| Research Article | ||
Open Vet. J.. 2026; 16(1): 71-80 Open Veterinary Journal, (2026), Vol. 16(1): 71-80 Research Article Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, ChinaZhuolei Yang1#, Jingyu Gao1#, Lvpeng Jiang1, Nuo Xu1, Sijia Sun1, Wanli Sha1,2,3, Baishuang Yin1,2,4, Wenlong Dong1,2,3*, and Guojiang Li51College of Animal Science and Technology, Jilin Agricultural Science and Technology College, Jilin City, China 2Jilin Provincial Key Laboratory of Preventive Veterinary Medicine, Jilin City, China 3Jilin Province Technology Innovation Center of Pig Ecological Breeding and Disease Preventing and Control, Jilin City, China 4Jilin Province Cross Regional Cooperation Technology Innovation Center of Porcine Main Disease Prevention and Control, Jilin City, China 5College of Life Sciences, Changchun Sci-Tech University, Changchun, China *Corresponding Author: Wenlong Dong. College of Animal Science and Technology, Jilin Agricultural Science and Technology College, Jilin City, China. Email: dongwenlong888 [at] jlnku.edu.cn Submitted: 15/09/2025 Revised: 02/12/2025 Accepted: 16/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
AbstractBackground: Recently, the misuse of antibiotics has led to the rapid evolution of bacterial resistance, with bacteria of animal origin increasingly recognized as reservoirs of resistance genes. The atypical streptococcal species Streptococcus hyovaginalis, originally isolated from porcine vaginal specimens, has been increasingly identified in respiratory tract samples. Its recovery from diseased livestock suggests a broader clinical relevance than previously understood. This expanded host and tissue range necessitates a reassessment of its antibiotic resistance profiles. Its pathogenicity remains unclear, and its resistance profile has not been systematically studied. Given its extensive multidrug resistance, which poses a potential risk for the spread of resistance genes, and its ability to transmit across species, which may breach host barriers, in-depth research on this bacterium is particularly urgent. Aim: This study aims to systematically reveal the multidrug-resistant phenotypes of S. hyovaginalis strains in swine in Jilin Province, analyze their resistance gene repertoire, and explore their population genetic diversity through phylogenetic analysis. Methods: A total of 16 strains isolated from nasal swabs of 213 sick piglets between March and July 2023 were identified as S. hyovaginalis based on 16S rRNA gene sequencing. These isolates were subjected to antimicrobial susceptibility testing using the agar dilution method to determine minimum inhibitory concentrations against 21 antimicrobial agents. The study first conducted second-generation sequencing analysis and then comprehensively screened for resistance genes by searching the Comprehensive Antibiotic Resistance Database up to January 2025. Results: These isolates exhibited multidrug resistance phenotypes, demonstrating antibiotic resistance spanning resistance to antibiotics spanning multiple drug classes. High resistance rates were observed for β-lactam antibiotics: ampicillin (100% of isolates) and lincomycin (94%). Genomic analysis using second-generation sequencing revealed 31 distinct antimicrobial resistance genes among the isolates. Functional annotation identified five genes, such as lsa (E) and lnu (B), associated with lincosamide resistance and four genes, such as patB, conferring aminoglycoside resistance mechanisms. Conclusion: This study provides the first systematic characterization of MD Rmultidrug resistance in porcine S. hyovaginalis from Jilin Province, revealing a complex resistance gene composition. These findings establish a critical foundation for future mechanistic studies and underscore the potential role of this bacterium as a reservoir for disseminating resistance genes. Keywords: Streptococcus hyovaginalis, Pig, Resistance genes, Second-generation sequencing analysis. IntroductionStreptococcus spp. are ubiquitously distributed in natural environments and typically colonize the cutaneous and mucosal surfaces of the respiratory, gastrointestinal, and urogenital systems in both humans and animals (Costinar et al., 2024). While predominantly existing as commensal organisms, certain Streptococcal species may exhibit opportunistic pathogenicity under specific conditions, potentially leading to clinical infections (Costinar et al., 2024). Recent advances in microbiological detection techniques have facilitated extensive characterization of various zoonotic Streptococcus species. Costinar et al. (2024) systematically investigated emerging pathogens such as Streptococcus canis, Streptococcus iniae, and Streptococcus porcinus (Costinar et al., 2024). Among these isolated streptococcal species, Streptococcus hyovaginalis, initially discovered in porcine genital tract specimens in 1997 (Devriese et al., 1997), stands out as an atypical streptococcal species. Current epidemiological data indicate that S. hyovaginalis mainly colonizes the vaginal and nasal mucosal surfaces of pigs, with occasional detection in cerebral mucosal tissues. However, the colonization of articular mucosal surfaces has not been documented (Renzhammer et al., 2020). In cases of bovine mastitis, S. hyovaginalis has been isolated not only from the central nervous system and vaginal secretions of affected pigs but also from inflammatory secretions (Moreno et al., 2016; Hassan et al., 2023). This suggests that the risk of cross-species transmission is potentially underestimated, which deserves further investigation. Furthermore, the isolates recovered from the respiratory tract in this study exhibited a high rate of antimicrobial resistance (AMR). This resistance appears to be directly related to the use of antimicrobial agents in the industry and could serve as a biomarker for resistance monitoring. AMR has emerged as a critical challenge in global healthcare systems. AMR is characterized by bacterial evolutionary adaptations that diminish therapeutic efficacy, thereby prolonging infection duration, increasing transmission risks, and elevating mortality rates (Hui-Jun et al., 2024). Currently, β-lactam antibiotics remain the first-line treatment for streptococcal infections, whereas macrolides are used as a second-line treatment option (Somogyi et al., 2023). This long-term reliance has led to a continuous rise in resistance rates among major pathogens, such as Streptococcus, which are typically part of the commensal microbiota in pigs (Costinar et al., 2024). Streptococcus hyovaginalis exhibits a high level of AMR, which may enable it to act as a reservoir of resistance genes, continuously providing resistance genes to other (pathogenic) bacteria. Therefore, S. hyovaginalis, a bacterium generally associated with cohabitation with streptococci in pigs, was specifically investigated as a potential resistance gene reservoir to reveal its public health risk. Epidemiological surveillance reveals that streptococci persist as the predominant pathogens in neonatal porcine infections. Although improved farm biosecurity measures have reduced clinical incidence, contemporary antimicrobial susceptibility data indicate alarming resistance trends (Segura et al., 2020). The scarcity of mechanistic studies on S. hyovaginalis resistance is particularly concerning, as it significantly limits comparative analyses. Materials and MethodsBacterial strainIn total, 16 strains (7.5%) were isolated from nasal swabs of 213 diseased piglets from 20 pig farms in Jilin Province from March to July 2023. The sick piglets exhibited respiratory symptoms, such as unsteady walking, coughing, and rapid breathing. These sampled piglets have all undergone antibiotic treatment; however, due to farm privacy concerns, the exact types and dosages of the drugs are currently unknown. We collected nasopharyngeal swabs from live diseased piglets and immediately placed them in sterile tubes, stored them at 4°C, and processed them within 12 hours. Samples were incubated in Brain Heart Infusion (BHI) agar medium from Qingdao Hope-Bio in China (HB8297-1) containing 5% fetal bovine serum at 37°C for 12 hours, and different colonies were selected (Su et al., 2025). These strains appeared as tiny, colorless, semi-transparent colonies with a diameter of 0.5 mm on BHI solid plates. Strain identificationThe DNA of the strain was extracted using the DNA extraction kit (DN1101) produced by Aidlab Biotechnologies Company Ltd (Beijing). Then, the temperature program for polymerase chain reaction (PCR) was as follows: heat at 94°C for 5 minutes, then heat at 95°C, 55°C, and 72°C for 1 minute each, and cycle 30 times. Finally, heat at 72°C for 10 minutes to complete the process. Examination of the PCR product band clarity was examined using 1% agarose gel electrophoresis to verify the quality of the obtained product. Samples that passed quality control were sent to GENERALBIOL Company Ltd (Anhui) for sequencing. Universal primers 27F (5′-AGAGTTTGATCCTGGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) were used to generate 16S rRNA gene amplicons, targeting the V3-V4 hypervariable regions (Sun et al., 2025). These amplicons were purified and sequenced bidirectionally using a Thermo Fisher Scientific 3730XL DNA Analyzer. Chromatograms from both sequencing directions were manually checked to obtain the final consensus sequence for strain identification. The GenBank databases were searched for sequences using the Basic Local Alignment Search Tool (BLAST) to identify the isolate strains. The 16S sequences obtained were used to construct a phylogenetic tree based on the isolates and selected species. After constructing the phylogenetic tree using sequences analyzed with MEGA 11.0, the Chiplot 2.6.1 was utilized to enhance and refine the appearance of the tree. Antimicrobial susceptibility assessmentAntimicrobial susceptibility experiments were conducted on these 16 strains to investigate antibiotic resistance (Su et al., 2025). The minimum inhibitory concentrations (MICs) of S. hyovaginalis were determined by agar dilution. Subsequent experiments were conducted using MH (Mueller-Hinton) medium (CN230697) (Ararat Biotechnology Company Ltd). The main steps of agar dilution for antimicrobial susceptibility testing consist of adjusting the drug concentration to 0.03 to 512 μg/ml (twofold dilution method: 0.03, 0.06, 0.125, 0.25, up to 512 μg/ml) and ensuring that the colony concentration is 1 × 107CFU/ml simultaneously. Finally, the strains were inoculated into a drug-containing medium and incubated at a constant temperature for 16 hours to observe the growth of the colonies. A total of 21 antibiotics were determined, including: amikacin, ciprofloxacin, vancomycin, lincomycin, trimethoprim, oxytetracycline, enrofloxacin, penicillin, sulfamethoxazole, erythromycin, ceftazidime, florfenicol, linezolid, tilmicosin, chloramphenicol, tetracycline, ampicillin, mupirocin, ceftizoxime, amoxicillin, and azithromycin. We adopted a hierarchical approach due to the incomplete establishment of veterinary-specific breakpoints for all antimicrobials (Kaczorek et al., 2017). For antimicrobials with defined criteria in CLSI VET09 (2024), these veterinary-specific breakpoints were prioritized. We referred to human-derived breakpoints from CLSI M100 (2025) or EUCAST (2025)for agents not covered by VET09, acknowledging the potential limitations of extrapolating from human medicine to porcine isolates. The specific standards applied to each drug are detailed in Figure 3.
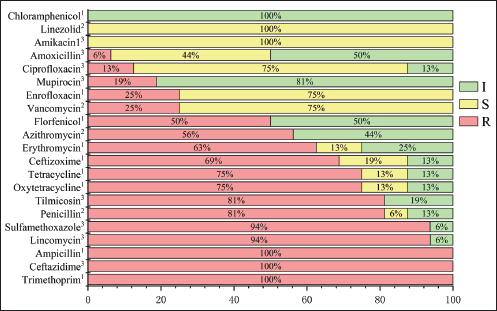
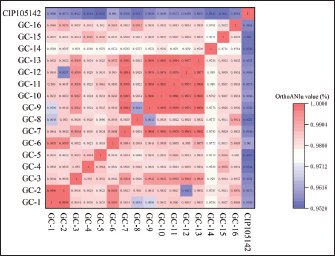
Fig. 3. Antibiotic resistance of S. hyovaginalis to 21 antibiotics The resistance of 16 S. hyovaginalis strains isolated from pig farms in Jilin Province to 21 antibiotics was analyzed. The red part represents resistance, the yellow part represents the intermediary, and the green part represents antibiotic susceptibility. The superscripts in the upper right corner of the antibiotics represent reference literature sources for the resistance breakpoints. Superscript 1: CLSI VET09 (2024 edition); superscript 2: CLSI M100 (2025 edition); superscript 3: EUCAST (2025 edition). All reference resistance breakpoints for Streptococcus species, with CLSI M100 specifically referencing resistance breakpoints for β-hemolytic Streptococcus. Sequence analysis of genes involved in antibiotic resistanceSecond-generation sequencing was used to acquire and analyze the sequences of antibiotic resistance genes. Bacteria were cultured to the logarithmic growth phase, colony concentration. Then, they were centrifuged at 7,000 rpm for 15 minutes to prepare the bacterial precipitate. DNA was extracted from the precipitate for analysis. The second-generation sequencing was performed by KAITAI-BIO (Hangzhou, China) using the Enzymic Universal DNAseq Library Prep Kit (KAITAI-BIO, AT4107) library preparation kit and AMPure XP beads for screening and recovery sequencing of 300—600 bp target products, first using NovaSeq 6000. After sequencing on the PE150 platform, the clean data were assembled using the unicycler software, and the preliminary assembly results were optimized, and the holes were filled to obtain the final assembly results. The Comprehensive Antibiotic Resistance Database was used for data comparison and analysis to predict the antibiotic resistance genes in the strains. After downloading the reference strain S. hyovaginalis (CIP105142) from the National Center for Biotechnology Information (NCBI) official website, the results of second-generation sequencing of this strain and the strains GC1-16 were uploaded to the EZEasyBioCloud website to calculate the average nucleotide identity (ANI) between each pair of strains (Chalita et al., 2024). Finally, Origin 2024 was used to create a heatmap (Fig. 2).
Fig. 2. Heatmap of the comparison of ANI among GC1-16 strains. The deeper the red color, the higher the ANI between the two strains, whereas the deeper the blue color, the lower the ANI between the compared strains. The ANI of strains GC-1 to GC-16 with the reference strain (CIP105142) from NCBI is above 0.95, which provides strong evidence that the isolated strains in the GC series are S. hyovaginalis. Ethical approvalSampling was conducted with the farm operators’ informed consent and was considered a routine veterinary inspection procedure, not an experimental animal study. All procedures strictly adhered to the animal welfare guidelines approved by the Animal Ethics Committee of Jilin Agricultural Science and Technology College (Ethics Approval No. LLSC202301055). ResultsPhylogenetic analysis and strain diversityFollowing PCR treatment, the 16S rRNA sequences of all strains were aligned with the sequences in the NCBI database using the BLAST program, revealing that the homology of all 16 study strains with the reference S. hyovaginalis strain was >97%. The 16S sequencing results of the strains have been uploaded to the NCBI database. The specific accession number is shown in Figure 1. ANI, which is strongly correlated with 16S rRNA gene similarity and is unaffected by horizontal gene transfer, provides a robust measure of genetic distance and offers subspecies-level resolution (Jain et al., 2018). Therefore, ANI values strongly support the taxonomic identification of the strains under study. The ANI values between the GC strains and the CIP105142 reference strain consistently exceeded 0.95 (Fig. 2). The key comparisons used for taxonomic identification, namely, the comparisons of each of our GC isolates with the S. hyovaginalis reference strain CIP105142, overall and consistently exceed the species threshold, with all exceeding 0.95 (95.1%), providing solid genomic evidence that all of our isolates belong to the species S. hyovaginalis and confirming the species identity of the strains.
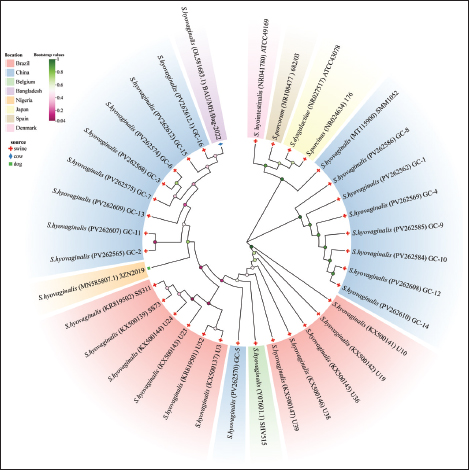
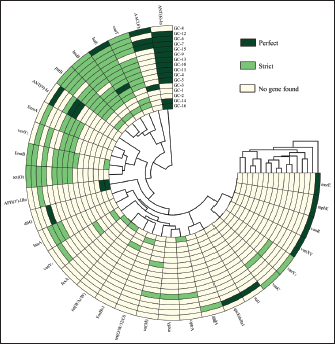
Fig. 1. Phylogenetic tree constructed based on 16S rRNA sequences of isolated strains and selected species. A total of 31 S. hyovaginalis strains, including 16 strains isolated from pigs in Jilin, China, were used to construct this phylogenetic tree. Four other Streptococcus species, including S. dysgalactiae, S. porcinus, S. porcorum, and S. hyointestinalis, were also included in the tree construction. These sequences represent strains isolated from different regions and sources. The color of the leaves indicates the regional origin of the strains, the shading of the circles on the branches represents bootstrap values, and different symbols between branches and strains indicate the strains’ species origin. Phylogenetic tree constructed based on the 16S rRNA sequences of isolates and selected species. This study constructed a phylogenetic tree based on the 16S rRNA sequences of the isolates and selected reference species, including a total of 31 pig-derived S. hyovaginalis strains, of which 16 were isolates from Jilin Province, China. To further clarify the phylogenetic position of S. hyovaginalis, four other streptococci isolated by scholars such as PerezSancho from the same clinical samples—Streptococcus dysgalactiae, S. porcinus, Streptococcus porcorum, and Streptococcus hyointestinalis—were selected as outgroup comparators using standard strains (sequences obtained from the NCBI database) (Pérez-Sancho et al., 2017). Phylogenetic analysis showed that S. hyovaginalis exhibited distinct differentiation characteristics during evolution. Notably, GC-1, GC-4, GC-8, GC-9, GC-10, GC-12, and GC-14 clustered into a highly supported monophyletic group (bootstrap value=1), suggesting a close genetic relationship among these strains. Strains GC-2, GC-3, GC-6, GC-7, GC-11, GC-13, GC-15, and GC-16 were positioned on branches with lower support, reflecting a certain degree of genetic diversity and evolutionary divergence within S. hyovaginalis. Notably, GC-5 was independent of these two branches, which may represent genetic variation during dissemination. Even strains isolated from the same region exhibited significant phylogenetic relatedness. Although S. hyovaginalis primarily hosts pigs, its phylogenetic structure suggests the potential for cross-species transmission. In the phylogenetic tree, S. dysgalactiae, S. porcinus, S. porcorum, and S. hyointestinalis formed distinct branches from S. hyovaginalis, further confirming that the strains isolated in this study are phylogenetically distant from these four Streptococcus species. Notably, the phylogenetic tree shows that one S. hyovaginalis strain from the NCBI database, SMM1052 (MT115900), clustered with reference strains such as S. dysgalactiae. This is likely due to historical mislabeling in public databases or a rare horizontal gene transfer event of the 16S rRNA gene within the genus Streptococcus. However, this isolated occurrence does not affect the phylogenetic independence of the main branches formed by the core isolates in this study. The phylogenetic tree construction results provide reliable support for the accurate identification of strains at the molecular phylogenetic level and reinforce the evolutionary independence and specificity of S. hyovaginalis. Determination of antimicrobial susceptibilityFigure 3 shows the AMR profiles of 16 S. hyovaginalis strains isolated from porcine sources. All isolates demonstrated complete resistance to ampicillin, oxytetracycline, and trimethoprim (100%). High-level resistance was observed for sulfamethoxazole and lincomycin (93.75% each), followed by tilmicosin and penicillin (81.25% each). Substantial resistance rates were also recorded for tetracycline (75%), ceftizoxime (68.75%), erythromycin (62.50%), and azithromycin (56.25%). Florfenicol (50%) and ceftazidime (43.75%) showed moderate resistance, whereas enrofloxacin and vancomycin (25% each), mupirocin (18.75%), and ciprofloxacin (12.5%) showed lower resistance. Notably, the isolates exhibited the lowest resistance rates (6.25%) to chloramphenicol and linezolid. Analysis of antibiotic-resistant genesA total of 31 antibiotic resistance genes were identified in 16 S. hyovaginalis isolates (Fig. 4). Strains GC-8 and GC-12 exhibited the highest genetic load with 13 resistance genes each. GC-8 carried the following genes: AAC(6’), vanT, lsa(E), lnu(B), patB, erm(B), tet(O), msrE, mphE, vanR, vanXY, vanY3, and vanC. GC-12 demonstrated a distinct resistance profile containing: ANT(6)-Ia, vanT, lsa(E), lnu(B), patB, ANT(9)-Ia, erm(A), vanY2, erm(B), tet(O), aadA, qacEdelta1, and sul1. The predominant resistance determinants were lincosamide-related genes lsa(E) (81.25%) and lnu(B) (75%), followed by glycopeptide resistance gene vanT (75%) and aminoglycoside resistance gene ANT(6)-Ia (75%). The fluoroquinolone efflux pump gene patB also showed a prevalence of 75%. Secondary prevalent genes included erm(B) (56.25%) and tet(O) (50%), while vanY2 and erm(A) each occurred in 31.25% of isolates. Lower detection rates were observed for: ANT(9)-Ia (25%), AAC(6’) (25%), vanY1 (18.75%), APH(3’)-IIIa (18.75%), lnu(A) (18.75%), vanY3 (12.5%), optrA (12.5%), vanC (12.5%), tet(M) (12.5%), vanXY (6.25%), vanR (6.25%), mphE (6.25%), dfrG (6.25%), fexA (6.25%), tet(W/N/W) (6.25%), FosBx1 (6.25%), tet(O/W/32/O) (6.25%), mefA (6.25%), aadA (6.25%), qacEdelta1 (6.25%), sul1 (6.25%), and msrE (6.25%).
Fig. 4. Resistance genes of 16 S. hyovaginalis strains in pigs This figure shows the carrying of resistance genes in 16 S. hyovaginalis strains in Jilin Province, China. Dark green indicates perfect resistance genes, light green indicates strict resistance genes, and yellow indicates no resistance genes. The vanY1 represents vanY gene in the vanB cluster, vanY2 represents the vanY gene in the vanG gene cluster, and vanY3 represents the vanY gene in the vanM gene cluster. The vanT gene is on the gene cluster of vanG, whereas both the vanR and vanXY genes are on the gene cluster of vanC. AAC(6’)-Ie-APH(2’’)-Ia is a bifunctional protein. Among the isolated strains, GC-8 and GC-12 contain the most antibiotic resistance genes. Due to their different antibiotic resistance genes, their drug susceptibility is also slightly different. In addition to vancomycin, florfenicol, linezolid, chloramphenicol, and mupirocin, GC-12 is sensitive to enrofloxacin and amoxicillin. GC-8 is also sensitive to sulfamethoxazole. GC-7 is resistant to only four drugs: linezolid, mupirocin, chloramphenicol, and florfenicol. DiscussionDrug susceptibility analysisA comprehensive antimicrobial susceptibility test was conducted on 16 porcine-derived S. hyovaginalis strains. All tested strains exhibited a 100% resistance rate to ampicillin, oxytetracycline, and trimethoprim. Resistance rates to sulfamethoxazole and lincomycin were also extremely high, reaching 93.75%. These findings reveal a severe multidrug-resistant phenotype, suggesting that these strains may carry and stably inherit resistance mechanisms against aminoglycosides, β-lactams, tetracyclines, and lincosamides. Notably, there was a significant variation in the resistance to certain antibiotics. For example, the resistance rates to tiamulin and penicillin were 81.25% and 81.25%, respectively (with penicillin showing 6.25% intermediate resistance), whereas resistance to tetracyclines, cefalexin, and macrolides (such as erythromycin and tilmicosin) ranged from 56.25% to 75%. Of particular concern were florfenicol and ceftazidime, with roughly equal proportions of resistant and susceptible strains, indicating that resistance mechanisms for these drugs may not yet be fixed in the population or may have a heterogeneous genetic background. In contrast, the strains were primarily sensitive to fluoroquinolones (such as enrofloxacin and ciprofloxacin) and mupirocin, with sensitivity rates ranging from 75% to 81.25%. They also showed near-complete or very high sensitivity to amikacin, chloramphenicol, and linezolid (87.5%–100%). The above phenotypic resistance patterns strongly indicate that these strains’ genomes harbor complex and diverse resistance genes. Antibiotic resistance gene profiles and genotype-phenotype associationsThe lsa gene, which belongs to the efflux pump superfamily, utilizes dual ATP-binding domains to export antimicrobial agents, including macrolides, lincosamides, and streptogramins. This multidrug efflux mechanism may result in coresistance to clindamycin, lincomycin, and dalfopristin (Salah et al., 2021). The lsa family consists of four isoforms (lsaA, B, C, and E) (Li BeiBei et al., 2013) and exhibits distinct AMR profiles. In particular, the lsa(E) variant has been molecularly characterized in clinically significant pathogens such as Staphylococcus spp., Enterococci, and Streptococcus agalactiae (Li BeiBei et al., 2013; Sanderson et al., 2020). Epidemiological investigations have identified lsa(E) as the predominant genetic determinant of macrolide resistance in Enterococcus, with 99.8% of resistant isolates harboring this gene (Chen et al., 2018). Originally identified in Streptococcus pluranimalium, this resistance determinant has demonstrated significant efficacy against lincomycin in experimental studies ((Xu et al., 2023)). Studies have shown that the resistance genes lsaE, lnuB, ant(6)-Ia, and ant(9)-Ia on Streptococcus suis coexist within a region of approximately 10 Kb, which is highly similar in nucleotide sequence to enterococcal fecal plasmids. The sequence identity of the lsaE gene reaches as high as 98% (Nicholson and Bayles, 2022), indicating that lsaE and other genes indeed exist in a mobile genetic context. The erm gene was initially identified as a widely distributed genetic element in environmental bacterial populations (Roberts, 2011). The relative abundance of erm(B) is significantly positively correlated with the use of lincosamides and macrolides (Yang et al., 2022). Resistance to macrolides is primarily mediated by ribosomal methylases encoded by erm genes, whereas it is less commonly mediated by large efflux pumps encoded by mef genes (Yang et al., 2022). The erm(B) gene is the most prevalent resistance gene among S. suis strains isolated from locally bred and commercial pigs in Jiangxi, China (Tan et al., 2025). In other countries, the erm(B) gene is widely carried by pathogens isolated from pigs (Muwonge et al., 2025). This may be related to the frequent use of macrolides as second-line antibiotics in veterinary practice. Researchers have also noted that the use of antibiotics in farming is closely linked to the development of AMR (Muwonge et al., 2025). Correspondingly, in the strains isolated in this study, the carriage rate of erm(B) (56.25%) is also considerable, which may contribute to the relatively high erythromycin resistance rate (62.50%) observed in these strains. Tet(M) and tet(O) proteins have been extensively characterized as ribosomal protection proteins with demonstrated ribosome-dependent GTPase activity (Hormeño et al., 2020). Analysis of 78 S. agalactiae strains isolated in Zimbabwe revealed that 94.5% of tetracycline-resistant isolates harbored the tet(M) gene and 3.9% contained the tet(O) gene (Mudzana et al., 2021). Notably, all erm(B)-positive isolates (100%) exhibited co-occurrence with tet(M), although no significant genetic linkage was observed. In this study, 50% of S. hyovaginalis isolates tested positive for the tet(O) gene. The elevated prevalence of resistance genes likely contributed to the observed high resistance rates to oxytetracycline (100%) and tetracycline (75%) in these strains. The analyzed strains harbored lincosamide resistance determinants, including lnu(A) and lnu(B), which demonstrated in vitro antibiotic inactivation via adenylylation. Erm gene-mediated resistance to macrolides and lincosamides was also observed. Most of these lincosamide resistance genes are plasmid- or transposon-associated, thereby facilitating their horizontal transfer across strain, species, and genus boundaries (Yang et al., 2024). In our cohort of 16 S. hyovaginalis isolates, lsa(E) exhibited the highest carriage rate (81.25%), followed by lnu(B) (75%) and lnu(A) (18.75%). Regarding erm gene distribution, the carriage rates were quantified as erm(A) (31.25%) and erm(B) (56.25%). The cumulative effect of lsa(E), lnu(A), lnu(B), erm(A), and erm(B) on lincomycin resistance mechanisms likely accounts for the observed 93.75% lincomycin resistance rate. This phenomenon is consistent with the findings in S. pneumoniae, where the coordinated expression of patA and patB genes mediated quinolone resistance (Emilie et al., 2012). Through heterologous expression in Escherichia coli, the functional relationship between genes and AMR was systematically investigated. Functional characterization using inside-out vesicles demonstrated that the activity and drug transport capacity of pervanadate-sensitive ATPase were strictly dependent on the co-expression of PatA and PatB proteins. While PatA and PatB demonstrated limited homodimerization capacity, neither homodimeric form exhibited detectable drug transport or ATPase activity, in sharp contrast to the functional heterodimeric complex. Collectively, these findings indicate that the PatA−PatB heterodimer is essential for constituting a functional multidrug efflux complex in vitro (Emilie et al., 2012). However, the genomic analysis of 16 S. hyovaginalis isolates from swine revealed the exclusive presence of the patB gene (75% carriage rate), which correlates with the observed low resistance rate to quinolones (e.g., enrofloxacin, 25%). These strains were susceptible to amikacin, a structurally related aminoglycoside. This phenomenon may be attributed to the exclusive dependence of amikacin inactivation on AAC(6’)-mediated enzymatic modification, which was detected in only 25% of isolates, thereby explaining the prevalence of amikacin sensitivity within this cohort. Functional vancomycin tolerance requires the cooperative activity of three encoded proteins: VanC-1, VanXYC, and VanT. Specifically, VanT catalyzes the conversion of L-serine to D-serine for autopeptidoglycan biosynthesis (Li et al., 2022). Despite the high prevalence of VanT (75%) in our isolates, the absence of VanC-1 and VanXYC genes resulted in limited resistance to vancomycin (25%). Phylogenetic analysis and species specificityThe phylogenetic tree in Figure 1 and the resistance gene circular map in Figure 4, GC-9 is distantly related to GC-13 and GC-5, yet it still carries the same resistance gene ANT-9 (Ia). The prevalence of this gene in other S. hyovaginalis strains is not high, effectively ruling out the possibility of vertical transmission of the resistance gene. ANT-9 (Ia) was first detected in S. aureus and was subsequently confirmed to be a resistance gene encoded on plasmids or transposable elements (Sheng et al., 2023), reasonably suggesting the occurrence of horizontal transfer of this resistance gene within the S. hyovaginalis population, indicating that the resistance genes carried by S. hyovaginalis may be horizontally transferred through mobile genetic elements. Similarly, GC-6 and GC-7 are on the same branch, but GC-8, which is on a different branch, is more distantly related; all carry the AAC(6’) gene, which belongs to the aminoglycoside N-acetyltransferase family. Genes of this family, particularly the common AAC(6’)-Ib and its variants, have been widely reported to be located on mobile genetic elements such as integrons, transposons, and plasmids (Ramirez et al., 2013). This genetic context strongly suggests that this resistance gene possesses a high potential for horizontal transfer, enabling its dissemination among different bacteria and potentially coexisting with other resistance genes, thereby collectively contributing to MDR phenotypes. Although clinical strains capable of confirming its pathogenicity have not yet been isolated, research indicates that the colonization of methicillin-resistant Staphylococcus aureus (MRSA) in the porcine upper respiratory tract is positively correlated with the colonization of S. hyovaginalis (Schlattmann et al., 2020). Moreover, S. hyovaginalis frequently coexists within the host alongside various multidrug-resistant Streptococcus species as Streptococcal clusters (Costinar et al., 2024). The isolated S. hyovaginalis strains exhibit high levels of antibiotic resistance—although this may be because S. hyovaginalis inherently possesses high resistance, allowing it to survive in pigs undergoing antibiotic treatment and co-colonize with MRSA or multidrug-resistant porcine Streptococcus species, S. hyovaginalis may also act as a conduit or reservoir for the dissemination of resistance genes. Limitations of research and future prospectsThe MICs of most pathogens in veterinary medicine are not as well established as they are in human medicine (Kaczorek et al., 2017). Therefore, breakpoints often need to be inferred from other animal species, different bacterial groups, or human medical standards. Accurately assessing the level of antibiotic resistance in veterinary pathogens becomes highly challenging. Monitoring the phenotypic (especially MIC values) and genotypic resistance profiles of pathogens isolated from various disease cases worldwide based on pharmacokinetic data will help researchers standardize or establish interpretive frameworks for susceptibility testing results. Accordingly, our study focuses on S. hyovaginalis isolated from pigs in northeastern China. To the best of our knowledge, studies on the resistance and resistance gene phenotypes of S. hyovaginalis in China remain limited. Although this study reveals its genetic diversity, the specific molecular and ecological mechanisms driving cross-species transmission and host adaptation of S. hyovaginalis require further investigation. Moreover, due to technical limitations, the genomic locations of resistance genes in the S. hyovaginalis GC series strains cannot yet be determined and can only be inferred based on fundamental research on resistance genes. The data provided in this study are intended for academic discussion and research purposes. It is not suitable as a reference for the clinical use of antibiotics to treat S. hyovaginalis. This retrospective study aimed to collect S. hyovaginalis strains isolated from pig farms to expand the understanding of the potential transmission risk of antibiotic resistance in S. hyovaginalis. Despite the lack of comprehensive information on other diseases in the sampled pig farms and previous antimicrobial treatments, we still consider this study worthy of publication because there are currently no available data on S. hyovaginalis in the intensive farming systems of northeastern China. Research on S. hyovaginalis strains remains at the basic level, with a focus on strain types and resistance phenotypes. This study explored the antibiotic resistance genes of S. hyovaginalis and provided preliminary inferences on the potential mechanisms of resistance gene transmission. It reveals previously unexplored areas at the genetic level regarding the multidrug resistance of S. hyovaginalis, and therefore, it is necessary to draw the attention of the academic community. Streptococcus hyovaginalis not only has the potential for cross-species transmission but may also serve as an important reservoir for AMR genes. Currently, PCR detection technology is still used to identify S. hyovaginalis. There is an urgent need to develop detection technologies that can accurately and quickly identify organisms at the species level to achieve precise identification of bacteria in complex samples. For example, nano-biosensors, with their high specificity and sensitivity, can clearly distinguish different species within the same genus in complex samples, providing a reliable solution for this purpose (Ayse et al., 2025). ConclusionThis study demonstrated that S. hyovaginalis from pigs has multi-antibiotic resistance, and 31 antibiotic resistance genes were identified in 16 S. hyovaginalis strains after second-generation sequencing analysis. This paper lays the foundation for a follow-up study on the mechanisms of antibiotic resistance and resistance genes in S. hyovaginalis. AcknowledgmentsWe extend our appreciation to the Jilin Province Science and Technology Innovation Platform Subsidy Project and the staff members of the Jilin Agricultural Science and Technology College. Conflict of interestThe authors declare that they have no conflicts of interest to declare. FundingFunding for this research was provided by Jilin Province Science and Technology Innovation Platform Subsidy Project (Grant number YDZJ202302CXJD038). Authors’ contributionsWriting - original draft: Zhuolei Yang. Visualization: Jingyu Gao. Data analysis: Nuo Xu. Methodology: Sijia Sun. Formal analysis: Lvpeng Jiang. Supervision: Wanli Sha, Baishuang Yin. Writing - review & editing: Wenlong Dong, Guojiang Li. Data availabilityData from the results of this study are not publicly available due to sensitivity. However, they can be obtained from the corresponding author upon reasonable request. Data are stored in a controlled-access data storage facility at Jilin Agricultural Science and Technology College. ReferencesAyse, B., Mehmet, F.B., Polat, I., Aziz, E., Cumali, K., Mehmet, N.A. and Mehmet, H.A. 2025. Green-synthesized nanoparticles for biomedical sensor technology nanosensors for healthcare diagnostics. Mardin Province, Turkey: Academic Press, pp: 355–80. Chalita, M., Kim, Y.O., Park, S., Oh, H.S., Cho, J.H., Moon, J., Baek, N., Moon, C., Lee, K., Yang, J., Nam, G.G., Jung, Y., Na, S.I., Bailey, M.J. and Chun, J. 2024. EzBioCloud: a genome-driven microbiome identification and discovery platform. Int. J. Syst. Evol. Microbiol. 74(6), 6421. Chen, M., Pan, H., Lou, Y., Wu, Z., Zhang, J., Huang, Y., Yu, W. and Qiu, Y. 2018. Epidemiological characteristics and genetic structure of linezolid-resistant E. faecalis. Infect. Drug. Resist. 11, 2397–2409. Costinar, L., Badea, C., Marcu, A., Pascu, C. and Herman, V. 2024. Multiple drug-resistant Streptococcus Strains—an actual problem in pig farms in Western Romania. Antibiotics 13(3), 277. Devriese, L.A., Pot, B., Vandamme, P., Kersters, K., Collins, M.D., Alvarez, N., Haesebrouck, F. and Hommez, J. 1997. S. hyovaginalis sp. nov. and S. thoraltensis sp. nov., from the genital tract of sows. Int. J. Syst. Evol. Microbiol. 47(4), 1073–1077. Emilie, B., Claire, D., Benoît, B., Christine, E., Marie, D.G.A., Jacques, C., Thierry, V. and Jean-Michel, J. 2012. PatA and PatB form a functional heterodimeric ABC multidrug efflux transporter responsible for S. pneumoniae fluoroquinolone resistance. Biochemistry. 51(39), 7755-7765. Hassan, J., Bag, M.A.S., Ali, M.W., Kabir, A., Hoque, M.N., Hossain, M.M., Rahman, M.T., Islam, M.S. and Khan, M.S.R. 2023. Streptococcus spp. diversity and Streptococcus uberis genomic characteristics isolated from clinical mastitis cattle in Bangladesh. Front. Vet. Sci. 10, 1198393. Hormeño, L., Campos, M.J., Vadillo, S. and Quesada, A. 2020. Occurrence of tet (O/M/O) mosaic gene in tetracycline-resistant Campylobacter. Microorganisms 8(11), 1710–1717; doi: 10.1016/j.microorganisms.2011.09.010 Hui-Jun, B., Qing-Feng, G., Fang, J. and Yong-Li, Y. 2024. Epidemiologic analysis of antimicrobial resistance in hospital departments in China from 2022 to 2023. J Health Popul Nutr. 43(1), 38-39. Jain, C., Rodriguez-R, L.M., Phillippy, A.M., Konstantinidis, K.T. and Aluru, S. 2018. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nature Commun. 9(1), 5114. Kaczorek, E., Małaczewska, J., Wójcik, R., Rękawek, W. and Siwicki, A.K. 2017. Phenotypic and genotypic antimicrobial susceptibility pattern of Streptococcus spp. isolated from cases of clinical mastitis in dairy cattle in Poland. J. Dairy Sci. 100(8), 6442–6453. Li, B., Wendlandt, S., Yao, J., et al. 2013. Detection and new genetic environment of the pleuromutilin-lincosamide-streptogramin A resistance gene lsa(E) in methicillin-resistant Staphylococcus aureus of swine origin. J Antimicrob Chemother. 68(6), 1251-1255. Li, G., Walker, M.J. and De Oliveira, D.M.P. 2022. Vancomycin resistance in Enterococcus and Staphylococcus aureus. Microorganisms 11(1), 24. Moreno, C., Romero, J. and Espejo, R.T. 2002. Polymorphism in repeated 16S rRNA genes is a common property of the genus’ type strains and environmental isolates. Vibrio. Microbiol. 148(4), 1233–1239. Moreno, L.Z., Matajira, C.E.C., Gomes, V.T.M., Silva, A.P.S., Mesquita, R.E., Christ, A.P.G., Sato, M.I.Z. and Moreno, A.M. 2016. Molecular and antimicrobial susceptibility profiling of atypical Streptococcus species from porcine clinical specimens. Infect. Genet. Evol. 44, 376–381. Mudzana, R., Mavenyengwa, R.T. and Gudza-Mugabe, M. 2021. Analysis of virulence factors and antibiotic resistance genes in group B streptococcus from clinical samples. BMC. Infect. Dis. 21, 125; doi:10.1016/j.bmcd.2008.08.010 Muwonge, A., Kakooza, T., Johnson, P.C., Kisuule, L., Kimaanga, M., Kankya, C., Bronsvoort, B.M.D.C. and Lembo, T. 2025. Drivers of antimicrobial resistance in Ugandan pig production systems. Commun. Earth. Environ. 6, 517. Nicholson, T.L. and Bayles, D.O. 2022. Comparative virulence and antimicrobial resistance distribution of Streptococcus suis isolates obtained from the United States. Front. Microbiol. 13, 1043529. Pérez-Sancho, M., Vela, A.I., García-Seco, T., González, S., Domínguez, L. and Fernández-Garayzábal, J.F. 2017. Usefulness of MALDI-TOF MS as a diagnostic tool for the identification of Streptococcus species recovered from clinical specimens of pigs. PLos One. 12(1), e0170784. Ramirez, M.S., Nikolaidis, N. and Tolmasky, M.E. 2013. Rise and dissemination of aminoglycoside resistance: the ac (6′)-Ib paradigm. Front. Microbiol. 4, 121. Renzhammer, R., Loncaric, I., Ladstätter, M., Pinior, B., Roch, F.F., Spergser, J., Ladinig, A. and Unterweger, C. 2020. Detection of various Streptococcus spp. and their antimicrobial resistance patterns in Austrian swine stock clinical specimens. Antibiotics 9(12), 893. Roberts, M.C. 2011. Environmental macrolide–lincosamide–streptogramin and TC-resistant bacteria. Front. Microbiol. 2, 40. Salah, A.N., Elleboudy, N.S., El-Housseiny, G.S. and Yassien, M.A. 2021. Cloning and sequencing of the lsaE efflux pump gene from MDR Enterococci and its role in ER Infect. Genet. Mol. Res. 94, 105010. Sanderson, H., Ortega-Polo, R., Zaheer, R., Goji, N., Amoako, K.K., Brown, R.S., Majury, A., Liss, S.N. and Mcallister, T.A. 2020. Comparative genomics of multidrug-resistant Enterococcus spp. isolated from wastewater treatment plants. BMC. Microbiol. 20(1), 1–7. Schlattmann, A., Von Lützau, K., Kaspar, U. and Becker, K. 2020. The porcine nasal microbiota with particular attention to livestock-associated methicillin-resistant Staphylococcus aureus in Germany — A culturomic approach. Microorganisms 8, 514. Segura, M., Aragon, V., Brockmeier, S.L., Gebhart, C., Greeff, A.D., Kerdsin, A., O’Dea, M. A., Okura, M., Saléry, M., Schultsz, C., Peter, V.W., Lucy, A.W., Jerry, M.W. and Marcelo, G. 2020. Streptococcus suis research and prevention in the era of antimicrobial restriction. 4th International Workshop on S. suis. Montreal, Canada. 9(5), 374. Sheng, X., Lu, W., Li, A., Lu, J., Song, C., Xu, J., Dong, Y., Fu, C., Lin, X., Zhu, M., Bao, Q. and Li, K. 2023. ANT(9)-Ic, a novel chromosomal encoded aminoglycoside nucleotidyltransferase from Brucella intermedia. Microbiol. Spectr. 11(9), 620–623. Somogyi, Z., Mag, P., Simon, R., Kerek., Makrai, L., Biks, I. and Jerzsele. 2023. Susceptibility of Actinobacillus pleuropneumoniae isolated from pigs in Hungary between 2018 and 2021. Antibiotics 12(8), 1298. Su, H., Yang, L., Liu, M., et al. 2025. Investigation of genomic islands involved in antimicrobial resistance in Proteus mirabilis isolated from swine. J. Glob. Antimicrob. Resist. 44, 160-165. Sun, Y., Wang, Q., Qiu, S., Chen, C., Wu, S., Gao, N. and Dai, J. 2025. Wuhapuella carnivora gen. Nov., sp. nov., a novel member of the family Weeksellaceae, a flocculent bacteria isolated from an urban sewage treatment plant. Curr. Microbiol. 82(11), 1–11. Tan, M.F., Liu, C.L., Zhou, Q.Y., Wei, J.Z., Hong, J.W., Wan, M.C., Zhang, F.L. and Ji, H. 2012. Prevalence and antimicrobial resistance of S. suis isolated from local pig breeds in Jiangxi Province, China. Front. Vet. Sci. 12, 1582461. Xu, C.W., Zhou, X., Zhang, X.L., et al. 2023. Whole genome sequence of Streptococcus pluranimalium SP21-2, a porcine strain harbouring optrA and lsa(E) with chromosomal location. J. Glob. Antimicrob. Resist. 35, 101–103. Yang, D., Heederik, D.J., Mevius, D.J., Scherpenisse, P., Luiken, R.E., Van Gompel, L., Skarżyńska, M., Wadepohl, K., Chauvin, C., Van Heijnsbergen, E. and Wouters, I.M. 2022. Risk factors for the abundance of antimicrobial resistance genes aph (3′)-III, erm (B), sul2, and tet (W) in pig and broiler feces in nine European countries. J. Antimicrob. Chemother. 77(4), 969–978; doi:10.1016/j.jantimicrob.2012.09.010 Yang, Z., Lan, T., Luo, H., Li, P., Wang, M., Jia, R., Chen, S., Liu, M., Zhao, X., Yang, Q., Wu, Y., Zhang, S., Huang, J., Ou, X., Mao, S., Gao, Q., Sun, D., Tian, B., Cheng, A. and Zhu, D. 2024. Emergence and mobilization of a novel lincosamide resistance gene lnu(I): from environmental reservoirs to pathogenic bacteria. Sci. Total Environ. 906, 167400. | ||
| How to Cite this Article |
| Pubmed Style Yang Z, Gao J, Jiang L, Xu N, Sun S, Sha W, Yin B, Dong W, Li G. Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Vet. J.. 2026; 16(1): 71-80. doi:10.5455/OVJ.2026.v16.i1.6 Web Style Yang Z, Gao J, Jiang L, Xu N, Sun S, Sha W, Yin B, Dong W, Li G. Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. https://www.openveterinaryjournal.com/?mno=283847 [Access: January 31, 2026]. doi:10.5455/OVJ.2026.v16.i1.6 AMA (American Medical Association) Style Yang Z, Gao J, Jiang L, Xu N, Sun S, Sha W, Yin B, Dong W, Li G. Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Vet. J.. 2026; 16(1): 71-80. doi:10.5455/OVJ.2026.v16.i1.6 Vancouver/ICMJE Style Yang Z, Gao J, Jiang L, Xu N, Sun S, Sha W, Yin B, Dong W, Li G. Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Vet. J.. (2026), [cited January 31, 2026]; 16(1): 71-80. doi:10.5455/OVJ.2026.v16.i1.6 Harvard Style Yang, Z., Gao, . J., Jiang, . L., Xu, . N., Sun, . S., Sha, . W., Yin, . B., Dong, . W. & Li, . G. (2026) Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Vet. J., 16 (1), 71-80. doi:10.5455/OVJ.2026.v16.i1.6 Turabian Style Yang, Zhuolei, Jingyu Gao, Lvpeng Jiang, Nuo Xu, Sijia Sun, Wanli Sha, Baishuang Yin, Wenlong Dong, and Guojiang Li. 2026. Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Veterinary Journal, 16 (1), 71-80. doi:10.5455/OVJ.2026.v16.i1.6 Chicago Style Yang, Zhuolei, Jingyu Gao, Lvpeng Jiang, Nuo Xu, Sijia Sun, Wanli Sha, Baishuang Yin, Wenlong Dong, and Guojiang Li. "Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China." Open Veterinary Journal 16 (2026), 71-80. doi:10.5455/OVJ.2026.v16.i1.6 MLA (The Modern Language Association) Style Yang, Zhuolei, Jingyu Gao, Lvpeng Jiang, Nuo Xu, Sijia Sun, Wanli Sha, Baishuang Yin, Wenlong Dong, and Guojiang Li. "Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China." Open Veterinary Journal 16.1 (2026), 71-80. Print. doi:10.5455/OVJ.2026.v16.i1.6 APA (American Psychological Association) Style Yang, Z., Gao, . J., Jiang, . L., Xu, . N., Sun, . S., Sha, . W., Yin, . B., Dong, . W. & Li, . G. (2026) Evaluation of the antimicrobial susceptibility and resistance gene mapping of Streptococcus hyovaginalis from piglets in Jilin Province, China. Open Veterinary Journal, 16 (1), 71-80. doi:10.5455/OVJ.2026.v16.i1.6 |