| Research Article | ||
Open Vet. J.. 2025; 15(12): 6298-6305 Open Veterinary Journal, (2025), Vol. 15(12): 6298-6305 Research Article Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red SeaMohammad Allam1, Ali H. Abu Almaaty2, Layla Omran Elmajdoub3* and Maryham M. Aziz21Zoology Department, Faculty of Science, Luxor University, Luxor, Egypt 2Zoology Department, Faculty of Science, Port Said University, Port Said, Egypt 3Zoology Department, Faculty of Science, Misurata University, Misurata, Libya *Corresponding Author: Layla Omran Elmajdoub. Zoology Department, Faculty of Science, Misurata University, Misurata, Libya. Email: elmajdoublayla [at] sci.misuratau.edu.ly Submitted: 16/08/2025 Revised: 01/11/2025 Accepted: 15/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
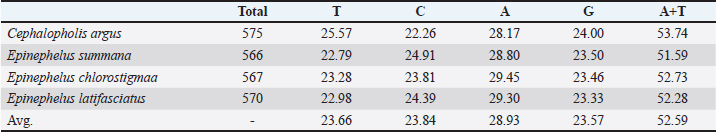
AbstractBackground: Grouper fishes have a diverse variety of shapes and colors, making it difficult to accurately perform morphological identification. In addition, certain unique grouper species may be included in the same species description due to morphological similarities. Aim: The main purpose of this study was to investigate the evolutionary relationships between various species of the family Epinephelinae using the large mitochondrial rRNA gene. Methods: Four Epinephelinae species (Cephalopholis argus, Epinephelus summana, Epinephelus chlorostigma, and Epinephelus latifasciatus) were gathered from the Red Sea and identified (Randall, 1982). Results: The accession numbers were obtained by displaying the 16S rRNA sequences in GenBank/NCBI (PQ661169.1—PQ661172.1). The average frequencies of adenine (A), thymine (T), cytosine (C), and guanine (G) were 28,93%, 23.66%, 23.84%, and 23.57%, respectively. Compared with C + G, the average proportion for A + T was more significant. Conclusion: Epinephelus summana and E. latifasciatus are closely linked, as evidenced by their low genetic distance. Keywords: 16S rRNA gene, Cephalopholis, Epinephelus, Grouper. IntroductionThe 16S rRNA gene is the ultimate molecular chronometer because of several of its characteristics. It is considered the most prevalent housekeeping genetic marker, making it a valuable target for phylogeny and clinical identification (Woese, 1978; Patel, 2001; Janda and Abbott, 2007). The function of the 16S rRNA has not changed in a long time; therefore, random sequence variations are more likely to represent microbial evolutionary (phylogeny) than selected sequences that change the function of the molecule (Woese, 1978). Moreover, the size of the 16S rRNA gene is approximately 1,500 bp, which is sufficient for informatics applications (Weisburg, 1991; Patel, 2001; Janda and Abbott, 2007). The presence of several conserved/hypervariable regions that provide a variety of PCR primer design options is a crucial feature that supports the use of the 16S rRNA gene (Van de Peer et al., 1996). The 16S rRNA gene has approximately 50 functional domains, and sequences in other domains are not significantly impacted by the introduction of specific modifications in one domain (Patel, 2001). To identify genes, regulatory elements, and non-coding RNAs in recently sequenced genomes, classify closely related species sequences, and refine phylogenetic trees, molecular phylogenetics is used to explain both ancient and modern individual genomes (Kellis et al., 2003; Pedersen et al., 2006). Phylogenetic analysis employs molecular techniques based on gene sequence analyses to categorize and comprehend the relationships between closely related species in systematics and taxonomy. This approach has grown in significance across all taxonomy domains due to the abundance of publicly accessible genetic sequence data (Yang and Rannala, 2010). Consequently, coral reef ecosystems have a high degree of biodiversity, particularly in fish species (Allen and Erdman 2012). Fish habitat is one of the most significant ecological roles of coral reef ecosystems, particularly as a place for spawning, feeding, nidification, and predator protection (Yuliana et al., 2020; Mujiyanto et al., 2021). Groupers are reef fish that belong to the Epinephelinae subfamily and the Serranidae family. In tropical and subtropical regions, groupers naturally live in shallow-water environments such as coral reefs, estuaries, mangroves, and seagrass (Kamal et al., 2019; Andriyono et al., 2020). There are 159 species in the Epinephelinae subfamily, including 31 genera of Epinephelus and 15 genera of Cephalopholis (Allen and Adrim, 2003). The Serranidae family has historically been regarded as a convenient pigeonhole for lower percoid fish with ambiguous affinities, despite more recent systematic treatments of the family offering a valid categorization. Our knowledge of the relationships of Serranidae has also been unclear (Craig and Hastings, 2007). Considering the complexity of the members of the subfamily Epinephelinae (Epinephelus, Serranidae), the limitations of studying lineages among fish in this family remain crucial (Craig and Hastings, 2007). The morphological characteristics of the Epinephelus and Cephalopholis genera are very similar. Consequently, misidentifications between the two genera may occur during morphological identification. Given the limitations of morphological traits alone, a molecular approach is required to support the use of morphological features in species characterization. Age and other environmental factors affect traits (Becker et al., 2015; Hulley et al., 2018; Dwifajri et al., 2022). The main purpose of this study was to use the 16S rRNA gene to examine the evolutionary relationships between various species of the Epinephelinae family. Materials and MethodsGathering samples and species identificationThe Red Sea served as the study sampling location, and four Epinephelinae species (Cephalopholis argus, Epinephelus summana, Epinephelus chlorostigma, and Epinephelus latifasciatus) were gathered and identified (Randall, 1982). To extract DNA, muscle tissue was dissected from the specimens and stored at −20°C. DNA isolation and polymerase chain reaction amplificationGenomic DNA was extracted from the stored muscles using the Biospin Genomic DNA Extraction Kit following the manufacturer’s instructions. The DNA quality was assessed by Agarose gel electrophoresis. A partial sequence of the mitochondrial 16S rRNA was amplified by PCR using recently reported primers (Simon et al., 1991). The PCR reactions were carried out in a total volume of 50 μl consisting of 25μl of 2X master mix, 1μl of genomic DNA, 1μl of each primer, and 22μl of nuclease-free water. Five minutes of denaturation at 95°C, 30 cycles of denaturation, annealing, and extension at 94°C, 48°C, and 72°C, respectively, for one minute each, followed by a final extension at 72°C for 10 minutes, were the conditions for PCR amplification. Electrophoresis was performed on a 1.5% agarose gel using a 100 bp DNA ladder and ethidium bromide to visualize the outcomes of PCR. Sequences and phylogenetic analysesMacrogen (Seoul, South Korea) completed the sequences. The sequences of the 16S rRNA were uploaded to GenBank/NCBI to obtain accession numbers. CLUSTAL W (Thompson et al., 1994) was used to align the sequences with the default settings. Neighbor joining and minimum evolution were the two methods used for phylogenetic structure tree using MEGA software version 7.0 (Kumar et al., 2016). To fully implement the sequence divergences (Felsenstein, 1985), Kimura two-parameter distances with 1,000 bootstrapped iterations (Kimura, 1980) were used. Ethical approvalThe Ethics of Animal Experiments Committee of the Faculty of Science of Port Said University accepted all animal experimentation practices (ERN: PSU. Sci. 79). ResultsThis study establishes the evolutionary lineages of Epinephelinae species (C. argus, E. summana, E. chlorostigma, and E. latifasciatus). This was achieved using large subunit ribosomal RNA (16S rRNA) sequences. In all four species, the 16S rRNA-produced bands ranged in length from 566 to 575 bp. The accession numbers were obtained by displaying the 16S rRNA sequences in GenBank/NCBI (PQ661169.1—PQ661172.1). The results indicate that Cephalopholis argus has the longest sequence (575 bp), whereas E. summana has the shortest sequence (566 bp). The average frequencies of adenine (A), thymine (T), cytosine (C), and guanine (G) were 28,93%, 23.66%, 23.84%, and 23.57%, respectively. Compared with C + G, the average proportion for A + T was more significant (Table 1). Table 1. Four species of the family Epinephelinae, along with their accession numbers, nucleotide frequencies, A+T concentrations, and averages of the 16S rRNA sequence.
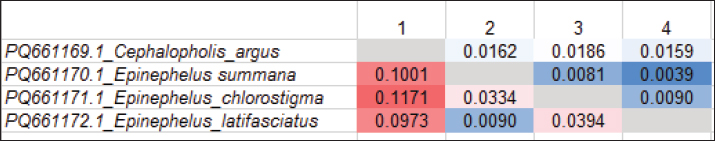
A total of 578 base pairs were included in the final alignments. The conserved and variable locations were 505 and 63, respectively. Throughout all fish, the P-distances varied from 0.0039% to 1.4114%. The P-distances of the understudied species varied from 0.0039% to 0.0186%. The greatest amount (0.0186) was found between C. argus and E. chlorostigma. The smallest value (0.0039) was found between E. summana and E. latifasciatus (Table 2 and Fig. 1).
Fig. 1. Heatmap representation of the 16S rRNA gene used to establish the P-distances among the four Epinephelinae species. Table 2. The pairwise distances between the outgroup and four members of the Epinephelinae family using the 16S rRNA gene.
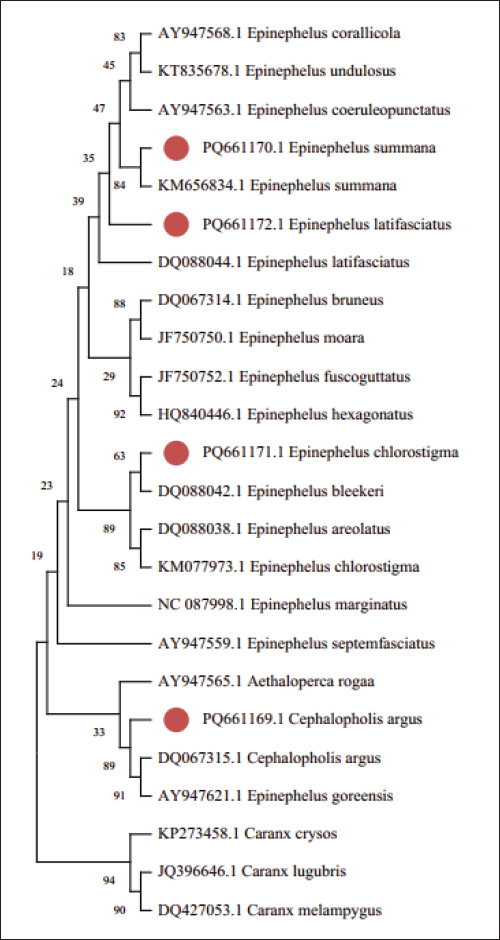
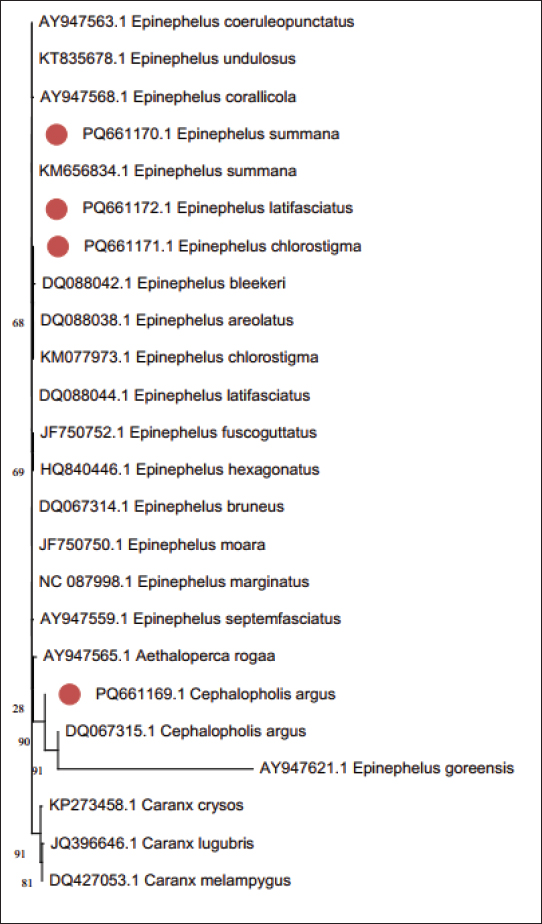
The sequences gathered from four species in the Epinephelinae family, 17 link sequences, and three species of the out-group from GenBank were employed in this study for a wide phylogenetic research to finalize the phylogenetic tree analysis utilizing the 16S rRNA sequence. Multiple phylogenetic techniques were used for the most compelling phylogenetic analysis employing the 16S rRNA gene: Neighbor joining and minimum evolution. Despite a small variation in the support rate, the approaches produced findings that were fundamentally similar and emphasized two key points: (1) A unique cluster is formed by the outgroup species and (2) With the exception of Epinephelus goreensis, all Epinephelus species were clustered close to one another (Figs. 2 and 3).
Fig. 2. The phylogenetic tree of Neighbor joining between species of the Epinephelinae family and the outgroup using the 16S rRNA gene.
Fig. 3. Minimum evolution phylogenetic tree between species of the Epinephelinae family and the outgroup using the DiscussionHeino (2014) stated that the morphological qualities of a living organism can vary according to the ecological conditions in which it lives. Fish exhibit a great deal of variation in their physical attributes, both between and within groups (Brraich and Akhter, 2015). The morphological changes in fish symbolize a kind of environmental adaptation (Hossain et al., 2010). Because conventional taxonomy frequently uses random morphometric datasets and lacks standards for character selection or coding, species identification can be challenging. In some situations, genetic analysis may be used as an additional method of determining taxonomic identification (Basheer et al., 2015). Owing to the diversity of shapes and colors in grouper fishes (Tapilatu et al., 2021), this study investigated the evolutionary relationships between various species of the Epinephelinae family using 16S rRNA sequences. Mitochondrial 16S rRNA has a slower rate of mutation and a lower rate of substitution than other mtDNA genes; thus, it is valuable for studying species, communities, and families (Garland and Zimmer, 2002). Furthermore, the 16S rRNA gene may be used to assess fish phylogenetic connections at both the species and generic levels (Moyer et al., 2004; Chakraborty and Iwatsuki, 2006). Thus, 16S rRNA is advised for the reconstruction of pertinent phylogenetic relationships and as an appropriate identification technique in fish evolutionary studies (Saad et al., 2019). Using 16S rRNA, nucleotide sequences from the understudied fish had an average length of approximately 550 bp. According to Simon et al. (1991), a 500–650 base fragment is amplified by the 16Sar and 16Sbr primers; therefore, this length was within expectations. Furthermore, several studies found the same outcome (Mar'ie and Allam, 2019; Alyamani et al., 2023; Allam et al., 2024; Eisa et al., 2025). Contrary to (C + G), this investigation revealed that the average bidder for the understudied fish was (A + T). This finding was consistent with further research. Bo et al. (2013) found that the whole 16S rRNA gene exhibits A+T affluence as opposed to C+G affluence. Basheer et al. (2015) discovered that 16S rRNA had a smaller C+G score than A+T in their study of Rastrelliger species. Furthermore, Mar’ie and Allam (2019) found that two puffer fish had a higher A+T ratio than C+G. Mahrous and Allam (2022) found that the A+T rate in various catfish species was higher than the C+G rate. Eisa et al. (2025) reported that the average content of the A+T sequence of the 16S rRNA gene in two Serranid species (Epinephelus aneus and Mycteroperca rubra) was higher than that of the C+G sequence. Complete alignments of incomplete 16S rRNA sequences in each of the four species of the Epinephelinae family showed high levels of conserved sites. Basheer et al. (2015) found 575 consistent locations of 590 bp in three Rastrelliger species utilizing 16S rRNA aligned sequences. Sokefun (2017) found 337 conserved regions totaling 463 bp of alignment in a phylogenetic study of Cichlids using the 16S gene. Aligning the incomplete 16S rRNA gene sequences of some Carangid fishes reveals several highly conserved areas (Alyamani et al., 2023). According to Ramadan et al. (2023), the (A + T) median value in the four Lutjanus fish species was greater than the (C + G) median value. According to Kaleshkumar et al. (2015), highly related species exhibited low genetic distance values, whereas the greatest genetic distance is responsible for situations with considerable genetic divergence. Epinephelus summana and E. latifasciatus were closely linked, as evidenced by their low genetic distance. AcknowledgmentsNone. Conflict of interestThere are no conflicts of interest. FundingNot applicable. Authors' contributionsAll authors contributed equally to this study. Data availabilityAll data are provided in the manuscript. ReferencesAllam, M., Eisa, S.G.A., Abu Almaaty, A.H. and Hassan M.K. 2024. Phylogenetic relationships and genetic diversity of some sparid species from the Egyptian Mediterranean Sea using 16S rRNA Gene. Egypt. J. Aquat. Biol. Fish. 28(3), 739–752. Allen, G.R and Adrim, M. 2003. Coral reef fishes of Indonesia. Zool. Stud. 42(1), 1–72. Allen, G.R. and Erdmann, M.V., 2012. Reef Fishes of the East Indies Volume IIII. Tropical Reef Research Institute, Perth, Australia. Alyamani, N.M., Fayad, E., Abu Almaaty, A.H., Ramadan, A. and Allam, M. 2023. Phylogenetic relationships among some carangid species based on analysis of mitochondrial 16S rRNA Sequences. Egypt. J. Aquat. Biol. Fish. 27(2), 61–74. Andriyono, S., Damora, A. and A. Hidayani, A. 2020. Genetic diversity and phylogenetic reconstruction of grouper (Serranidae) from Sunda Land, Indonesia. Egypt. J. Aquat. Biol. Fish. 24(3), 403–415; doi:10.21608/ejabf.2020.92320 Brraich, O.S. and Akhter, S. 2015. Morphometric characters and meristic counts of a fish, Crossocheilus latius latius (Hamilton-Buchanan) from Ranjit Sagar Wetland, India. Int. J. Fish. Aquat. Stud. 2(5), 260–265. Basheer, V.S., Mohitha, C., Vineesh, N., Divya, P.R., Gopalakrishnan, A. and Jena, J.K. 2015. Molecular phylogenetics of three species of the genus Rastrelliger using mitochondrial DNA markers. Mol. Biol. Rep. 42(4), 873–879; doi:10.1007/s11033-014-3710-8 Becker, R.A., Sales, N.G., Santos, G.M., Santos, G.B. and Carvalho, D.C. 2015. DNA barcoding and morphological identification of neotropical ichthyoplankton from Upper Paraná and São Francisco. J. Fish. Biol. 87(1), 159–168; doi:10.1111/jfb.12707 Bo, Z., Xu, T., Wang, R., Jin, X. and Sun, Y. 2013. Complete mitochondrial genome of the Bombay duck Harpodon nehereus (Aulopiformes, Synodontidae). Mitochondrial DNA. 24(6), 660–662. doi:10.3109/19401736.2013.772988 Chakraborty, A. and Iwatsuki, Y. 2006. Genetic Variation at the Mitochondrial 16S rRNA Gene among Trichiurus lepturus (Teleostei: trichiuridae) from Various Localities: Preliminary Evidence of a New Species from West Coast of Africa. Hydrobiologia 563(1), 501–513; doi:10.1007/s10750-006-0105-4 Craig, M.T. and Hastings, P.A. 2007. A molecular phylogeny of the groupers of the subfamily Epinephelinae (Serranidae) with a revised classification of the Epinephelini. Ichthyol. Res. 54, s1–17; doi:10.1007/s10228-006-0367-x Dose, C.R. 1987. Bacterial evolution. Microbiol. Rev. 51(2), 221–271; doi:10.1128/mr.51.2.221-271.1987 Dwifajri, S., Tapilatu, R.F., Pranata, B. and Kusuma, A.B. 2022. Molecular phylogeny of grouper of Epinephelus genus in Jayapura, Papua, Indonesia inferred from Cytochrome Oxidase I (COI) gene. Biodiversitas 23, 1449–1456. Eisa, S.G.A., Hassan, M.K., Abu Almaaty, A.H. and Allam, M. 2025. Genetic Variations of Male and Female E. aeneus Fishes from the Egyptian Mediterranean Sea Using Mitochondrial 16S rRNA Sequences. Egypt. J. Aquat. Biol. Fish. 29(2), 1673–1687. Felsenstein, J. 1985. Confidence Limits on Phylogenies: an Approach Using The Bootstrap. Evolution 39(4), 783–791; doi:10.1111/j.1558-5646.1985.tb00420.x Garland, E.D. and Zimmer, C.A. 2002. Techniques for identifying bivalve larvae. Mar. Ecol. Prog. Ser. 225, 299–310; doi:10.3354/meps225299 Heino, M. 2014. Quantitative Traits. In: Cadrin SX, Karr LA, Mariani S (eds) Stock Identification Methods: Applications in Fishery Science. 2nd Edition. Academic Press – Elsevier, London, UK. Hossain, M.A.R., Nahiduzzaman, M.D., Saha, D., Khanam, M.H. and Alam, M.S. 2010. Landmark-based morphometric and meristic variations of the endangered carp, Kalibaus Labeo calbasu, from stocks of two isolated rivers, the Jamuna and Halda and a Hatchery. Mar. Ecol. Biol. Zool. Stud. 49(4), 556–563. Hulley, E.N., Taylor, N.D.J., Zarnke, A.M., Somers, C.M., Manzon, R.G., Wilson, J.Y. and Borehamd, D.R. 2018. DNA barcoding vs. morphological identification of larval fish and embryos in Lake Huron: Advantages to a molecular approach. J. Great Lakes Res. 44(5), 1110–1116. doi: 10.1016/j.jglr.2018.07.013 Janda, J.M. and Abbott, S.L. 2007. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J. Clin. Microbiol. 45(9), 2761–2764; doi:10.1128/JCM.01228-07 Kaleshkumar, K., Rajaram, R., Vinothkumar, S., Ramalingam, V. and Meetei, K.B. 2015. Note: DNA barcoding of selected species of pufferfishes (Order : Tetraodontiformes) of Puducherry coastal waters along south-east coast of India. Indian J. Fish. 62(2), 98–103. Kamal, M.M., Hakim, A.A., Butet, N.A., Fitrianingsih, Y. and Astuti, R. 2019. Autentikasi spesies ikan kerapu berdasarkan marka gen MT-COI dari perairan Peukan Bada, Aceh. J. Biol. Tropis. 19(2), 116–123; doi:10.29303/jbt.v19i2.1245 Kellis, M., Patterson, N., Endrizzi, M., Birren, B. and Lander, E.S. 2003. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423(6937), 241–254; doi:10.1038/nature01644 Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences.J. Mol. Evo. 16(2), 111–120; doi:10.1007/BF01731581 Kumar, S., Stecher, G. and Tamura, K. 2016. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 33(7), 1870–1874; doi:10.1093/molbev/msw054 Marie, Z. and Allam, M. 2019. Molecular phylogenetic linkage for Nile and marine puffer fishes using mitochondrial DNA sequences of cytochrome b and 16S rRNA. Egypt. J. Aquat. Biol. Fish. 23(5), 67–80. Moyer, G.R., Burr, B.M. and Krajewski, C. 2004. Phylogenetic relationships of thorny catfishes (Siluriformes: Doradidae) inferred from molecular and morphological data. Zool. J. Linn. Soc. 140(4), 551–575; doi:10.1111/j.1096-3642.2004.00114.x Mujiyanto, M., Sugianti, Y., Afandy, Y.A., Rahayu, R., Budikusuma, R.A., Nastiti, A.S., Syam, A.R. and Purnamaningtyas, S.E. 2021. Reef fish community structure in the islands of Paraja Bay, Pandeglang District, Banten, Indonesia. Biodiversitas 22(10), 4402–4413; doi:10.13057/biodiv/d221033 Parous, N.S. and Allam, M. 2022. Phylogenetic relationships among some catfishes assessed by small and large Mitochondrial rRNA sequences. Egypt. J. Aquat. Biol. Fish. 26(6), 1069–1082. Patel, J.B. 2001. 16S rRNA gene sequencing for bacterial pathogen identification in the clinical laboratory. Mol. Diagn. 6(4), 313–321; doi:10.1054/modi.2001.29158 Pedersen, J.S., Bejerano, G., Siepel, A., Rosenbloom, K., Lindblad-Toh, K., Lander, E.S., Kent, J., Miller, W. and Haussler, D. 2006. Identification and classification of conserved RNA secondary structures in the human genome. PLos Comput. Biol. 2(4), e33; doi:10.1371/journal.pcbi.0020033 Randall, J.E. 1982. The diver guide to Red Sea reef fishes. Publishing limited 20 Berkeley street, Berkeley square London Wix 5AE Saad, Y., Abdulkader, S.-O. and Gharbawi, W. (2019). Evaluation of molecular diversity in some Red Sea parrotfish species based on mitochondrial 16S ribosomal RNA gene sequence variations. Res. J. Biotec. 2014;14:30–38. Simon, C., Franke, A. and Martin, A. 1991. The polymerase chain reaction: DNA extraction and amplification. In Molecular Techniques in taxonomy. Eds. G.M. Hewitt, A.W.B. Johnston and J.P.W. Young. NATO AS1 Series H 57: 329 355, 57: 329 355. Sokefun, O.B. 2017. The cichlid 16S gene as a phylogenetic marker: Limits of its resolution for analyzing global relationship. Int. J. Genet. Mol. Biol. 9(1), 1–7. doi:10.5897/ijgmb2016.0131 Tapilatu, R.F., Tururaja, T.S., Sipriyadi. and Kusuma, A.B. 2001. Molecular Phylogeny Reconstruction of Grouper (Serranidae: Epinephelinae) at Northern Part of Bird’s Head Seascape - Papua Inferred from COI Gene. Fish Aquatic Sci. 24(5), 181–190. doi: 10.47853/FAS.2021.e18. Thompson, J.D., Higgins, D.G. and Gibson, T.J. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22(22), 4673–4680. doi: 10.1093/nar/22.22.4673 Van de Peer, Y., Chapelle, S. and De Wachter, R. 1996. A quantitative map of nucleotide substitution rates in bacterial rRNA. Nucleic Acids Res. 24(17), 3381–3391. doi:10.1093/nar/24.17.3381 Weisburg, Barns, S.M., Pelletier, D.A. and Lane, D.J. 1991. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173(2), 697–703; doi:10.1128/jb.173.2.697-703 Yang, Z. and Rannala, B. 2010. Bayesian species delimitation using multilocus sequence data. Proc. Natl. Acad. Sci. U. S. A. 107(20), 9264–9269; doi:10.1073/pnas.0913022107 Yuliana, E., Farida, I., Nurhasanah, Boer, M., Fahrudin A. and Kamal, M.M. 2020. Habitat quality and reef fish resources potential in Karimunjawa National Park, Indonesia. AACL. Bioflux. 13(4), 1836–1848. | ||
| How to Cite this Article |
| Pubmed Style Allam M, Abu-almaaty AH, Elmajdoub LO, Aziz MM. Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Vet. J.. 2025; 15(12): 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 Web Style Allam M, Abu-almaaty AH, Elmajdoub LO, Aziz MM. Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. https://www.openveterinaryjournal.com/?mno=277698 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.13 AMA (American Medical Association) Style Allam M, Abu-almaaty AH, Elmajdoub LO, Aziz MM. Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Vet. J.. 2025; 15(12): 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 Vancouver/ICMJE Style Allam M, Abu-almaaty AH, Elmajdoub LO, Aziz MM. Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 Harvard Style Allam, M., Abu-almaaty, . A. H., Elmajdoub, . L. O. & Aziz, . M. M. (2025) Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Vet. J., 15 (12), 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 Turabian Style Allam, Mohammad, Ali H. Abu-almaaty, Layla Omran Elmajdoub, and Maryham M. Aziz. 2025. Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Veterinary Journal, 15 (12), 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 Chicago Style Allam, Mohammad, Ali H. Abu-almaaty, Layla Omran Elmajdoub, and Maryham M. Aziz. "Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea." Open Veterinary Journal 15 (2025), 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 MLA (The Modern Language Association) Style Allam, Mohammad, Ali H. Abu-almaaty, Layla Omran Elmajdoub, and Maryham M. Aziz. "Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea." Open Veterinary Journal 15.12 (2025), 6298-6305. Print. doi:10.5455/OVJ.2025.v15.i12.13 APA (American Psychological Association) Style Allam, M., Abu-almaaty, . A. H., Elmajdoub, . L. O. & Aziz, . M. M. (2025) Utility of large mitochondrial rRNA sequences in phylogenetic inference of some epinephelinae species from the Egyptian Red Sea. Open Veterinary Journal, 15 (12), 6298-6305. doi:10.5455/OVJ.2025.v15.i12.13 |