K., Kulyar, M.F.E.A. and Pu, W. 2022. Tracking infection and genetic divergence of methicillin-resistant Staphylococcus aureus at pets, pet owners, and environment interface. Front. Vet. Sci. 9, 900480. Shoaib, M., Aqib, A.I., Muzammil, I., Majeed, N., Bhutta, Z.A., Kulyar, M.F.E.A., Fatima, M., Zaheer, C.N.F., Muneer, A., Murtaza, M. and Kashif, M. 2023. MRSA compendium of epidemiology, transmission, pathophysiology, treatment, and prevention within one health framework. Front. Microbiol. 13, 1067284. Shrestha, A., Bhattarai, R.K., Luitel, H., Karki, S. and Basnet, H.B. 2021. Prevalence of methicillin-resistant Staphylococcus aureus and pattern of antimicrobial resistance in mastitis milk of cattle in Chitwan Nepal. BMC Vet. Res. 17, 239. Soccol, C.R., Prado, M.R., Garcia, L.M., Rodrigues, C., Medeiros, A.B. and Soccol, V.T. 2014. Current developments in probiotics. J. Microb. Biochem. Technol. 7, 11–20. Spirescu, V.A., Chircov, C., Grumezescu, A.M., Vasile, B.S. and Andronescu, E. 2021. Inorganic nanoparticles and composite films for antimicrobial therapies. Int. J. Mol. Sci. 22, 4595. Sungkaworn, T. and Triampo, W. 2007. The effects of TiO2 nanoparticles on tumor cell colonies: fractal dimension and morphological properties. J. Nanopart. Res. 9, 519–527. Syed, M.A., Jamil, B., Ramadan, H., Rukan, M., Ali, S., Abbasi, S.A., Woodley, T.A. and Jackson, C.R. 2021. Genetic diversity of Staphylococcus aureus strains from a tertiary care hospital in Rawalpindi, Pakistan. Microorganisms 9, 2301. Syed, M.A., Shah, S.H.H., Sherafzal, Y., Shafi-Ur-Rehman, S., Khan, M.A., Barrett, J.B., Woodley, T.A., Jamil, B., Abbasi, S.A. and Jackson, C.R. 2018. Detection and molecular characterization of methicillin-resistant Staphylococcus aureus from table eggs in Haripur, Pakistan. Foodborne. Pathog. Dis. 15, 86–93. Talha, M., Islam, N.U., Zahoor, M., Sadiq, A., Nawaz, A., Khan, F.A., Gulfam, N., Alshamrani, S.A., Nahari, M.H., Alshahrani, M.A., Mahnashi, M.H. and Hassan, S.S.U. 2022. Biological evaluation, phytochemical screening, and fabrication of Indigofera linifolia leaves extract-loaded nanoparticles. Molecules 27, 4707. Tanuğur Samanci, A.E., Bayar Muluk, N., Samanci, T. and Cingi, C. 2024. Honey: overview. In: Propolis: prevention and healing effects in otorhinolaryngology. Cham, Switzerland: Springer Nature, pp: 29–41; doi: https://doi.org/10.1007/978-3-031-68546-0 Tariq, N., Muhammad Tariq Javed., Aftab Ahmad Anjum., Hafza Zahira Manzoor., Muhammad Hunain Ahmed., Aira Tariq. and Sami Ullah Khan Bahadur. 2023. Antibiotic resistance and molecular detection of nuc and mecA genes of Staphylococcus aureus isolated from mastitic milk. Agric. Sci. J. 5, 17–24. Tominaga, T. and Ishii, M. 2020. Detection of microorganisms with lateral flow test strips. Methods in Microbiology 47, Cambridge, MA: Academic Press, pp: 351–394; doi: 10.1016/bs.mim.2019.11.007 Tyasningsih, W., Effendi, M.H., Budiarto, B. and Syahputra, I.R. 2019. Antibiotic resistance to Staphylococcus aureus and methicillin resistant Staphylococcus aureus (MRSA) isolated from dairy farms in Surabaya, Indonesia. Indian Vet. J. 96(11), 27–31. Vestergaard, M., Frees, D. and Ingmer, H. 2019. Antibiotic resistance and the MRSA problem. Microbiol. Spectr. 7, GPP3-0057-2018. Waqar, N., Amin, Q., Munir, T., Ikram, M.S., Shahzad, N., Mirza, A., Ali, A. and Arshad, M.I. 2019. A cross-sectional study of methicillin-resistant Staphylococcus aureus at the equine-human interface. Trop. Anim. Health. Prod. 51, 1927–1933. Yunita, M.N., Effendi, M.H., Rahmaniar, R.P., Arifah, S. and Yanestria, S.M. 2020. Identification of spa gene for strain typing of methicillin-resistant Staphylococcus aureus (MRSA) isolated from nasal swab of dogs. Biochem. Arch. 20(1), 2999–3004. Zaatout, N. and Hezil, D. 2022. A meta-analysis of the global prevalence of methicillin-resistant Staphylococcus aureus (MRSA) isolated from clinical and subclinical bovine mastitis. J. Appl. Microbiol. 132, 140–154. Zaheer, Z., Hussain, I., Rahman, S.U., Younas, T., Zaheer, I., Abbas, G. and Nasir, M. 2017. Occurrence and antibiotic susceptibility of methicillin-resistant Staphylococcus aureus recovered from oropharynx of live cockerels. J. Appl. Microbiol. 123, 134–141. | ||
| How to Cite this Article |
| Pubmed Style Khan AY, Effendi MH, Shaukat A, Muneeb M. Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Vet. J.. 2026; 16(3): 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 Web Style Khan AY, Effendi MH, Shaukat A, Muneeb M. Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. https://www.openveterinaryjournal.com/?mno=293145 [Access: March 31, 2026]. doi:10.5455/OVJ.2026.v16.i3.3 AMA (American Medical Association) Style Khan AY, Effendi MH, Shaukat A, Muneeb M. Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Vet. J.. 2026; 16(3): 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 Vancouver/ICMJE Style Khan AY, Effendi MH, Shaukat A, Muneeb M. Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Vet. J.. (2026), [cited March 31, 2026]; 16(3): 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 Harvard Style Khan, A. Y., Effendi, . M. H., Shaukat, . A. & Muneeb, . M. (2026) Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Vet. J., 16 (3), 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 Turabian Style Khan, Asfand Yar, Mustofa Helmi Effendi, Aftab Shaukat, and Muhammad Muneeb. 2026. Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Veterinary Journal, 16 (3), 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 Chicago Style Khan, Asfand Yar, Mustofa Helmi Effendi, Aftab Shaukat, and Muhammad Muneeb. "Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan." Open Veterinary Journal 16 (2026), 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 MLA (The Modern Language Association) Style Khan, Asfand Yar, Mustofa Helmi Effendi, Aftab Shaukat, and Muhammad Muneeb. "Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan." Open Veterinary Journal 16.3 (2026), 1438-1450. Print. doi:10.5455/OVJ.2026.v16.i3.3 APA (American Psychological Association) Style Khan, A. Y., Effendi, . M. H., Shaukat, . A. & Muneeb, . M. (2026) Prevalence, risk factors, and innovative therapies for methicillin-resistant Staphylococcus aureus in livestock and companion animals in Pakistan. Open Veterinary Journal, 16 (3), 1438-1450. doi:10.5455/OVJ.2026.v16.i3.3 |