| Research Article | ||
Open Vet. J.. 2025; 15(12): 6823-6835 Open Veterinary Journal, (2025), Vol. 15(12): 6823-6835 Research Article Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offalAya A. Khairy1, Rasha M. El Bayomi2, Zohair S. Mulla3*, Abdelsalam E. Hafez2, Mai F. Saad4, Dalia E. Elatriby5, Saad Elsaeed Elfiky6, Wageh Sobhy Darwish2 and Nafissa A. Mustafa71Faculty of Veterinary Medicine, Zagazig University, Zagazig, Egypt 2Food Hygiene, Safety and Technology Department, Faculty of Veterinary Medicine, Zagazig University, Zagazig, Egypt 3Department of Public Health, College of Veterinary Medicine, King Faisal University, Al-Hofuf, Saudi Arabia 4Department of Veterinary Public Health, Faculty of Veterinary Medicine, Zagazig University, Zagazig, Egypt 5Infection Control Unit, Specialized Internal Medicine Hospital, Mansoura University, Mansoura, Egypt 6Alain Municipality Slaughterhouse, Alain, United Arab Emirates 7Educational Veterinary Hospital, Faculty of Veterinary Medicine, Zagazig University, Zagazig, Egypt *Corresponding Author: Zohair S. Mulla. Department of Public Health, College of Veterinary Medicine, King Faisal University, Al-Hofuf, Saudi Arabia. Email: zmulla [at] kfu.edu.sa Submitted: 09/09/2025 Revised: 09/11/2025 Accepted: 15/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
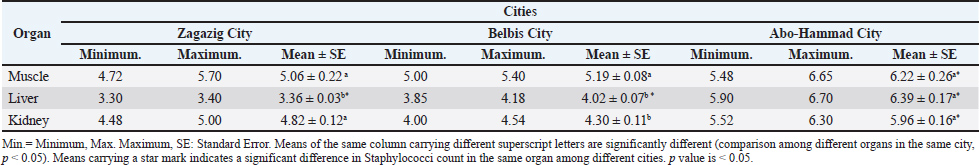
AbstractBackground: Camel's meat is a nutrient-dense and curative food. It is abundant in animal-derived proteins, vital amino acids, vitamins, and minerals. However, this type of meat source is susceptible to contamination with foodborne pathogens such as Salmonella spp. and Staphylococcus aureus. Aim: This study aimed to determine the prevalence of S. aureus and Salmonella spp. in camel meat and offal. Additionally, staphylococcal enterotoxins and virulence-associated genes were screened in the recovered isolates. The antibiograms of the obtained isolates were further examined. Methods: Camel meat and offal, including the liver and kidney, were collected. The isolation and identification of S. aureus and Salmonella spp. were conducted using standard methods. The detection of virulence-associated genes was performed using Polymerase Chain Reaction. Antimicrobial resistance was screened using the disk diffusion method. Results: The overall prevalence rates of S. aureus in the examined muscle, liver, and kidney were 73.33%, 90%, and 100%, respectively. Only 10 of the 79 isolated S. aureus strains were enterotoxigenic (12.7%), carrying Staphylococcus enterotoxin type B (SEB), Staphylococcus enterotoxin type A (SEA)+ Staphylococcus enterotoxin type C (SEC), and SEA enterotoxins. The recovered S. aureus isolates showed marked antimicrobial resistance against kanamycin (100%), ampicillin (96.2%), and tetracycline (84.6%). The overall prevalence of Salmonella spp. in the current study was 27.8% (25 of 90 examined samples). The serological identification of Salmonella isolates identified S. Typhimurium, Salmonella Enteritidis, Salmonella Takoradi, Salmonella Rissen, Salmonella Anatum, and Salmonella Tsevie was detected at 3 (3.3%), 10 (11.1%), 2 (2.2%), 3 (3.3%), 4 (4.4%), and 3 (3.3%), respectively. Salmonella isolates in the current study were resistant to nalidixic acid (100%), oxacillin, and tetracycline (87.5%). The isolated strains harbored virulence genes such as invasion A gene, Salmonella enterotoxin gene, Salmonella plasmid virulence gene, and hyper-invasive locus gene. Conclusion: Camel meat and offal can pose a risk of food poisoning due to the presence of bacteria, including Salmonella spp. and S. aureus. Consequently, camel meat must be processed with the utmost cleanliness. Keywords: Camel meat, Offal, S. aureus, Salmonella spp., Antimicrobial resistance. IntroductionCamel meat is considered a nutrient-rich and curative food, rich in vitamins, minerals, and essential amino acids, such as lysine, leucine, and arginine, as well as carnosine, anserine, and glutathione, which serve as bioactive elements, and omega-3 essential fatty acids. Camels contain less intramuscular fat than beef, lamb, and goat, whereas the camel hump has a high level of saturated fatty acids (Kadim et al., 2018; El-Ghareeb et al., 2019). Camels have recently become a subject of interest in many countries as a safe source of high-nutrient food with high protein content. Furthermore, it contains higher levels of vitamin B and E (Tang et al., 2020; Elkady et al., 2024). The number of diseased cases, the degree of illness, and the number of deaths among the recorded cases were used to estimate the urgency of human health disturbances resulting from microbiological hazards in animal meat. Salmonella spp. and Staphylococcus spp. are the primary microbial hazards most frequently linked to meat (EFSA Panel on Biological Hazards (BIOHAZ), 2013). Staphylococci typically inhabit the nasal cavity, nasopharynx, skin, and mucous membranes. Nearly 30 species of Staphylococci are found, most of which are found in animals. These bacteria are opportunistic pathogens, with only a few being truly pathogenic. Staphylococcal infections are often acute and pyogenic. The coagulase-positive Staphylococcus aureus and S. pseudintermedius are considered the most pathogenic. Staphylococcus aureus is known to cause human food poisoning by secreting enterotoxins types A E as virulent factors, which stimulate the emetic center reflexively, mainly through type A enterotoxin. Staphylococci contamination of food is a life-threatening issue. Enterotoxins produced by S. aureus are heat-stable and resist nearly all sanitizing agents (Darwish et al., 2022). The transmission of harmful bacteria, such as Salmonella, to meat from other food sources, such as cutting boards, utensils, contaminated water, feces, and workers’ hands, during animal slaughtering and meat handling, is the main source of human infection by food-borne illness (Saeed et al., 2013). Salmonella is the most common foodborne pathogen that causes diarrhea, fever, abdominal cramps, enterohemorrhagic colitis, and hemolytic uremic syndrome. Meat-producing animals, such as camels, sheep, and goats, can be carriers, shedding pathogenic bacteria in their feces. The incidence of meat contamination during the dehiding process is high, which serves as a pathway for disease transmission to humans, leading to foodborne illness (Bosilevac et al., 2015). The widespread use of antibiotics in animal breeding without considering the withdrawal time before slaughter is a serious issue. Bacterial resistance to an antibiotic can lead to cross-resistance to other drugs within the same class, with multiple resistance genes carried on plasmids and mobile elements, resulting in multidrug-resistant isolates, such as Salmonella Typhimurium DT104 and methicillin-resistant S. aureus (Alsayeqh et al., 2021). The present study aimed to investigate the prevalence of South aureus and Salmonella spp. in camel meat and offal. Second, the antimicrobial susceptibility of the recovered isolates was studied. Third, the virulence associated with the recovered pathogens was investigated. Materials and MethodsCollection of the samplesNinety samples were collected, including camel muscles, livers, and kidneys (30 of each). The samples were randomly collected from three cities in the Sharkia governorate, Egypt: Zagazig, Abo Hammad, and Belbis cities. The collected samples were immediately sent in an ice box to the Meat Hygiene Laboratory, Faculty of Veterinary Medicine, Zagazig University, Egypt, for bacteriological analysis. Preparation of samplesThe camel meat samples were prepared, and serial dilutions were performed (ISO 6887). A 25-g sample from each sample was aseptically homogenized in 225 ml of 0. 1 buffered peptone water BPW HIMEDIA, M614 using a stomacher Colworth, 400 for 2for 5 minutes at 25. After that, it was left to stand for 5 minutes to produce a homogenate representing a 10-1 dilution. Then, 10-fold serial dilutions were created. Isolation and identification of Staphylococcus aureusA Staphylococcus species count was conducted following APHA’s guidelines (APHA, 2002). Half a milliliter of the diluted solution was added to plates that already contained Baird Parker medium Himedia, M043 100G, Mumbai, India, with an additional 50 ml of egg yolk tellurite emulsion Oxoid SR54. Plates were then incubated at 37°C for at least 1 day. Both typical and atypical coagulase-positive Staphylococci colonies were counted. Colonies often have a shiny, convex appearance, a zone of clearing around them, and a diameter of 1–1.5 mm and 1.5–2.5 mm after 24 and 48 hours of incubation, respectively. Two consecutive plates from each sample were used to determine the overall staphylococcal count; plates with 20 200 colonies were counted (Public Health England, 2016). Gram staining, catalase test, growth on mannitol salt agar base, slide and tube coagulase tests, and detection of hemolysis were some of the biochemical and morphological tests performed on each suspected colony in accordance with ISO (2013). Isolation of Salmonella speciesPre-enrichment in selective liquid mediaThe previously prepared homogenate was placed in an incubator to pre-enrich it with Salmonella. Salmonella species were isolated using the method suggested by Vassiliadis (1983). In brief, 9.0 mL of Rappaport Vassiliadis enrichment broth (HIMEDIA M880) was directly inoculated with 1 ml of the pre-enriched homogenate and left at 41.5°C ± 0.5°C for 24 hours. Selective plating and identificationThe culture was then taken and streaked onto Xylose Lysine Deoxycholate (HIMEDIA M031) agar plates. After incubating at 37°C ± 1°C for 24 hours ± 3 hours, the plates were checked for the presence of suspected Salmonella colonies. These colonies have a black center and a slightly transparent reddish zone, which is caused by the indicator's color change. In accordance with ISO 6579-1 (2017), we collected and stored individual colonies of bacteria in brain heart infusion broth containing 10% (v/v) added glycerol for further biochemical and serological analysis. Serological identification of SalmonellaeThe Kauffman White system for determining somatic O and flagellar H antigens was used to perform the serological identification of Salmonellae, as explained by Kauffman (1974). Antimicrobial susceptibility testing and virulence-associated gene testingTo determine the antibiotic susceptibility of the recovered S. aureus and Salmonella spp. strains, the single diffusion method was employed following Deresse et al. (2012). The susceptibilities of isolated strains were assessed using sensitivity discs from Oxoid Limited (Basingstoke, Hampshire, UK) that had different concentrations. The antibiotic sensitivity of the bacteria was examined using the agar plate method, which involves growing the bacteria on nutrient agar. The nutrient agar was evenly covered with the bacterial culture. Once the inoculation plate was covered, the antibiotic discs were added. After placing the plate in an incubator set at an appropriate temperature (37°C) for 1–2 days, bacterial growth was observed. The National Committee for Clinical Laboratory Standards "NCCLS" (2001) recommended methods for antimicrobial susceptibility testing. We selected the most commonly used antimicrobials during animal farming. The examined isolates were classified as either susceptible, intermediate, or resistant. Each strain’s Multiple Antibiotic Resistance (MAR) index was calculated using the formula laid out by Singh et al. (2010). Enterotoxin detection and typing in the recovered S. aureus isolates were performed according to the method of Shingaki et al. (1981). Virulence analysis of the recovered Salmonella isolates was conducted according to previously established methods (Elafify et al., 2019). Statistical analysisA one-way analysis of variance with post hoc Tukey–Kramer was used to assess statistical significance, with a p-value < 0.05 indicating statistical significance. The SPSS 23 statistical software for the social sciences was used to obtain the Pearson correlation coefficient r; Chicago, IL, USA. Ethical approvalAll experiments were conducted according to the ethical guidelines of Zagazig University. ZU-IACUC/2/F/209/2023. ResultsResults revealed that the mean ± SE of Staphylococci count in the examined camel meat samples was 5.06 ± 0.22, 3.36 ± 0.03 and 4.82 ± 0.12 in the examined camel muscle, liver and kidney samples, respectively from Zagazig city, 5.19 ± 0.08, 4.02 ± 0.07 and 4.30 ± 0.11 in the examined camel muscle, liver and kidney samples, respectively from Belbis city, and 6.22 ± 0.26, 6.39 ± 0.17 and 5.96 ± 0.16 in the examined camel muscle, liver and kidney samples, respectively from Abo-Hammad city (Table 1). Table 2. Prevalence of Staphylococcus spp. in the examined camel meat samples (N=10 of each).
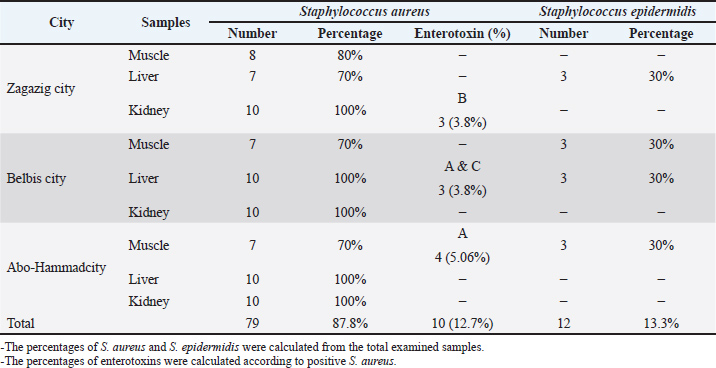
As described in the serological identification of Staphylococcus spp. in the examined camel meat samples (N=10 of each) declared that S. aureus was detected in all kidney samples (100%), in 8 (80%), 7 (70%), and 7 (70%) of muscle samples collected from Zagazig, Belbis, and Abo-Hammad cities, respectively, in 7 (70%) of liver samples collected from Zagazig city and in all liver samples (100%) collected from Belbis and Abo-Hammad cities. Staphylococcus epidermidis was detected in 3 (30%) of muscle samples collected from Belbis and Abo-Hammad cities and in 3 (30%) of liver samples collected from Zagazig and Belbis cities. Results recorded in this study also indicated that the overall prevalence of S. aureus was 87.8% (79 samples out of 90), while the overall prevalence of S. epidermidis was 13.3% (12 samples out of 90). The obtained results revealed that the overall prevalence rates of S. aureus in the examined muscle, liver, and kidney were 73.33%, 90%, and 100%, respectively. The obtained data also showed the prevalence of enterotoxins in the isolated S. aureus strains. It was found that only kidney isolates from Zagazig city carried the SEB gene, with a percentage of 3.8%. Similarly, the SEA + SEC genes were found in S. aureus isolated from liver samples in Belbis city, also at a rate of 3.8%. The SEA gene was detected in muscle isolates from Abo-Hammad city samples at a percentage of 5.06%. In contrast, liver and kidney isolates from Abo-Hammad city did not produce any types of enterotoxins. The obtained results declared that 10 isolates out of 79 positive S. aureus contained enterotoxins with a percentage of 12.7%. The SEA was the most common enterotoxin produced in the 79 tested isolates, with a rate of 5.06%, followed by SEB and (SEA + SEC), with percentages of 3.8% for each. In contrast, S. aureus isolates from the examined camel meat samples failed to produce SED (Table 2). Table 2. Total Staphylococci count Log10 (CFU/g) in the examined camel meat samples (N=10 of each).
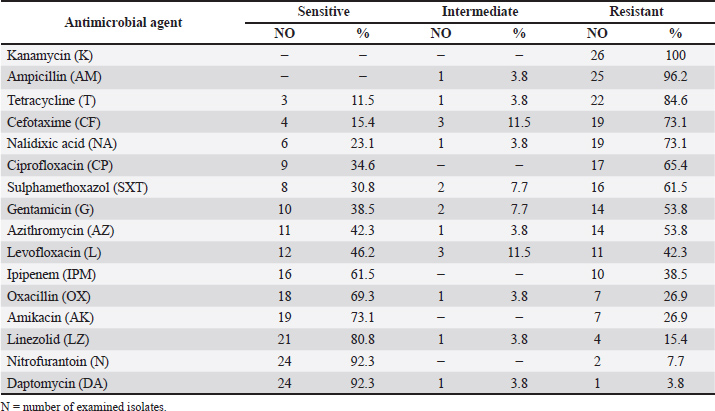
Camel meat isolates were tested for antimicrobial susceptibility profile (n=26). Data declared that all isolates were resistant to kanamycin (100%). Staphylococcus aureus isolates had high resistance against ampicillin (96.2%), tetracycline (84.6%), cefotaxime (73.1%), nalidixic acid (73.1%), ciprofloxacin (65.4%), and sulphamethoxazole (61.5%). Staphylococcus aureus isolates were highly sensitive to nitrofurantoin and daptomycin (92.3%), linezolid (80.8%), amikacin (73.1%), oxacillin (69.3%), and ipipenem (61.5%) (Table 3). Table 3. Antimicrobial susceptibility of Staphylococcus aureus isolated from the examined camel meat samples (N=26).
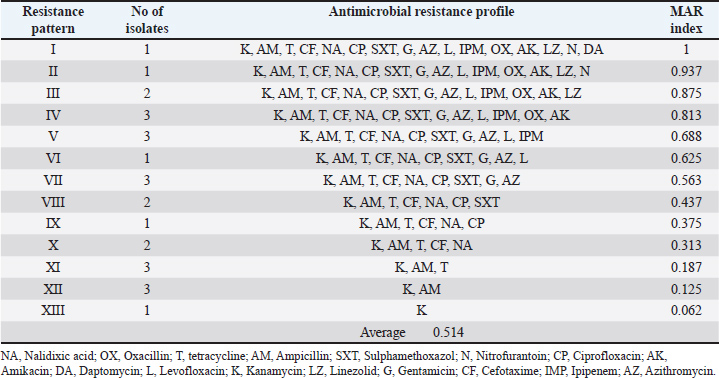
The results clarified that the antibiotic resistance profile of the identified S. aureus isolates to sixteen antibiotics and revealed 13 resistance patterns with a high number (22) of Multidrug resistance S. aureus isolates, with a percentage of 84.6% (Table 4). Table 4. Antimicrobial resistance profile (MAR index) of Staphylococcus aureus isolated from the examined camel meat samples (N=26).
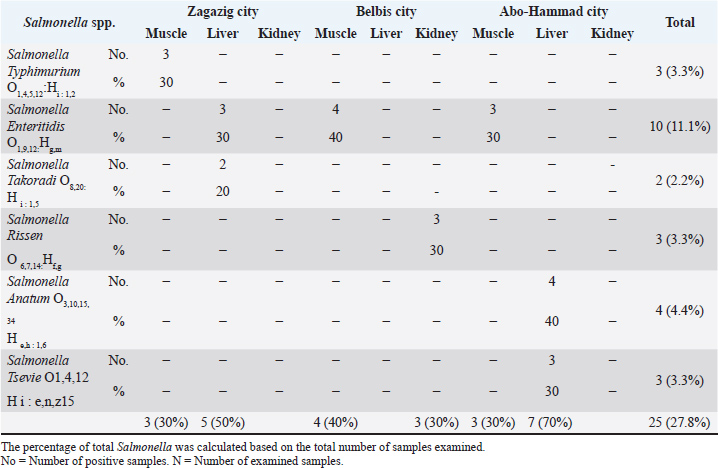
The results showed that the prevalence of Salmonella in camel muscle and liver samples from Zagazig city was 3 (30%) and 5 (50%), respectively, while kidney samples were free from Salmonella. Regarding Belbis samples, Salmonella prevalence in camel muscle and kidney samples was 4 (40%) and 3 (30%), respectively, while Salmonella was not detected in liver samples from the same city. Camel muscle and liver samples from Abo-Hammad city had Salmonella prevalence rates of 3 (30%) and 7 (70%), respectively, with no prevalence in kidney samples. It was declared that 25 samples (27.8%) of the total examined 90 camel meat and offal samples were positive for Salmonella. It was clear that Zagazig and Abo-Hammad liver samples had the highest Salmonella incidence rates; 5 samples (50%) and 7 samples (70%), respectively, followed by Belbis muscle samples with an incidence rate of 4(40%), while the lowest incidence rates were recorded in Zagazig, Abo-Hammad muscle samples, and Belbis kidney samples with an incidence rate of 3(30%) for each. The serological identification of Salmonella isolates declared that S. Typhimurium, S. Enteritidis, S. Takoradi, S. Rissen, S. Anatum and S. Tsevie were detected in 3 (3.3%), 10 (11.1%), 2 (2.2%), 3 (3.3%), 4 (4.4%) and 3 (3.3%), respectively, from the total examined camel muscle, liver and kidney samples from the three cities (Table 5). Table 5. Prevalence of Salmonella spp. in the examined camel meat samples (N=10 of each).
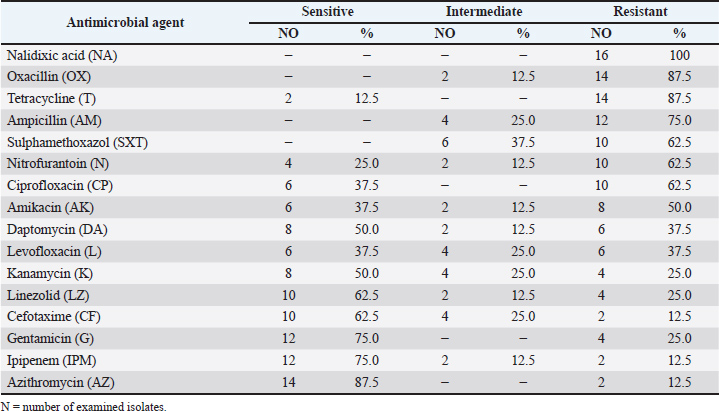
Camel meat isolates were tested for the antimicrobial susceptibility profile of Salmonella spp. (n=16). Data obtained declared that a high number of isolates were resistant to most antibiotics. Isolates were resistant to nalidixic acid (100%), with high resistance against oxacillin and tetracycline (87.5%). It was obvious that isolates had the same resistance rate against sulphamethoxazole, nitrofurantoin, and ciprofloxacin (62.5%), while 50% of the examined Salmonella isolates showed resistance against amikacin. Our findings declared that Salmonella isolates were highly sensitive to azithromycin (87.5%), had the same sensitivity rate to ipipenem and gentamicin (75%), and a lower sensitivity rate to cefotaxime and linezolid (62.5%). While 50% of the examined Salmonella isolates were sensitive to kanamycin (Table 6). Table 6. Antimicrobial susceptibility of Salmonella spp. isolated from the examined camel samples (N=16).
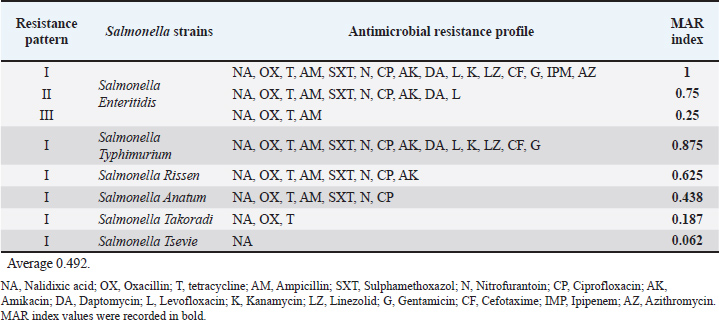
The results clarified the antibiotic resistance profile of the identified Salmonella isolates to sixteen antibiotics and revealed that S. Enteritidis showed multidrug resistance with three resistance patterns, and MAR index values ranged from 0.25 to 1. While S. Typhimurium, S. Rissen, S. Anatum, S. Takoradi, and S. Tsevie had one resistance pattern with MAR index values of 0.875, 0.625, 0.438, 0.187, and 0.062, respectively. The isolates' MAR index ranged from 0.187 to 1, with an average of 0.492 (Table 7). Table 7. Antimicrobial resistance profile (MAR index) of Salmonella spp. isolated from the examined camel meat samples (N=16).
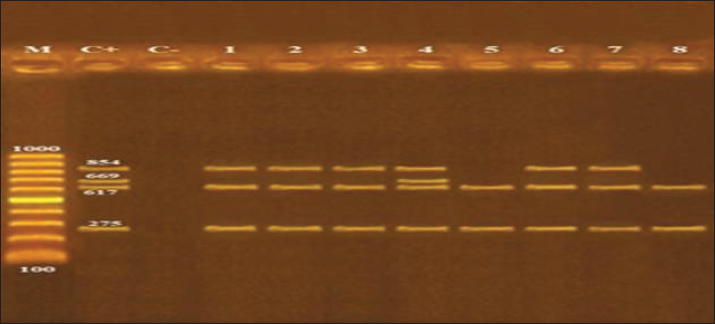
As illustrated, the molecular characterization of Salmonella spp. using Polymerase Chain Reaction clarified that the isolated strains harbored virulence genes such as invasion A gene (invA), enterotoxin gene (stn), Salmonella plasmid virulence gene (spvC), and hyper-invasive locus gene (hilA). Invasion A gene and Salmonella enterotoxin gene (stn) were detected in all identified Salmonella strains, while spvC was detected only in S. Typhimurium. Regarding the hilA gene, it was detected in S. Anatum, S. Enteritidis, S. Tsevie, and S. Typhimurium (Table 8 and Fig. 1).
Fig. 1. Agarose gel electrophoresis of multiplex PCR of invA (275 bp), stn (617 bp), spvC (669 bp) and hilA (854 bp) virulence genes for characterization of Salmonella species. Lane M: 100 bp ladder as molecular size DNA marker. Lane C+: Control positive strain for invA, stn, spvC and hilA genes. Lane C-: Control negative. Lanes 1, 2, 3 S. Enteritidis; 6 S. Tsevie and 8 S. Anatum: Positive strains for invA, stn and hilA genes. Lane 4 S. Typhimurium: Positive strain for invA, stn, spvC and hilA genes. Lane 5 S. Rissen and 8 S. Takoradi: Positive strains for invA and stn gene. DiscussionA significant prevalence of Staphylococcus species is associated with meat contamination from surfaces that come into contact with meat, including floors, cutting boards, tables, and workers’ hands (Çetin et al., 2006). Camel muscle samples collected from Belbis city had a higher level of Staphylococcus contamination than liver and kidney samples from the same city. A highly significant difference at p value of 0. 05 was observed in the mean of Staphylococcus count in the examined camel samples from Abo Hammad city compared with samples from other cities. Bad hygienic practices of some workers, such as coughing, sneezing, and failure to follow health requirements during meat handling, make meat and meat products prone to contamination by pathogenic bacteria, such as S. aureus. Higher figures were obtained by Alaboudi et al. (2012), who investigated the prevalence rate of Staphylococcus in 147 camel meat samples collected from abattoirs and meat markets in Irbid Governorate, Jordan, and found that approximately 97.27% of the examined camel meat samples were contaminated by S. aureus. In addition, higher rates were recorded by Rahimi et al. (2013), who conducted a study on 50 camel meat samples from slaughterhouses in Isfahan, Iran. The bacteriological examination of the samples indicated that 46% of the camel meat was contaminated by S. aureus. Furthermore, Tegegne et al. (2019) conducted a study on 140 camel meat samples collected from slaughterhouses and retail shops and found that the mean ± SE of S. aureus was 6.19 ± 0.10 log10 CFU/g-1 and 6.33 ± 0.04 log10 CFU/g-1 in abattoir and retail shop samples. Results recorded by Ras (2019) who carried out a study on camel liver and kidney samples (10 from each) collected from Sharkia province, Egypt, and found that the mean count of S. aureus in camel kidney and liver samples was 4.2 ± 0.24 log10 CFU/g and 4.1 ± 0.15 log10 CFU/g, respectively were lower than those recorded in camel samples from Abo-Hammad city and then camel muscle samples from Zagazig and Belbis cities in the current study. Meanwhile, the mean counts in liver samples collected from Zagazig city were higher than those in liver and kidney samples collected from Belbis city in the current study. Naas et al. (2019) performed a study on 21 camel meat samples and found that from 5 (23.8%) selected isolates for S. aureus, only one sample (20%) was positive for S. aureus by partial sequencing of 16S rDNA. Besides, Gwida et al. (2019) conducted a study on 200 samples from camel meat and liver (100 for each) from Zagazig abattoir, Sharkia province, Egypt, and found that camel meat and liver samples were contaminated by S. aureus with a prevalence of 45% and 25%, respectively, with an overall incidence of 35%. Furthermore, Abdi Hassen et al. (2021) conducted a study on 68 raw camel meat samples from slaughterhouses and retail shops (34 samples each) in Nagelle town, Southern Oromia, Ethiopia, and recorded that the mean of S. aureus counts was 2.76 log10 CFU/g for abattoir samples and 3.07 log10 CFU/g for butchery samples. In addition, Salman et al. (2024) conducted a study on 100 camel meat samples collected from El-Basatien slaughterhouses in Cairo governorate, Egypt, and noticed that 24% of the examined samples were contaminated by S. aureus. The findings of the present work were in accordance with Abdal-Rudha et al. (2024), who performed a study on 35 camel meat samples from Najaf governorate/ Iraq, and revealed that S. epidermidis incidence in camel meat samples was 14.2%; meanwhile, S. aureus incidence in camel meat samples (14.2%) was lower than that recorded in the present study. Staphylococcal food poisoning can be caused by consuming foods or drinks that contain one or more preformed enterotoxins, which are categorized according to their antigenic properties: SEA, SEB, SEC, SED, and SEE (Pinto et al., 2005). Less than 1 mg of SE is required to produce drunkenness. When the amount of germs in food surpasses 100,000 /g, we approach this toxic level. For those who are really sensitive, even 100–200 ng of enterotoxin can trigger symptoms. Hospitalization may be necessary depending on the severity of the illness (Bennett 2012). Staphylococcal food intoxication is most commonly caused by eating meat with sufficient levels of one or more preformed enterotoxins. According to Argudín et al. (2010), SEA stands out as the leading cause of Staphylococcal food poisoning worldwide among the several enterotoxins produced by S. aureus. Rahimi et al. (2013) also discovered that of 23 camel meat samples from Isfahan, Iran, four S. aureus isolates 17 %) produced Staphylococcal enterotoxins, one for each of the four types A, B, C, and A C. Alaboudi et al. (2012) tested 58 S. aureus isolates from camel meat samples in Jordan’s Irbid Governorate for enterotoxigenicity and found higher quantities. A total of 41 isolates (70.7% of the total) were determined to have Staphylococcus enterotoxigenic genes. Further information was gleaned from the fact that the SEA toxin gene ranked highest, followed by the SEA+SEB and SEC genes. No evidence of SEE or SED genes was found in any of the samples. Ras (2019) discovered that S. aureus isolates from camel liver and kidney samples taken from the Sharkia abattoir in Egypt contained Staphylococci enterotoxins. A total of 69.2% of the examined S. aureus isolates were found to be negative for enterotoxin genes, whereas 15.4%, 7.7%, and 7.7% of the tested isolates were found to have SEA, SED, and SEA + SEB genes, respectively. Of all the S. aureus isolates tested, only one had the SEA+SED gene, whereas none of the others had the SEC gene. The widespread use of antibiotics in animal breeding, without considering the withdrawal time before slaughter, and the unintended use of supplements and growth promoters in animal husbandry play a significant role in the emergence of MDRB. The emergence of multidrug-resistant bacteria, which are not susceptible to one or more antibacterial agents in three or more different antibacterial categories, is considered a public health hazard because there are fewer or nearly no antibacterial agents that can effectively treat infections caused by these pathogenic bacteria (Alsayeqh et al., 2021). Raji et al. (2016) recorded similar results, who found that methicillin-resistant Staphylococcus aureus and methicillin-sensitive Staphylococcus aureus isolates were susceptible to linezolid and sulfamethoxazole. Salman et al. (2024) reported similar results, finding that S. aureus isolates were highly ampicillin-resistant and cefotaxime-sensitive to imipenem. Torki Baghbaderani et al. (2020) assessed the phenotypic profile of antibiotic resistance of S. aureus isolates from camel meat and found that S. aureus had high resistance against tetracycline and penicillin, with incidence rates of 79. 16 and 72. 91, respectively, and lower resistance against levofloxacin and azithromycin, with incidence rates of 22. 91 and 25, respectively. In contrast to Al-Amery et al. (2019), S. aureus isolates from camel meat in Egypt exhibited 86.2% resistance to oxacillin. The MAR index of the isolates ranged from 0.125 to 1, with an average of 0.514. MAR index ˃ 0.2 indicates a high contamination rate and misuse of antimicrobial agents at the source of the isolates (Abd El-Hamid and Bendary, 2015). Of the 53 S. aureus isolates from camel meat samples, 11 isolates (20.7%) were sensitive to cefoxitin, oxacillin, levofloxacin, erythromycin, and trimethoprim/ sulfamethoxazole, while 35.8% (19/53) were resistant to penicillin. MAR index ranged from 0.05 to 0.3 (Alkuraythi et al., 2024). Camel meat and edible offal may be a source of the enteric pathogens Salmonella spp., causing human salmonellosis, which is considered a public health hazard (Raufu et al., 2015). The obtained results were comparable to Molla et al. (2004), who conducted a study on camel muscle and liver samples from Ethiopia, and found that Salmonella was isolated with a prevalence rate of 20.1% and 11.8%, respectively, which is lower than the current study (33.3% and 40%) in camel muscle and liver samples, respectively. The serological identification also revealed that S. Typhimurium and S. Enteritidis had the lowest prevalence rate of 0.9% for each, which was lower than our study results. The overall Salmonella isolation rate obtained in the current study was nearly similar to the Salmonella prevalence rate (29.4%) recorded by Abdi Hassen et al. (2021) in camel meat samples from Nagelle butcher shops, Ethiopia, and higher than the results recorded in the same study in abattoir camel meat samples (23.5%). A lower prevalence rate of Salmonella (8%) was recorded by Münch et al. (2012) in liver and kidney samples from camels slaughtered in the United Arab Emirates. Salmonella was isolated from 20%, 10% and 5% in camel meat samples from Elbagour, Menouf and Shibin-Elkom, respectively, which were lower than the current study results, while the serological identification of Salmonella isolates revealed that S. Enteritidis and S. Typhimurium were identified in 20% and 5% of the isolates, respectively which were higher than the results recorded in the current study (Edris et al., 2013). Unlikely, Mandour et al. (2014) could not isolate Salmonella spp. from camel meat samples in Al-Ahsaa central abattoir, Saudi Arabia. Nossair et al.'s (2016) study on camel muscle, liver, and kidney samples from Behera Province, Egypt, revealed that muscle samples were contaminated by Salmonella with an incidence rate of 16% followed by liver and kidney samples with incidence rates of 8% and 4%, respectively. The serological identification revealed that the most incriminated serotypes were S. Enteritidis with an incidence rate of 8% and 4% in muscle and liver samples, respectively, while S. Typhimurium was detected in 4% of muscle samples. Besides, Alkhalaf et al. (2024) who conducted a study on camel meat, liver and kidney vended in Zagazig city, Egypt, and found that Salmonella incidence rate in liver samples was 30% followed by kidney samples with an incidence rate of 20% and the lowest incidence was in muscle samples (10%), the most prominent isolated serotypes were S. Typhimurium and S. Enteritidis. Salmonella spp. can easily acquire resistance to antimicrobial agents, especially to the new antibacterial drugs, causing medical failure for the pathogenic bacterial infections in humans and animals (Al-Shafee and Abdulwahid, 2024). Comparable results were obtained by Scharawe et al. (2009), who declared that Salmonella isolates from camel muscle swabs, Cairo abattoir, Egypt, were resistant to ampicillin and nalidixic acid. Raufu et al. (2015) found that most of the isolated Salmonella serotypes from camel liver samples, Nigeria, were sensitive to gentamycin, while they differed from our findings of being sensitive to nalidixic acid and amoxicillin. Alkhalaf et al. (2024) found that the isolated Salmonella serotypes from camel meat, liver, and kidney samples, Zagazig City, Egypt, had antimicrobial resistance against nalidixic acid, ampicillin, and oxacillin. Likely, Salman et al. (2024) study, who recorded that all Salmonella isolates from camel meat samples, Beheira Governorate, Egypt had resistance against ampicillin, tetracycline, and nalidixic acid, while both studies disagreed in cefotaxime resistance; Salmonella isolates in the current study were sensitive to cefotaxime antibiotic. However, Münch et al. (2012) recorded that Salmonella isolates from camel liver and kidney samples, United Arab Emirates, had lower resistance to nalidixic acid, ciprofloxacin, sulphamethoxazole, and ampicillin, while the susceptibility to cefotaxime and gentamycin agreed with the current study. Several issues can cause antimicrobial resistance by pathogenic bacteria, including modification in the antibiotic of choice, or the antibiotic can’t pass through the bacterial cell, or is pumped out of the bacterial cytoplasm by efflux system, or is destroyed (Iyamba et al., 2022). A nearly similar MAR index of 0.472 was recorded by Sallam et al. (2024) in Salmonella isolates from camel meat samples. While our findings differed from those of Alkhalaf et al. (2024), who reported that the MAR index for S. Enteritidis ranged from 0.71 to 1 and for S. Typhimurium ranged from 0.36 to 1 in camel muscle, liver, and kidney samples. A number of chromosomal and plasmid factors are responsible for Salmonella virulence, such as invA, spv, fimA, and stn genes. The invA gene is harbored in a genome and codes for a protein on the inner membrane, which is responsible for attacking host epithelial cells (Elkhadragy et al., 2020). Salmonella enterotoxin gene is a virulence factor distinctive to Salmonella spp. and their serotypes, which plays a principal role in the bacterium's enterotoxic effects, manifesting as gastroenteritis and diarrheal illness (Fekry et al., 2020). Hyper-invasive locus gene represents an important Salmonella virulence gene. It is involved in the type III secretion genes regulation; the type III secretion system secretes proteins responsible for host epithelial cell invasion (Cardona-Castro et al., 2002). Salmonella plasmid virulence gene is an essential Salmonella virulence gene that is responsible for the viability and proliferation of bacteria inside host macrophage cells (Mohamed et al., 2021). In agreement with our study, Saad et al. (2018) detected Stn, hilA, and fimH virulence genes in S. Typhimurium and S. Enteritidis isolates from camel meat at Menouf slaughterhouse, Egypt. Similar results were recorded in Ibrahim et al. (2022), who found the invA gene with 100% incidence in the positive camel samples retailed in Tripoli, Libya. In addition, Sallam et al. (2024) study recorded stn, spvc, and hilA virulence genes in 91.4%, 20.3% and 80.5%, respectively, of Salmonella isolates detected in the examined camel meat retailed in Beheira Governorate, Egypt. ConclusionThe results of this study indicated that the examined camel meat samples were contaminated with various microorganisms, reflecting unhygienic measures and unfavorable environmental conditions during handling, transportation, and storage. The results of this study revealed that the camel muscle, liver, and kidney were bacteriologically contaminated with Staphylococci and Salmonella spp. Staphylococcus aureus enterotoxigenic genes, including SEB, SEA, SEC, and SEA genes, were detected alongside Salmonella virulence genes, such as invA, san, spvC, and hilA. Therefore, the strict application of hygienic rules at each step during slaughtering, dressing, evisceration, and loading, as well as effective cleaning and disinfection procedures, are strongly recommended. AcknowledgmentsNone. Conflict of interestThe authors declare that there is no conflict of interest. FundingNot applicable. Authors’ contributionsAll authors contributed equally. Data availabilityAll data were provided in the manuscript. ReferencesAbd El-hamid, M.I. and Bendary, M.M. 2015. Comparative phenotypic and genotypic discrimination of methicillin-resistant and susceptible Staphylococcus aureus in Egypt. Cell. Mol. Biol. 61(4), 101–112. Abdal-Rudha, A.M.H., Al-Nasiry, B.S.A.N. and Dakheel, M.M. 2024. Evaluation of bacterial contamination and heavy metals in cow and camel meat. Arch. Zootechnica. 27(1), 146–154. Abdi Hassen, K., Abdureman Omer, S. and Abdi Hassen, N. 2021. Assessment of hygienic practice on camel meat handlers, and identification of bacterial contamination in Abattoir and Butcheries of Nagelle Town, Southern Oromia, Ethiopia. Inter. Res. J. Sci. Technol. 2(2), 384–397. Alaboudi, A.R., Jaradat, Z.W. and Shatnawi, M.M. 2012. Biotypes and enterotoxigenicity of Staphylococci isolated from Camel’s meat in Jordan. Microbiol. Res. J. Inter. 2(1), 23–35. Al-Amery, K., Elhariri, M., Elsayed, A., El-Moghazy, G., Elhelw, R., El-Mahallawy, H., El Hariri, M. and Hamza, D. 2019. Vancomycin-resistant Staphylococcus aureus isolated from camel meat and slaughterhouse workers in Egypt. Antimicrob. Res. Infect. Cont. 8(129), 1–8. Alkhalaf, A.A., El-Ghareeb, W.R., Raheem, S.M.A., Seliem, M.M., Shosha, A.M. and Elzawahry, R.R. 2024. A study on the prevalence of multidrug-resistant food poisoning Salmonella spp. in camel meat and offal with a reduction trial using organic acids. J. Adv. Vet. Res. 14(3), 526–530. Alkuraythi, D.M., Alkhulaifi, M.M., Binjomah, A.Z., Alarwi, M., Mujallad, M.I., Alharbi, S.A., Alshomrani, M., Gojobori, T. and Alajel, S.M. 2024. Comparative genomic analysis of antibiotic resistance and virulence genes in Staphylococcus aureus isolates from patients and retail meat. Front. Cell. Infect. Microbiol. 13, 1339339. Alsayeqh, A.F., Baz, A.H.A. and Darwish, W.S. 2021. Antimicrobial-resistant foodborne pathogens in the Middle East: a systematic review. Environ. Sci. Pollut. Res. 28(48), 68111–68133. Al-Shafee, A.A.J. and Abdulwahid, M.T. 2024. Occurrence, antimicrobial resistance, and molecular characterization of Salmonella enterica from chicken products and human in Wasit Governorate of Iraq. Open Vet. J. 14(5), 1117. American Public Health Association (APHA) 2002. Methods for the microbiological examination of foods. 4th Ed., Technical Committee on Microbiological Methods for Foods. Washington, DC: American Public Health Association. Argudín, M., Mendoza, M.C. and Rodicio, M.R. 2010. Food poisoning and Staphylococcus aureus enterotoxins. Toxins (Basel). 2(7), 1751–1773. Bennett, R. 2012. Bad Bug Book (2nd Edition). Food and Drug Administration. Handbook of Food borne Pathogenic Microorganisms and Natural Toxins. Gram-positive bacteria. Staphylococcus aureus. Laurel, MD, USA. Pp. 87–88. Bosilevac, J.M., Gassem, M.A., Al Sheddy, I.A., Almaiman, S.A., Al-Mohizea, I.S., Alowaimer, A. and Koohmaraie, M. 2015. Prevalence of Escherichia coli O157: h7 and Salmonella in camels, cattle, goats, and sheep harvested for meat in Riyadh. J. Food Prot. 78(1), 89–96. Cardona-Castro, N., Restrepo-Pineda, E. and Correa-Ochoa, M. 2002. Detection of hilA gene sequences in serovars of Salmonella enterica subgroups enterica. Memórias. Do Instituto Oswaldo Cruz 97, 1153–1156. Çetin, O., Kahraman, T. and Büyükünal, S.K. 2006. Microbiological evaluation of food contact surfaces at red meat processing plants in Istanbul, Turkey. Ital. J. Anim. Sci. 5(3), 277–283. Darwish, W.S., El-Ghareeb, W.R., Alsayeqh, A.F. and Morshdy, A.E.M. 2022. Foodborne intoxications and toxicoinfections in the Middle East. In Food safety in the Middle East. Savvaidis, I.N. and Osaili, T.M. (eds), Academic Press, pp: 109–141, London, United Kingdom Edris, A.M., Hassan, M.A., Shaltout, F. and El-Hosseny, S. 2013. Detection of E. coli and Salmonella organisms in cattle and camel meat. Benha Vet. Med. J. 24(2), 198–204. EFSA Panel on Biological Hazards (BIOHAZ). 2013. Scientific Opinion on the public health hazards to be covered by inspection of meat (bovine animals). EFSA. J. 11(6), 3266. Elafify, M., Darwish, W.S., Al-Ashmawy, M., Elsherbini, M., Koseki, S., Kawamura, S. and Abdelkhalek, A. 2019. Prevalence of Salmonella spp. in Egyptian dairy products: molecular, antimicrobial profiles and a reduction trial using d-tryptophan. J. Cons. Prot. Food Saf. 14(4), 399–407. El-Ghareeb, W.R., Darwish, W.S. and Meligy, A.M.A. 2019. Metal contents in the edible tissues of camel and sheep: human dietary intake and risk assessment in Saudi Arabia. Jpn. J. Vet. Res. 67(1), 5–14. Elkady, S., Darwish, W., Tharwat, A., Said, M., Elatriby, D., Seliem, M., Alfifi, A., Ghareeb, W. and Gad, T. 2024. Prevalence and antibiogram of shigatoxin producing E. coli in camel meat and offal. Open Vet. J. 14(1), 571. Elkhadragy, M.F., Al-masoud, A.A. and Al-Dagal, M.M. 2020. invA Gene to Detect Salmonella Enterica Serovar Typhimurium Supported by Serum Anti-Salmonella Antibodies and Protein Profiles for Chicken Carcass Isolates. Version 1. Fekry, A., Fahim, S., Hussein, W., Hanafy, W. and Elbeltagy, A. 2020. Investigation of virulent and toxin genes expression in some Salmonella serovars. Menoufia. J. Agricul. Biotechnol. 5(2), 67–81. Gwida, M., Zakaria, A., El-Sherbiny, H., Elkenany, R. and Elsayed, M. 2019. Prevalence of Campylobacter, Enterococcus and Staphylococcus aureus in slaughtered camels. Vet. Med. 64(12), 521–530. Ibrahim, K., Aljfaeri, S., Aslougi, A., Alsonosi, A., Saad, M., Eltaib, F.I. and Fadel, M.E. 2022. Study on the Incidence of Salmonella Species in Different Food Samples by Using Cultural and Rapid invA Gene Specific PCR-Based Assay. Inter. J. Microbiol. Biotechnol. 7(1), 11–15. ISO 6579-1. 2017. Microbiology of the food chain — Horizontal method for the detection, enumeration and serotyping of Salmonella — Part 1: Detection of Salmonella spp. International Standards Organization (ISO), 2013. Microbiology of food and animal feeding stuffs: Preparation of test sample, initial suspension and decimal dilutions for microbiological examination, International Standards Organization (ISO 6887-1), Geneva. Iyamba, J.M.L., Lukukula, C.M., Unya, J.W., Ngbandani, B.K., Bissingou, E., Mabankama, M. and Takaisi-Kikuni, N.B. 2022. Antibiotic Resistance Pattern and Biofilm Formation of Staphylococcus and Enterobacteriaceae Isolates from Clinical Samples of Patients with Urinary Tract and Surgical Site Infections in Kinshasa, Democratic Republic of Congo. J. Pharm. Pharmacol. Res. 6(4), 158–168. Kauffman. 1974. Kauffman's white scheme. J. Acta. Pathol. Microbiol. Sci. 61, 385. Kadim, I.T., Al-Amri, I.S., Alkindi, A.Y. and Mbaga, M. 2018. Camel Meat production and quality: a review. J. Camel Pract. Res. 25(1), 9–23. Mandour, M.A., Altabary, G.F., El-Ghareeb, W.R. and Al –Shami, S.A. 2014. Microbial assessment for camel and mutton carcasses slaughtered at Al-Ahsaa Abattoir, Saudi Arabia. J. Anim. Vet. Adv. 13(21–24), 1179–1184. Mohamed, M., Mohamed, R., Gharieb, R., Amin, M. and Ahmed, H. 2021. Antimicrobial resistance, virulence-associated genes and biofilm formation of Salmonella species isolated from different sources. Zagazig Vet. J. 49(2), 208–221. Molla, B., Mohammed, A. and Salah, W. 2004. Salmonella prevalence and distribution of serotypes in apparently healthy slaughtered camels (Camelus dromedarius) in Eastern Ethiopia. Trop. Anim. Health Prod. 36(5), 451–458. Münch, S., Braun, P., Wernery, U., Kinne, J., Pees, M., Flieger, A., Tietze, E. and Rabsch, W. 2012. Prevalence, serovars, phage types, and antibiotic susceptibilities of Salmonella strains isolated from animals in the United Arab Emirates from 1996 to 2009. Trop. Anim. Health Prod. 44(7), 1725–1738. Naas, H.T., Edarhoby, R.A., Garbaj, A.M., Azwai, S.M., Abolghait, S.K., Gammoudi, F.T., Moawad, A.A., Barbieri, I. and Eldaghayes, I.M. 2019. Occurrence, characterization, and antibiogram of Staphylococcus aureus in meat, meat products and some seafood from Libyan retail markets. Vet. World. 12(6), 925–931. National Committee for Clinical Laboratory Standards "NCCLS" 2001. Performance standards for antimicrobial susceptibility testing. Supplement M100-S11. Villanova, PA, USA. Nossair, M.A., Samaha, I.A. and Khalifa, E. 2016. Camels' meat and offal as vehicles of E. coli and Salmonella with special emphasis on their zoonotic importance. In 3rd International Food Safety Conference, 2016 Damanhur University, pp 1–8, Damanhour city, Egypt Pinto, B., Chenoll, E. and Aznar, R. 2005. Identification and typing of food-borne Staphylococcus aureus by PCR-based techniques. Systematic Appl. Microbiol. 28(4), 340–352. Public Health England, 2016. Enumeration of coagulase-positive staphylococci (Staphylococcus aureus and other species). National Infection Service, Food, Water and Environmental Microbiology Standard Method FNES8 (F12), version 4. Rahimi, E., Nonahal, F. and Ataye Salehi, E. 2013. Detection of Classical Enterotoxins of Staphylococcus aureus Strains Isolated from Raw Meat in Esfahan, Iran. Health. Scope. 2(2), 95–98. Raji, M.A., Garaween, G., Ehricht, R., Monecke, S., Shibl, A.M. and Senok, A. 2016. Genetic Characterization of Staphylococcus aureus Isolated from Retail Meat in Riyadh, Saudi Arabia. Front. Microbiol. 7, 911. Ras, R. 2019. Parasitic and bacterial assessment of edible offal of slaughtered animals at Sharkia abattoirs, Egypt. Egy. Vet. Med. Soc. Parasitol. J. 15(1), 64–88. Raufu, I.A., Odetokun, I.A., Oladunni, F.S., Adam, M., Kolapo, U.T., Akorede, G.J., Ghali, I.M., Ameh, J.A. and Ambali, A. 2015. Serotypes, antimicrobial profiles and public health significance of Salmonella from camels slaughtered in Maiduguri central abattoir, Nigeria. Vet. World 8(9), 1068–1072. Saad, S., Abou-Elroos, N. and Abou Elghar, I. 2018. Occurrence of enterobacteriaceae in fresh camel meat with special reference to salmonellae. Benha Vet. Med. J. 35(2), 79–95. Saeed, A.A., Hasoon, M.F. and Mohammed, M.H. 2013. Isolation and molecular identification of Salmonella typhimurium from chicken meat in Iraq. J. World’s. Poult. Res. 2(2), 63–67. Sallam, K.I., Kasem, N.G., Abdelkhalek, A. and Elshebrawy, H.A. 2024. Extensively drug-, ciprofloxacin-, cefotaxime-, and azithromycin-resistant Salmonella enterica serovars isolated from camel meat in Egypt. Inter. J. Food. Microbiol. 411, 110538. Salman, M., Zin Eldin, A., Ata, N., Abdelfatah, E. and Eissa, N. 2024. Prevalence, Antimicrobial Resistance and Risk Factors of Zoonotic Foodborne Pathogens Isolated from Camel Meat. J. Curr. Vet. Res. 6(1), 272–284. Scharawe, H., Ibrahim, H. and Safwat, E. 2009. Studies on some Salmonella serovars isolated from slaughtered imported camels. Vet. Med. J. 57(1), 23–33. Shingaki, M., Igarashi, H., Fujikawa, H., Ushioda, H., Terayrna, T. and Sakai, S. 1981. Study on Reversed Passive Latex Agglutination for detection of staphylococcal enterotoxins A, B, and C. Annu.; Rep. Tokyo, metro p. Res. Lab. Public Health, 32(1), 128–131. Singh, S., Yadav, A.S., Singh, S.M. and Bharti, P. 2010. Prevalence of Salmonella in chicken eggs collected from poultry farms and marketing channels and their antimicrobial resistance. Food Res. Inter. 43(8), 2027–2030. Tang, H., Darwish, W.S., El-Ghareeb, W.R., Al-Humam, N.A., Chen, L., Zhong, R.M., Xiao, Z.J. and Ma, J.K. 2020. Microbial quality and formation of biogenic amines in the meat and edible offal of Camelus dromedaries with a protection trial using gingerol and nisin. Food. Sci. Nutr. 8(4), 2094–2101. Tegegne, H.A., Berhanu, A., Getachew, Y., Serda, B., Nölkes, D., Tilahun, S. and Sibhat, B. 2019. Microbiological safety and hygienic quality of camel meat at abattoir and retail houses in Jigjiga city, Ethiopia. J. Infect. Develop. Count. 13(3), 188–194. Torki Baghbaderani, Z., Shakerian, A. and Rahimi, E. 2020. Phenotypic and genotypic assessment of antibiotic resistance of Staphylococcus aureus bacteria isolated from retail meat. Infect. Drug. Resist. 7(13), 1339–1349. Vassiliadis. 1983. The Rappaport—Vassiliadis (RV) enrichment medium for the isolation of salmonellas: an overview. J. Appl. Microbiol. 54(1), 69–76. | ||
| How to Cite this Article |
| Pubmed Style Khairy AA, Bayomi RME, Mulla ZS, Hafez AE, Saad MF, Elatriby DE, Elfiky SE, Darwish WS, Mustafa NA. Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Vet. J.. 2025; 15(12): 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 Web Style Khairy AA, Bayomi RME, Mulla ZS, Hafez AE, Saad MF, Elatriby DE, Elfiky SE, Darwish WS, Mustafa NA. Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. https://www.openveterinaryjournal.com/?mno=289299 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.62 AMA (American Medical Association) Style Khairy AA, Bayomi RME, Mulla ZS, Hafez AE, Saad MF, Elatriby DE, Elfiky SE, Darwish WS, Mustafa NA. Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Vet. J.. 2025; 15(12): 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 Vancouver/ICMJE Style Khairy AA, Bayomi RME, Mulla ZS, Hafez AE, Saad MF, Elatriby DE, Elfiky SE, Darwish WS, Mustafa NA. Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 Harvard Style Khairy, A. A., Bayomi, . R. M. E., Mulla, . Z. S., Hafez, . A. E., Saad, . M. F., Elatriby, . D. E., Elfiky, . S. E., Darwish, . W. S. & Mustafa, . N. A. (2025) Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Vet. J., 15 (12), 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 Turabian Style Khairy, Aya A., Rasha M. El Bayomi, Zohair S. Mulla, Abdelsalam E. Hafez, Mai F. Saad, Dalia E. Elatriby, Saad Elsaeed Elfiky, Wageh Sobhy Darwish, and Nafissa A. Mustafa. 2025. Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Veterinary Journal, 15 (12), 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 Chicago Style Khairy, Aya A., Rasha M. El Bayomi, Zohair S. Mulla, Abdelsalam E. Hafez, Mai F. Saad, Dalia E. Elatriby, Saad Elsaeed Elfiky, Wageh Sobhy Darwish, and Nafissa A. Mustafa. "Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal." Open Veterinary Journal 15 (2025), 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 MLA (The Modern Language Association) Style Khairy, Aya A., Rasha M. El Bayomi, Zohair S. Mulla, Abdelsalam E. Hafez, Mai F. Saad, Dalia E. Elatriby, Saad Elsaeed Elfiky, Wageh Sobhy Darwish, and Nafissa A. Mustafa. "Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal." Open Veterinary Journal 15.12 (2025), 6823-6835. Print. doi:10.5455/OVJ.2025.v15.i12.62 APA (American Psychological Association) Style Khairy, A. A., Bayomi, . R. M. E., Mulla, . Z. S., Hafez, . A. E., Saad, . M. F., Elatriby, . D. E., Elfiky, . S. E., Darwish, . W. S. & Mustafa, . N. A. (2025) Prevalence, virulence attributes, and antimicrobial resistance profiling of Staphylococcus aureus and Salmonella spp., isolated from camel meat and offal. Open Veterinary Journal, 15 (12), 6823-6835. doi:10.5455/OVJ.2025.v15.i12.62 |