| Research Article | ||
Open Vet. J.. 2025; 15(12): 6237-6244 Open Veterinary Journal, (2025), Vol. 15(12): 6237-6244 Research Article Detection of virulence factors of Escherichia Coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, SyriaAseem Albaker1*, Maher Saleh1 and Ashraf Alsaleh21Department of Public Health and Preventive Medicine, College of Veterinary Medicine, University of Hama, Hama, Syria 2Department of Microbiology, College of Veterinary Medicine, University of Hama, Hama, Syria *Corresponding Author: Aseem Albaker. Department of Public Health and Preventive Medicine, College of Veterinary Medicine, University of Hama, Hama, Syria. Email: aseem7g41 [at] gmail.com Submitted: 22/08/2025 Revised: 08/11/2025 Accepted: 15/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
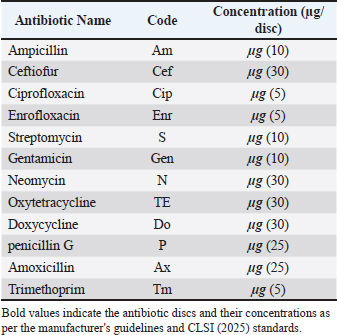
AbstractBackground: Diarrhea poses a major threat to calf herds, causing severe economic losses. Escherichia coli is one of the most important causes of diarrhea in animals, such as Enterotoxogenic E. coli (ETEC), Shiga toxin-producing E. coli (STEC), Enteropathogenic E. coli (EPEC), and Enterohemolytic E. coli (EHEC). Multidrug-resistant E. coli is emerging as a serious public health hazard. Aim: This study aimed to detect some virulence factors in Escherichia coli (sta, eae, stx1, and stx2) and to detect the antibiotic resistance rates of these bacterial strains. Methods: Escherichia coli strains were isolated from 60 diarrheic calves on MacConkey and Eosin Methylene Blue agar, following Gram staining. Biochemical characterization for the subsequent steps was performed using the Hi25™ Enterobacteriaceae Identification Kit (KB003). Multiplex polymerase chain reaction was employed to detect virulence genes (sta, eae, stx1, stx2). Antimicrobial susceptibility testing was performed using the disk diffusion method. Results: Escherichia coli was found in 53 of 60 diarrheic calves. Screening for virulence-associated genes revealed the following pathotype distribution: ETEC (carrying sta gene) 16.98%, EPEC (carrying eae gene) 13.21%, EHEC (carrying stx + eae genes) 13.21%, STEC carrying stx1 and/or stx2 only) 32.07%, and hybrid strains (carrying mixed virulence genes from multiple pathotypes): 5.66%, while nonpathogenic strains (lacking all tested virulence genes) 18.87%. Regarding antimicrobial susceptibility testing, the study demonstrated high resistance to penicillin-G (92.5%) and amoxicillin (88.7%), while showing the highest sensitivity to ceftiofur (82.1%) and ciprofloxacin (79.2%). Conclusion: The pathogenic E. coli strains represent one of the most significant causative agents of diarrhea in the calf. Furthermore, the emergence of multidrug-resistant bacterial strains in animals poses a potential threat to human public health. Keywords: Escherichia coli, Calves diarrhea, Virulence factors, Antibiotic resistance. IntroductionDiarrhea is one of the most critical veterinary health issues and a leading cause of high mortality rates in newborn calves during the suckling period. It leads to generalized weakness, reduced vitality, and poor health, making affected calves more susceptible to secondary infections that may result in death (Foster and Smith, 2009). Calf diarrhea continues to plague livestock operations worldwide, with E. coli being consistently identified as one of the main causes. The economic impact goes far beyond temporary digestive upset. Affected calves frequently experience lasting growth retardation and failure to thrive. Han et al. (2017) quantified these losses, showing how a single outbreak can decimate an entire season’s profit margins. What makes these infections particularly frustrating is their persistence despite improved management practices. Colibacillosis frequently occurs in calves due to infection with one or more pathogenic strains of E. coli, such as Enterotoxogenic E. coli (ETEC), Shiga toxin-producing E. coli (STEC), Enteropathogenic E. coli (EPEC), and Enterohemolytic E. coli (EHEC) (Cabrera-González et al., 2024). Aref et al. (2018) found that pathogenic Escherichia coli strains were highly prevalent in both diarrheic and healthy calves, and that even clinically normal animals can act as reservoirs for zoonotic transmission. Their study also identified a novel hybrid strain of E. coli (ETEC-STEC) that causes diarrhea in calves and may pose a risk to public health. The most common cause of colibacillosis in neonatal calves is intestinal pathogenic E. coli ETEC (Nagy and Fekete, 2005). Taghadosi et al. (2018) characterized the serotypes of STEC isolated from humans, calves, and goats with diarrhea in Kerman, Iran, and identified the O26 and O91 serogroups. STEC secretes cytotoxic toxins (Stx1 and Stx2) that damage intestinal and renal endothelial cells ECs (Schouten, 2004). Ibrahim (2012) detected virulence factors in E. coli strains isolated from cattle farms with diarrheic calves (Maskana, Joreen, and Deir ez-Zor). The study found:6% of isolates carried the heat-stable enterotoxin A (sta) and Fimbrial adhesion (K99) genes, 14% harbored the Shiga toxin 1 (stx1) and Shiga toxin 2 (stx2) genes, and 1.33% carried the intimin (eaeA) gene. Additionally, Altaweel and Aldaoud (2023) isolated 1.04% of E. coli K99 from diarrheic calves in rural Aleppo and Idlib. The disturbing patterns of antibiotic resistance found in diarrheagenic E. coli from calves, as revealed by a series of empirical studies, add to the growing concern about the Antimicrobial Resistance (AMR) epidemic in veterinary medicine. Naderi et al. (2016) investigated 170 E. coli strains from diarrheic calves and revealed alarming antimicrobial resistance. All isolates were resistant to penicillin and sulfamethoxazole, with high resistance rates to tetracycline (73.5%), streptomycin (60%), trimethoprim-sulfamethoxazole (56.5%), and kanamycin (53.5%). In contrast, Srivani et al. (2017) established a connection between the prevalence of diarrheagenic E. coli among buffalo calves and the presence of multidrug-resistant strains, with a prevalence rate of 85.04%. Notably, 35% of the isolates were of the STEC type. Resistance of the highest magnitude was demonstrated against tetracycline (63.21%) and ampicillin (48.11%), while marked susceptibility to chloramphenicol and gentamicin (96.33%) was also exhibited. Notably, the Shiga toxin-producing isolates registered a multidrug resistance rate of 69.81%, indicating their high potential of posing a threat. This study aimed to detect some virulence factors in E. coli (sta, eae, stx1, stx2) and to determine the antibiotic resistance rates of these bacterial strains. Materials and MethodsStudy samplesSamples were collected from various regions in Hama Governorate, Syria, between June 2, 2024, and September 23, 2024. A total of 60 swab samples were obtained from diarrheic calves aged 1 day to 6 months. To preserve bacterial viability, each swab was placed in a peptone water transport tube. All samples were transferred to the laboratory for further microbiological testing on the same day. Bacterial isolationClinical samples were initially streaked onto MacConkey agar (HiMedia®) and incubated aerobically at 37°C for 24 hours to isolate Gram-negative enteric bacteria. Following incubation, two lactose-fermented colonies (pinkish) were selected and subcultured onto Eosin Methylene Blue (EMB) agar (HiMedia®) and incubated at 37°C for 24 hours. Characteristic E. coli colonies exhibiting a distinctive metallic green sheen were selected, purified, and preserved on nutrient agar for subsequent tests (Gibbons et al., 2014). All selected isolates were subjected to Gram staining to confirm their morphological characteristics. Biochemical testingPresumptive E. coli isolates were subjected to comprehensive biochemical profiling using the Hi25™ Enterobacteriaceae Identification Kit (KB003) according to the manufacturer’s instructions. Gram staining was also performed on E. coli isolates. Antibiotic susceptibility testingThe Kirby-Bauer disk diffusion method was used to assess antibiotic susceptibility. Bacterial suspensions were standardized to McFarland turbidity of 0.5 and swabbed onto Mueller-Hinton agar plates (HiMedia®). Commercially prepared antibiotic minidisks (Oxoid®) on inoculated plates. The tested antibiotics with their respective concentrations and codes are detailed in Table 1. Plates were incubated at 37°C for 24 hours under aerobic conditions. Zones of inhibition were measured in millimeters using calibrated digital calipers (CLSI, 2025) following incubation. Table 1. Antibiotics used in antimicrobial susceptibility testing of Escherichia coli isolates.
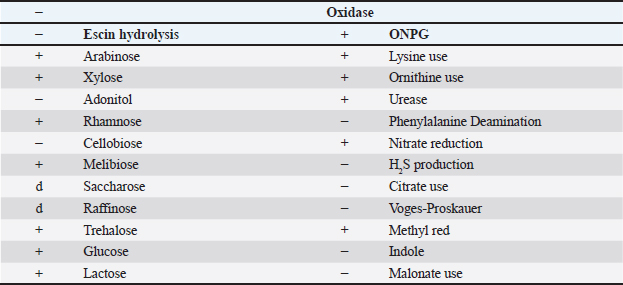
Detection of virulence genesDeoxyribonucleic acid (DNA) extractionDNA templates were prepared by transferring one pure bacterial colony into 50 μl of sterile distilled water. The tubes were then boiled in a water bath for 10 minutes, followed by centrifugation at 12,000 × g for 15 minutes. The supernatant was carefully collected and used as the DNA template for multiplex polymerase chain reaction (PCR) (Cengiz and Mehmet, 2020). Multiplex PCR assayThe virulence factor genes (sta, eae, stx1, and stx2) were detected using specific primers (Table 2). The PCR reaction mixture (25 μl final volume) consisted of the following: (3 μl DNA template, 1 μl each of forward and reverse primers (100 pmol/ul), 2.5 μl PCR buffer, 1.5 μl MgCl2 (25 mM), 0.5 μl dNTP mix, 0.5 μl Taq polymerase, and nuclease-free water up to 25 μl). The thermal cycling protocol was performed as follows (Cengiz and Mehmet, 2020): (Initial denaturation: 94°C for 3 minutes, 30 amplification cycles: (Denaturation: 94°C for 30 seconds, Annealing: 50°C for 45 seconds, and Extension: 72°C for 1.5 minutes) and Final extension: 72°C for 10 minutes. Table 2. Oligonucleotide primers used for the detection of E. coli virulence genes (Cengiz and Mehmet, 2020).
Detection of PCR productsPCR amplicons were analyzed by electrophoresis on a 2% agarose gel (GeneDire®, USA) prepared in 1× TBE buffer and stained with ethidium bromide (1 µg/ml). Electrophoresis was conducted at 100 V for 1 hour, and the band intensities were determined using a Gel documentation system (UviPro platinum, Uvitec, UK). The presence of specific DNA bands corresponding to the expected sizes of the tested virulence factors (Table 3) was confirmed by comparison with a molecular weight marker (DNA ladder). Table 3. Biochemical characteristics of Escherichia coli isolates from diarrheic calves.
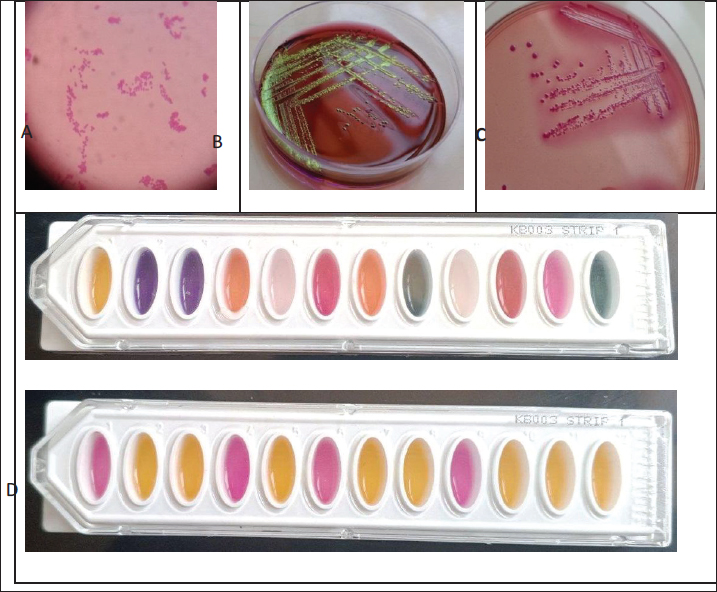
Statistical analysisData processing and graphical representation were performed using Microsoft Excel 2019 and GraphPad Prism (v8.2.1). Ethical approvalEthical approval was provided by the College of Veterinary Medicine, Hama University, Hama, Syria (No. 479, Date 20/2/2024). Rectal fecal samples were obtained from animals without anesthesia or tranquilizers after obtaining the permission of the farmers. ResultsEscherichia coli was found in 53 of 60 diarrheic calves on MacConkey and EMB media, giving an occurrence prevalence of 88.3%; in contrast, 7 remaining cases were negative for E. coli. The biochemical test results are summarized in Table 4 (Fig. 1).
Fig. 1. Isolation and biochemical characterization of Escherichia coli. A) Gram staining: Escherichia coli is a Gram-negative bacterium. B) Culture on EMB agar: E. coli produces greenish colonies with a distinctive metallic sheen on EMB agar. C) Culture on MacConkey agar: E. coli appears as pink colonies on MacConkey agar because of its ability to ferment lactose. D) Biochemical testing: Biochemical reactions were observed using the Hi25™ Enterobacteriaceae Identification Kit (KB003), which provides a profile for accurate identification. Table 4. Prevalence of virulence genes in diarrheic calves’ E. coli isolates.
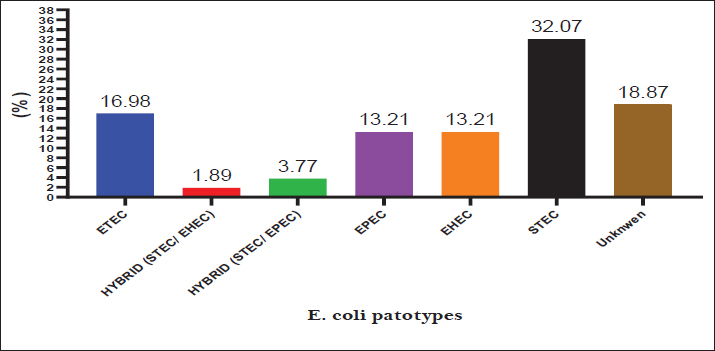
Virulence gene detection in pathogenic E. coli strainsMultiplex PCR revealed that 81.13% (43/53) of E. coli isolates carried various virulence genes. Screening for virulence-associated genes revealed the following pathotype distribution: ETEC (carrying sta gene) 16.98%, EPEC (carrying eae gene) 13.21%, EHEC (carrying stx + eae genes) 13.21%, STEC (carrying stx1 and/or stx2 only) 32.07%, hybrid strains (carrying mixed virulence genes from multiple pathotypes): 5.66% while Non-pathogenic strains (lacking all tested virulence genes) 18.87%. For detailed distribution, refer to Table 5, Figure 2, and Figure 4.
Fig. 2. Prevalence of Escherichia coli strains in diabetic calves.
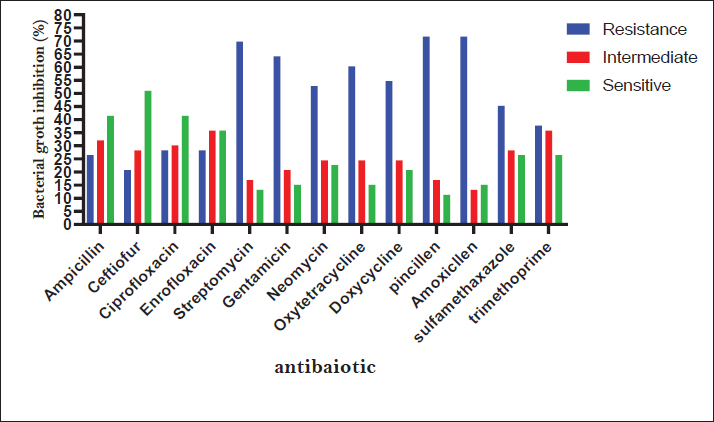
Fig. 3. Antibiotic susceptibility profile of E. coli isolates.
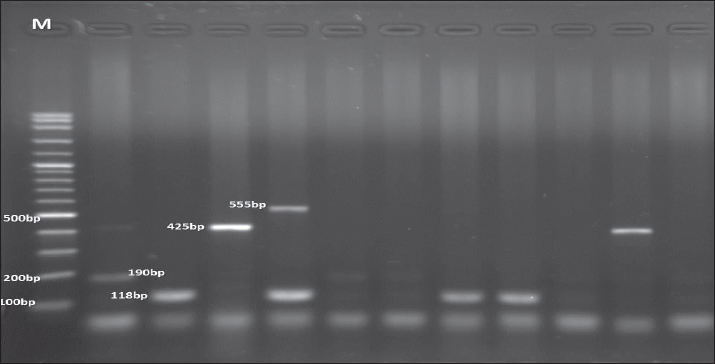
Fig. 4. Agarose gel electrophoresis product of multiplex PCR for the presence of stx1 (555 bp), stx2 (118 bp), eaeA (425 bp), and sta (190 bp) genes in different E. coli strains. Lane M: 100 bp ladder as a molecular size DNA marker. Table 5. Antibiotic susceptibility profile of E. coli isolates.
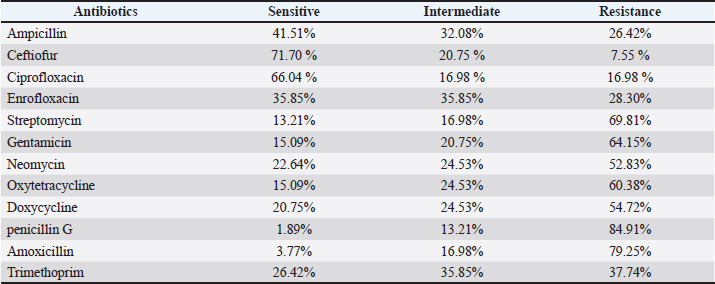
The study revealed the antibiotic resistance patterns among the tested isolates. Resistance rates were highest for penicillin G (84.91%) and amoxicillin (79.25%). Conversely, the most effective antibiotics showed susceptibility rates of cefquinome (71.70%) and ciprofloxacin (66.04%) (Table 5 and Fig. 3). DiscussionOur results demonstrate a remarkably high prevalence of virulence genes among E. coli isolates, occurring both individually and in various combinations of hybrids. These findings substantially exceed the rates reported by Ibrahim (2012) in Syrian cattle farms, where only 6% of isolates carried sta genes, 14% possessed Stx1/Stx2 genes, and 1.33% harbored the eae A gene. More recent data from Cabrera-González et al. (2024) revealed an even more complex virulence profile, 83.15% prevalence of sta genes, 1.05% for stx2, and 21.05% for eae. Notably, we observed significant frequencies of hybrid pathotype combinations EAEC/ETEC (44.21%), STEC/EAEC (3.15%), EPEC/ETEC (2.10%), STEC/EAEC/ETEC (21.05%), EPEC/EAEC/ETEC (6.31%), and EPEC/STEC/EAEC/ETEC (7.36%). Ok et al. (2009) investigated diarrheagenic E. coli in calves and reported sta gene 18.9%, Stx1 gene 13.5%, Stx2 gene 5.4%, and eae gene 8.1%. Nguyen et al. (2011) subsequently identified stx1 alone, 13.3% stx2 alone, 21.2%, and co-occurrence of stx1/stx2, 16.8%. They found substantially lower rates for the sta gene 0.6%, the eae gene 9.8%, and the eae/stx combination 6.7%. More recent data from Cengiz and Adigüzel (2020) revealed stx1 3.03%, stx2 9.09%, sta 21.21%, and eae 15.15%. Their pathotype distribution showed that ETEC was predominant (51.5%), EPEC (15.2%), EHEC (6.1%), and STEC/EHEC hybrid (12.1%). The widespread prevalence of pathogenic E. coli strains observed in this study may be attributed to the absence of E. coli vaccination in the geographical area where the research was conducted, in addition to the presence of some of these virulence genes in phages and plasmids, which facilitate horizontal gene transfer and the emergence of hybrid strains. The observed variations between our findings and those of previous studies may be attributed to differences in geographical regions, the epidemiology of pathogenic E. coli strains, and breeding types and feeding practices. The emergence of AMR stems from genetic modifications through multiple mechanisms, including spontaneous mutations in chromosomal DNA, horizontal gene transfer of resistance determinants via conjugation (plasmid-mediated), transduction (phage-mediated), and transposons (mobile genetic elements) (Normark and Normark, 2002). The findings revealed that E. coli bacteria causing calf diarrhea exhibit widespread resistance to antibiotics. In comparison with previous research, numerous studies have also indicated broad resistance in E. coli isolates from diarrheic calves. Cengiz and Adigüzel (2020) found that the highest resistance rates were to oxytetracycline (78.9%), trimethoprim-sulfamethoxazole (60.9%), and neomycin (58.6%). In contrast, the highest susceptibility rates were to ceftiofur (92.5%), cefoperazone (90.2%), and cefuroxime (90.2%). Paul et al. (2010) discovered that E. coli isolates from diarrheic calves were highly sensitive to enrofloxacin and ciprofloxacin, showed moderate sensitivity to cephalexin and amoxicillin, but were resistant to nalidixic acid and erythromycin. Bok et al. (2015) isolated E. coli from diarrheic calves. These isolates showed the highest resistance rate to neomycin (51%) and the lowest resistance rate to norfloxacin (0.7%). The isolates from cattle showed the highest resistance rate to neomycin (24.6%) and the lowest resistance rate to cefapirin (0.8%). Li et al. (2018) observed that E. coli strains exhibited varying degrees of resistance to 19 commonly used antibiotics, particularly tetracycline, cephradine, doxycycline, carbenicillin, ampicillin, ciprofloxacin, and norfloxacin. Resistance rates for these drugs exceeded 40%, with tetracycline resistance being the most common at 53.7%. No strains showed resistance to neomycin or polymyxin B. Antibiotic resistance in E. coli could also arise from the irresponsible use of antibiotics, the use of multiple antibiotics without testing their efficacy, and the non-completion of treatment. These actions tend to cause bacteria to develop resistance to the applied antibiotics. The discrepancy between the results of this study and previous studies may be attributed to differences in treatment methods and the commonly used medications for managing animal diseases. ConclusionPathogenic E. coli strains are among the most significant causes of diarrhea in calves. These pathogens also represent a zoonotic concern, as they can transmit and infect humans. Extensive resistance to multiple antibiotics poses a major public health threat because multidrug-resistant strains in animals pose a potential hazard to human health. Consequently, E. coli infection treatment must be guided by antibiotic sensitivity testing results. In addition, we recommend limiting the indiscriminate use of antibiotics and conducting more research on the dissemination of pathogenic Escherichia coli strains among animal and human populations. AcknowledgmentsNone. Conflict of interestThe authors declare no conflict of interest. FundingThe main funder of this research is the University of Hama, which provided the laboratories in which the research was conducted. Authors’ contributionsASEEM ALBAKER is the primary researcher, implementer, and writer of the research, as this research is part of the doctoral thesis that is currently being prepared. Dr. MAHER SALEH supervises the research as a scientific supervisor and assistant in directing and writing the thesis, and Dr. ASHRAF ALSALEH is the assistant practical supervisor during the implementation of the research. Data availabilityAll data are provided in the revised manuscript. ReferencesAltaweel, H. and Aldaoud, F. 2023. Investigation of E. coli K99 and rotaviruses causing diarrhea in newborn calves in the northwest of Syria. Idlib. Univ. Res. J. 6, 245–271. Aref, N.M., Abdel-Raheem, A.A., Kamaly, H.F. and Hussien, S.Z. 2018. Clinical and sero-molecularcharacterization of Escherichia coli with an emphasis on hybrid strain in healthy and diarrheic neonatal calves in Egypt. Open Vet. J. 8, 351–359. Bok, E., Mazurek, J., Stosik, M., Wojciech, M. and Baldy-Chudzik, K. 2015. Prevalence of virulence determinants and antimicrobial resistance among commensal Escherichia coli derived from dairy and beef cattle. Int. J. Environ. Res. Public Health 12, 970–985. Cabrera-González, M., Quilcate-Paírazamán, C., Alvarez-García, W., Cabrera, H., Tayca-Saldaña, A., Aliaga-Tambo, F., Rojas-Valdez, D. and Cueva-Rodríguez, M. 2024. Molecular identification of the most frequent pathotypes of Escherichia coli in calves with diarrhoea in the Cajamarca region of Peru. Open Veterinary Journal, 14(9), 2170 -2180. Cengiz, S. and Adigüzel, M.C. 2020. Determination of virulence factors and antimicrobial resistance of E. coli isolated from calf diarrhea, part of eastern Turkey. Ankara. Univ. Vet. Fak. Derg. 67, 365–371. CLSI. 2025. Performance standards for antimicrobial susceptibility testing (35th ed). CLSI supplement M100. Wayne, PA: Clinical and Laboratory Standard Institute. Foster, D.M. and Smith, G.W. 2009. Pathophysiology of diarrhea in calves. Vet. Clin. North Am. Food Anim. Pract. 25, 13–36. Gibbons, J.F., Boland, F., Buckley, J.F., Butler, F., Egan, J., Fanning, S., Markey, B.K. and Leonard, F.C. 2014. Patterns of antimicrobial resistance in pathogenic Escherichia coli isolates from cases of calf enteritis during the spring-calving season. Vet. Microbiol. 170, 73–80. Han, Z., Li, K., Shahzad, M., Zhang, H., Luo, H., Qiu, G., Lan, Y., Wang, X., Mehmood, K. and Li, J. 2017. Analysis of the intestinal microbial community in healthy and diarrheal perinatal yaks by high-throughput sequencing. Microb. Pathog. 111, 60–70. Ibrahim, S.K. 2012. Detection of virulence factors of Escherichia coli strains isolated from cases of calf diarrhea by polymerase chain reaction. Tishreen Univ. J. For Res. Scientific Stud. - Biol. Sci. Ser. 34, 51–62. Lez, M., N, C., A, W., Cabrera, H., A, A., Tambo, F., Valdez, D. and Guez, M. 2024. Molecular identification of the most frequent pathotypes of Escherichia coli in calves with diarrhoea in the Cajamarca region of Peru. Open Vet. J. 14, 2170–2180. Li, K., Wang, X.Q., Shahzad, M., Zhang, H., Zhao, X.D., Jiang, X., Mehmood, K., Han, Z.Q., Wang, L. and Li, J.K. 2018. Antibiotic resistance and screening of the resistant genes of Escherichia coli (E.coli) isolated from diarrheal yak calves in Sichuan Province, China. Trop. Biomed. 35, 478–486. Nagy, B. and Fekete, P.Z. 2005. Enterotoxigenic Escherichia coli in veterinary medicine. Int. J. Med. Microbio. 295, 443–454. Naderi, Z., Ghanbarpour, R. and Sami, M. 2016. Antimicrobial resistance characteristics and phylogenetic groups of Escherichia coli isolated from diarrheic calves in southeast of Iran. Int. J. Enteric Pathog. 4(4), 1-7. Nguyen, T.D., Vo, T.T. and Vu-Khac, H. 2011. Virulence factors in Escherichia coli isolated from calves with diarrhea in Vietnam. J. Vet. Sci. 12, 159–164. Normark, B.H. and Normark, S. 2002. Evolution and spread of antibiotic resistance. J. Intern. Med. 252, 91–106. Ok, M., Güler, L., Turgut, K., Ok, U., Şen, I., Gündüz, I.K., Birdane, M.F. and Güzelbekteş, H. 2009. The studies on the aetiology of diarrhoea in neonatal calves and determination of virulence gene markers of Escherichia coli strains by multiplex PCR. Zoonoses. Public. Health. 56, 94–101. Paul, S.K., Khan, M.S.R., Rashid, M.A., Hassan, J. and Mahmud, S.M.S. 2010. Isolation and characterization of Escherichia coli from buffalo calves in some selected areas of Bangladesh. Bangladesh J. Vet. Med. 8, 23–26. Srivani, M., Reddy, Y. N., Subramanyam, K. V., Reddy, M. R. and Rao, T. S. 2017. Prevalence and antimicrobial resistance pattern of Shiga toxigenic Escherichia coli in diarrheic buffalo calves. Veterinary world, 10(7), 774. Schouten, J. 2004. Prevalence Estimation and Risk Factors for Escherichia coli O157 on Dutch dairy farms. Prev. Vet. Med. 64, 49–61. Taghadosi, R., Shakibaie, M. R., Alizade, H., Hosseini-Nave, H., Askari, A., & Ghanbarpour, R. (2018). Serogroups, subtypes and virulence factors of shiga toxin-producing Escherichia coli isolated from human, calves and goats in Kerman, Iran. Gastroenterol. Hepatol. Bed Bench.11, 60. | ||
| How to Cite this Article |
| Pubmed Style Albaker AK, Saleh M, Alsaleh A. Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Vet. J.. 2025; 15(12): 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 Web Style Albaker AK, Saleh M, Alsaleh A. Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. https://www.openveterinaryjournal.com/?mno=279032 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.8 AMA (American Medical Association) Style Albaker AK, Saleh M, Alsaleh A. Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Vet. J.. 2025; 15(12): 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 Vancouver/ICMJE Style Albaker AK, Saleh M, Alsaleh A. Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 Harvard Style Albaker, A. K., Saleh, . M. & Alsaleh, . A. (2025) Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Vet. J., 15 (12), 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 Turabian Style Albaker, Aseem K., Maher Saleh, and Ashraf Alsaleh. 2025. Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Veterinary Journal, 15 (12), 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 Chicago Style Albaker, Aseem K., Maher Saleh, and Ashraf Alsaleh. "Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria." Open Veterinary Journal 15 (2025), 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 MLA (The Modern Language Association) Style Albaker, Aseem K., Maher Saleh, and Ashraf Alsaleh. "Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria." Open Veterinary Journal 15.12 (2025), 6237-6244. Print. doi:10.5455/OVJ.2025.v15.i12.8 APA (American Psychological Association) Style Albaker, A. K., Saleh, . M. & Alsaleh, . A. (2025) Detection of virulence factors of Escherichia coli strains isolated from calves diarrhea and their resistance to antibiotics in Hama Governorate, Syria. Open Veterinary Journal, 15 (12), 6237-6244. doi:10.5455/OVJ.2025.v15.i12.8 |