| Research Article | ||
Open Vet. J.. 2025; 15(12): 6506-6513 Open Veterinary Journal, (2025), Vol. 15(12): 6506-6513 Research Article Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, IraqAsaad Chasib Al-Atabi1, Nabeel Ahmed Al-Anbagi1, Hind A. Abdulghafoor2 and Murtadha abbas3*1Department of Veterinary Clinical Sciences, Faculty of Veterinary Medicine, University of Kufa, Al-Najaf, Iraq 2Department of Pathology and Forensic Medicine, College of Medicine, University of Fallujah, Baghdad, Iraq 3Department of Public Health, Faculty of Veterinary Medicine, University of Kufa, Al-Najaf, Iraq *Corresponding Author: Murtadha Abbas. Department of Public Health, Faculty of Veterinary Medicine, University of Kufa, Al-Najaf, Iraq. Email: murtadhaa.alghazali [at] uokufa.edu.iq Submitted: 13/08/2025 Revised: 11/11/2025 Accepted: 19/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
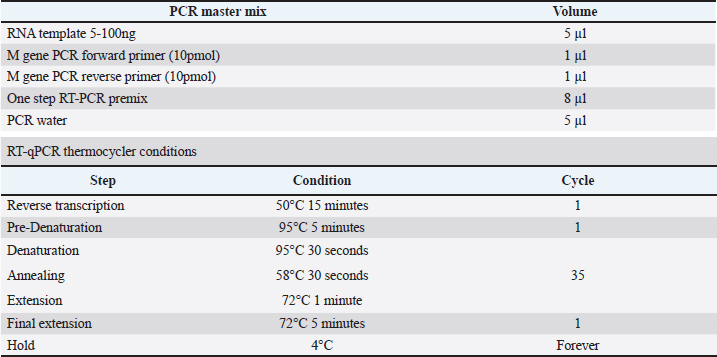
AbstractBackground: Numerous infectious agents and risk factors cause diseases of the respiratory system in farm animals. Parainfluenza virus type 3 is a common cause of respiratory tract infections in cattle, sheep, and goats worldwide. Aim: The goal of this study was to investigate the clinical and molecular identification, as well as phylogenetic tree analysis of Caprine para influenza virus 3 (CPIV3) in goats in various regions of Al-Najaf, Iraq. Methods: During December 2022 and November 2023, 100 nasal swabs showing signs of pneumonia were obtained from two separate goat fields belonging to the College of Veterinary Medicine, University of Kufa. The quantitative findings were presented as a chi-square test (χ2) was performed to assess the significance of the observed variances. A total of 100 nasal swabs were collected from goats, with CPIV3 possessing a molecular incidence of 38%. The clinical manifestations included fever, cough, difficulty breathing (rapid, short breaths), decreased appetite, and weight loss. There are no significant differences between the three areas of AL-Najaf province and sex (p > 0.05). Results: Infection rates were significantly higher in age groups older than 1 year. A phylogenetic study of the (M) gene in Caprine parainfluenza virus 3 revealed that the local Iraqi isolate accession numbers (OR978482.1–OR978486.1) were found to be closely related to NCBI-BLAST Caprine parainfluenza virus 3 strain JS2013 (KJ850331.1) at a total genetic alteration (0.01%). Conclusion: CPIV-3 is a serious respiratory infection that affects goats all over the world, including Iraq. Clinical manifestations of CPIV-3 infection range from mild respiratory problems to severe pneumonia, causing significant economic losses in the goat industry. Keywords: Caprine parainfluenza virus 3, Molecular detection, PCR, Al-Najaf, Iraq. IntroductionRespiratory diseases are a main concern in small ruminant flocks, producing significant economic losses in goat farming worldwide. These infections frequently affect both individual animals and entire herds, posing a persistent challenge to the industry (AL-abudy and Alrodhan, 2014; Al-Bayati et al., 2023). Also, one of the main factors contributing to morbidity and death in flocks of sheep and goats is the small ruminant respiratory syndrome and factors that promote an animal's risk to bacterial and viral infections and it is often includes a mix of infectious causes as well as predisposing management factors include stress, animal movement, exposure to unfavorable weather, and overcrowding, possibly leading to significant losses (Shanshol et al., 2018; Al Atabi et al., 2023). In sheep, goats, and cattle, respiratory infections are frequently caused by the parainfluenza virus type 3. Nevertheless, there are now four known serotypes of the parainfluenza virus, which belongs to the Paramyxoviridae family and the Respirovirus genus, which also includes the cattle, camels, sheep, and human parainfluenza viruses (Al-Bayati et al., 2023; Baghezza et al., 2024). CPIV-3 is a respiratory pathogen that affects goats worldwide, and the virus was first detected in the United States in 1959 (Branche and Falsey, 2016). It conversely can produce severe pneumonia in calves when combined with other bacterial and viral infections under stressful conditions; this pneumonia is commonly called "shipping fever" or "enzootic pneumonia” (Al Salihi and Yahia, 2018). However the severity of the disease can vary, ranging from mild respiratory symptoms to severe pneumonia and the clinical manifestations of CPIV-3 infection in goats include fever, coughing, nasal discharge, and respiratory distress, as well as pneumonia and bronchitis, resulting in major economic losses for the goat business, Understanding the epidemiology and molecular features of CPIV-3 is critical for successful management and prevention methods (Emikpe et al., 2019, 2017; Eser et al., 2023). Additionally, in young goats, the infection can be particularly devastating, leading to high mortality rates, and CPIV-3 infection can predispose goats to secondary bacterial infections, further exacerbating the direct contact and aerosols are the main transmission ways (Gafer et al., 2009; Sandip et al., 2014). Several studies have reported the presence of CPIV-3 in goats from different parts of the world in Mexico 2017 (Contreras-Luna et al., 2017), in Egypt 1971 (Gafer et al., 2009), and in Sudan (Saeed et al., 2016), these studies data reflect the susceptibility of sheep and goats to infection by PI3V due to a species cohabitation and exposition to predisposing factors as weather conditions, inappropriate handling, stress, and so on. Genetic material from Respiratory Syncytial Virus and PI3V was identified by Reverse Transcription Polymerase Chain Reaction in nasal swab samples from clinically healthy lambs and confirmed by sequencing and phylogenetic analysis (Eser et al., 2023). However, limited information is available regarding the prevalence and phylogenetic relationships of CPIV-3 strains in Iraqi goat populations. In this study, we aim to investigate the clinical and molecular characteristics of CPIV-3 in goats from Iraq by Molecular detection techniques. Materials and MethodsStudy areas and sample collectionThe current diagnostic study was done between December 2022 and November 2023. This cross-sectional study examined the detection of CPI3V in goats with a history of respiratory illness. A total of 100 nasal swab samples were collected from two separate goat fields belonging to the College of Veterinary Medicine, University of Kufa; they were entirely separate from each other. All samples were deposited in sterile test tubes, sealed in an ice bag, and added to special transport media (RNA was later stored at −20°C until analysis). The animals had not been vaccinated against CPIV3. All data (age, sex, and clinical signs) were recorded in the clinical examination. Molecular detection of Caprine parainfluenza virus 3Viral RNA was collected from respiratory viral transport medium (VTM) swab samples using the AccuZolTM Total RNA extraction kit (Bioneer, Korea) and following the manufacturer's instructions. The extracted RNA was calculated using a Nanodrop spectrophotometer, which measured the RNA concentration and purity at absorbance 260/280 nm at a ratio of (1.8–2.0) for pure RNA. RT-PCR was done for finding CPIV-3 based on the matrix protein (M) gene by using (One-Step RT-PCR Premix Kit) to amplify the expected size of 670bp for the matrix (M), which were designed in this study using the NCBI-Genbank database (GenBank: OR472985.1) and Primer3 plus. The protein gene sequence of CPIV-3 (Table 1) and done according to the company instructions. One-Step RT-PCR master mix was utilized; these RT-PCR reaction mixes were put in sterile 0.2 ml PCR tubes, combined, and centrifuged using an Exispin centrifuge for three minutes at 3,000 rpm, PCR thermocycler conditions were set by using Optimize protocol writer and according to primer annealing temperature as in (Table 2). Table 1. The one step RT-qPCR primers.
Table 2. One-step RT-PCR master mix and thermocycler condition.
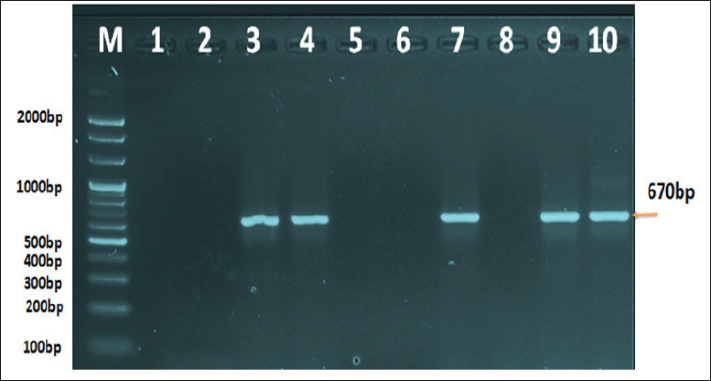
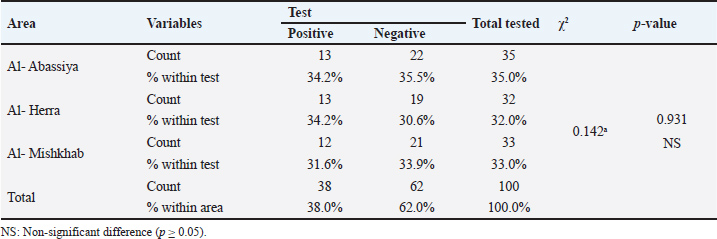
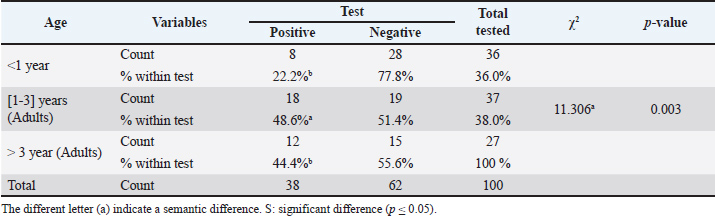
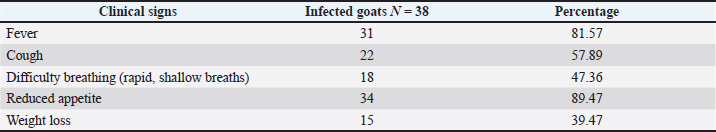
The following methods were used to examine the RT-PCR results using agarose gel electrophoresis: A 1% Agarose gel was made using 1X TBE, dissolved in the microwave for three minutes, and then allowed to cool to 50°C. Next, 3 µl of ethidium bromide stain was added to the solution of agarose gel. After adjusting the comb to its correct position, agarose gel solution was added to the tray. After fixing the gel tray in the electrophoresis chamber, 1X TBE buffer was added to it. Each comb well held 10 µl of the RT PCR product, while the first well had 5 µl of the 100 bp ladder. After that, an electric current was run for 1.5 hours at 100 V and 80 mA. The results of the RT-PCR were seen using a UV transilluminator. DNA sequencing methodMatrix protein (M) gene RT-PCR products that were positive for CPIV-3 were sent to Macrogen Company in Korea for DNA sequencing using an AB DNA sequencing System. Multiple alignment analysis based on Clustal W alignment analysis, NCBI-BLAST for homology sequence identity, and the phylogenetic tree UPGMA method (MEGA 6.0 version) were used in the DNA sequencing analysis. The one-step RT-qPCR primers are shown in Table 1, and the One-Step RT-PCR master mix and thermocycler conditions are shown in Table 2. Ethical approvalThis study was conducted in conformity with the rules of the Animal Ethics Committee of the Faculty of Veterinary Medicine at the University of Kufa (license number: 27865 on 13/10/2023). ResultsAccording to the PCR data, CPIV-3 infection was present in 38% (38 out of 100) of the three regions of AL Najaf (Al Abassiya, Al Herra, and Al-Mishkhab). Every location showed no significant difference; all animals had clinical examinations. Table 3 gives the molecular percentages of CPIV-3 positive samples for each location. The percentage of viral infection and risk factor was not significantly different between the study's male and female animals, as indicated by Table 4. Conversely, the age groups in Table 5 confirmed significant differences based on age groups, with the corresponding values being (22.2%, 48.6%, and 44.4 %). Table 6 displays the frequent clinical symptoms, which include fever, coughing, dyspnea (rapid, shallow breathing), decreased appetite, and weight loss. All positive samples were confirmed using the matrix gene (expected size bands: 670 bp); segments that matched the size range of 2,000–100 bp on the universal ladder were considered positive (Fig. 1).
Fig. 1. Agarose gel electrophoresis image that showed the RT-PCR product analysis of matrix protein (M) gene in Caprine parainfluenza virus 3 based A from VTM swab samples. Where M: marker (2,000–100bp) and Lane (1-10) showed some positive samples for M gene in Caprine parainfluenza virus 3 at (670bp) RT-PCR product. Table 3. The infection rate of CPIV-3 relation to the area by PCR.
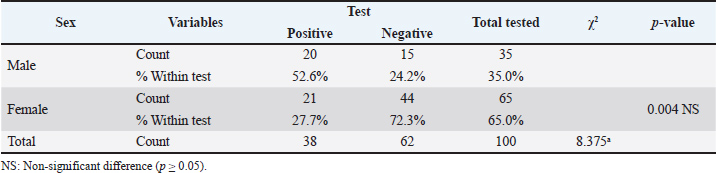
Table 4. The infection rate of CPIV-3 relation to the Sex by PCR.
Table 5. The infection rate of CPIV3 relation to the age.
Table 6. Clinical signs of infected goats with CPIV-3.
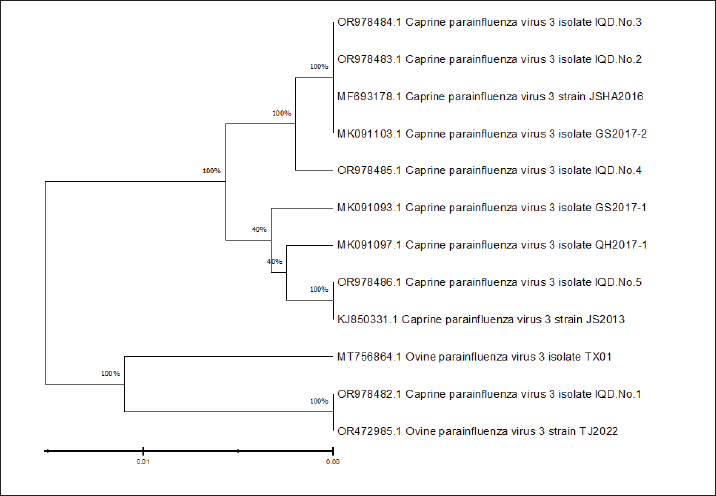
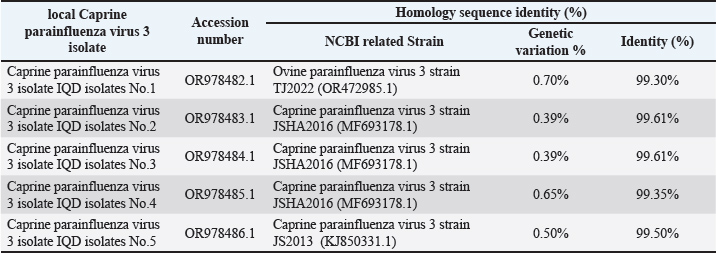
DNA sequence resultsDNA sequencing was done for genetic relationship analysis, Multiple sequence alignment of matrix protein (M) gene in local Caprine parainfluenza virus 3 isolate IQD isolates, as well as NCBI-Genbank Caprine and ovine parainfluenza virus 3 isolate. It revealed nucleotide alignment similarities and substitution mutations in the matrix protein (M) gene through isolates, as seen in Figure 1. The local Caprine parainfluenza virus 3 isolates (No.1) were shown to be highly linked to the NCBI-BLAST Ovine parainfluenza virus 3 strain TJ2022 (OR472985.1), according to the phylogenetic tree analysis. The NCBI-BLAST Caprine parainfluenza virus 3 strain JSHA2016 (MF693178.1) was closely associated with the local isolates of the virus (Nos. 2, 3, and 4). With just 0.01 percent genetic differences, the local isolates of the Caprine parainfluenza virus (No. 5) were determined to be closely linked to the NCBI-BLAST strain of the virus, JS2013 (KJ850331.1) (2). Additionally, there was a range of 99.35%–99.61% genetic homology sequence similarity between the NCBI-BLAST related Caprine parainfluenza virus 3 strains and the local IQD isolates of the virus, as shown in Table 7. As a final point, the Caprine parainfluenzavirus 3 isolate IQD isolates were submitted to Genbank and documented as accession numbers (OR978482.1-OR978486.1), Figure 2. The infection rate of CPIV-3 in relation to the Area by PCR showed in Table 3. The infection rate of CPIV-3 relation to the sex by PCR showed in Table 4. The infection rate of CPIV3 relation to the age showed in Table 5. The clinical signs of infected goats with CPIV-3 showed in Table 6, the NCBI-BLAST homology sequence identity percentage between local Caprine parainfluenza virus 3 isolate IQD isolates and NCBI-BLAST closed related Caprine and ovine parainfluenza virus 3 strain showed in Table 7.
Fig. 2. Phylogenetic tree analysis based on matrix protein (M) gene partial sequence in in local Caprine parainfluenza virus 3 isolate IQD isolates that used for genetic relationship analysis. Table 7. The NCBI-BLAST Homology Sequence identity percentage between local Caprine parainfluenza virus 3 isolate IQD isolates and NCBI-BLAST closed related Caprine and ovine parainfluenza virus 3 strain.
DiscussionParainfluenza-3 (PI-3) is the most common virus that affects the respiratory systems of sheep and goats worldwide, while it can cause pneumonia on its own (Rawat et al., 2019; Chakraborty et al., 2014). In this study, the identification of CPIV-3 as the causal agent of pneumonia in goats was examined. This study focused on the matrix protein (M) gene to analyze the genetic relationships within local Caprine parainfluenza virus 3 isolates. Caprine parainfluenza virus type 3 is an important respiratory infection in goats, producing disease with symptoms such as coughing, fever, and pneumonia (Baghezza et al., 2024). Females were more likely than males to have CPIV3, this outcome may be explained by the increased number of females (since breeding strategies were the major focus of the farms that were chosen (Baghezza et al., 2024). With regard to the animals' ages, adult goats had a higher incidence of CPIV-3 than goats in the other age groups (Table 1). These results agree with a study conducted in Mexican sheep (Contreras-Luna et al., 2017). The increased incidence of infection of CPIV3 in adults may be related to their history of several infections at this age (Saeed et al., 2016). On the other hand, it is also thought that animals reared for a longer period of time are more likely to have the illness. In the context of CPIV-3, RT-PCR is used to identify infected animals and isolate the viral RNA for further analysis, and the RT-PCR, culture, viral isolation, and electron microscopy are used to determine the incidence of the CPIV-3 virus in naturally infected flocks worldwide (Mao et al., 2019; Rawat et al., 2019; Ren et al., 2023). The M gene is relatively conserved across different CPIV3 strains, making it a reliable target for broad-spectrum detection (Sandip et al., 2014; Saeed et al, 2016). A similar related study concerning the epidemiological investigation and phylogenetic analysis of caprine parainfluenza virus type 3 in sheep of China (Mao et al., 2019; Li et al., 2016). The analysis involved aligning these sequences with M gene sequences of related CPIV-3 and ovine parainfluenza virus 3 isolates retrieved from NCBI-Blast. The findings revealed valuable insights into the genetic makeup of the local CPIV-3 isolates. Phylogenetic tree analysis further elucidated the evolutionary relationships between these isolates and reference strains. Interestingly, isolate No. 1 clustered most closely with the ovine parainfluenza virus 3 strain TJ2022 (OR472985.1) from NCBI-BLAST, suggesting a potential interspecies transmission event. Isolates No. 2, No. 3, and No. 4 displayed a closer genetic relationship to the caprine parainfluenza virus 3 strain JSHA2016 (MF693178.1) from NCBI-BLAST. Notably, isolate No. 5 exhibited the highest degree of homology (99.61% genetic identity) with the caprine parainfluenza virus 3 strain JS2013 (KJ850331.1) from NCBI-BLAST, with a minimal genetic divergence of only 0.01% (Fig. 2). This close association suggests a potentially recent circulation of a highly similar CPIV3 strain within the local goat population. Overall, the homology sequence identity between the local IQD isolates and reference strains ranged from 99.35% to 99.61% (Table 1), indicating a high degree of genetic conservation within the M gene. This observation aligns with previous studies highlighting the M gene as a reliable target for CPIV3 detection due to its relative conservatism (Al Salihi and Yahia, 2018; Werid et al., 2024). However, the identification of isolate No. 1 clustering with an ovine strain underscores the potential for genetic variation and cross-species transmission of CPIV3, warranting further investigation. Finally, the successful submission of the IQD isolates to GenBank under accession numbers OR978482.1-OR978486.1 contributes valuable genetic data to the global repository of CPIV3 sequences. This data resource can be instrumental for future studies aiming to understand the global distribution and evolution of CPIV3 strains, facilitating the development of more comprehensive diagnostic tools and vaccines. ConclusionCPIV-3 is a serious respiratory infection that affects goats all over the world, including Iraq. Clinical manifestations of CPIV-3 infection range from mild respiratory problems to severe pneumonia, causing significant economic losses in the goat industry. The PCR assay demonstrated high sensitivity and specificity in detecting the targeted gene. Further research into the sequencing of the complete CPIV-3 genome circulating in various species in Iraq is recommended. It was assumed that vaccination should be done for protection. AcknowledgmentsThe author is inclined to thank the University of Kufa Faculty of Veterinary Medicine, each veterinarian working in the hospital's laboratory, and the authors established no funding for this study. Conflict of interestThere are no conflicts of interest numerous of the authors concerning the publication of this manuscript. FundingNone. Authors' contributionAll listed authors have made significant intellectual and academic contributions to this study. Each author has reviewed and approved the final manuscript for publication. Data availabilityAll the datasets formed or examined during this work are included in this publication form. ReferencesAl Salihi, K. and Yahia, Z. 2018. Histopathological, Bacteriological and Molecular study of enzootic respiratory complex of Small Ruminants Slaughtered at Al Muthanna abattoir. 28(1), 102–110. AL-Abudy, L.S. and Alrodhan, M.A. 2014. Seroprevalence and molecular detection of Bovine Parainfluenza-3 Virus (BPI-3V). AL Qadisiya J. Vet. Med. Sci. 13, 27–32. Baghezza, S., Azizi, A., Derrar, F., Adnane Smadi, M., Djeghim, H., Ghougal, K., Gradi, E.A., Bennoune, O. and Mamache, B. 2024. Epidemiological Study of Bovine Parainfluenza 3 Virus in Sheep: seroprevalence, Risk Factors, and Distribution in Two Regions of Algeria. Iran. J. Vet. Med. 18(2), 159–168. Branche AR. and Falsey, A.R. 2016. Parainfluenza virus infection.In Seminars in respiratory and critical care medicine. Thieme Medical Publishers, 37, pp: 538–54. Chakraborty, S., Kumar, A., Tiwari, R., Rahal, A., Malik, Y., Dhama, K., Pal, A. and Prasad, M. 2014. Advances in diagnosis of respiratory diseases of small ruminants. Vet. Med. Int. 2014. Contreras-Luna, M.J., Ramírez-Martínez, L.A., Sarmiento Silva, R.E., Cruz Lazo, C., Pérez Torres, A. and Sánchez-Betancourt, J.I. 2017. Evidence of respiratory syncytial virus and parainfluenza-3 virus in Mexican sheep. VirusDisease 28(1), 102–110. Emikpe, B.O., Jarikre, T.A., Akpavie, S.O., Opoku-Agyemang, T., Asare D. and Folitse, R.D. 2019. Histological and immunohistochemical assessments of pneumonia in sheep slaughtered at Ibadan, Nigeria and Kumasi, Ghana. J. Immunoass. Immunochem. 40(3), 300–313. Eser, G., Sağlam, Y.S. and Yıldırım, S. 2023. Immunohistochemical investigation of parainfluenza 3 virus in sheep pneumonia.In Veterinary Research Forum. Urmia, Iran: Faculty of Veterinary Medicine, Urmia University, 14, p: 461. Gafer, J.A.M., Hussein, H.A. and Reda, I.M. 2009. Isolation and characterization of PI-3 virus from sheep and goats. Int. J. Virol. 5(1), 28–35. Jarikre, T.A. and Emikpe, B.O. 2017. First report of immunohistochemical detection of Peste des petit ruminants, parainfluenza 3 and respiratory syncytial viral antigens in lungs of Nigerian goats. J. Immunoass. Immunochem. 38(5), 555–568. Li, J., Li, W., Mao, L., Hao, F., Yang, L., Zhang, W. and Jiang, J. 2016. Rapid detection of novel caprine parainfluenza virus type 3 (CPIV3) using a TaqMan-based RT-qPCR. J. Virol. Methods 236, 126–131. Al-bayati, H.A., Shamkhi, G.J., Al-Aidy, S.R. and Gharban, H.A.J. 2023. Serological Detection, Isolation and Molecular Confirmation of Parainfluenza Virus-3 in Camels, Iraq. Bionatura 8(1), 1–10. Mao, L., Yang, L., Li, W., Liang, P., Zhang, S., Li, J., Sun, M., Zhang, W., Wang, L., Zhong, C., Liu, M., Jiang, J., Cai, X. and Luo, X. 2019. Epidemiological investigation and phylogenetic analysis of caprine parainfluenza virus type 3 in sheep of China. Transbound. Emerg. Dis. 66(3), 1411–1416. Rawat, N., Gilhare, V.R., Kushwaha, K.K., Hattimare, D.D., Khan, F.F., Shende, R.K. and Jolhe, D.K. 2019. Isolation and molecular characterization of Mannheimia haemolytica and Pasteurella multocida associated with pneumonia of goats in Chhattisgarh. Vet. World. 12(2), 331. Ren, Y., Tang, C. and Yue, H. 2023. Prevalence and molecular characterization of bovine parainfluenza virus type 3 in cattle herds in China. Animals 13(5), 793. Saeed, I., Ali, Y., Taha, K., Mohammed, N., Nouri, Y., Mohammed, B., Mohammed, O., Elmagboul, S. and Alghazali, F. 2016. Para influenza virus 3 infection in cattle and small ruminants in Sudan. J. Adv. Vet. Anim. Res. 3(3), 236–241. Sandip, C., Amit, K., Ruchi, T., Anu, R., Yash, M., Kuldeep, D. and Minakshi, P. 2014. Advances in diagnosis of respiratory diseases of small ruminants. Vet. Med. Int. Shanshol, R.H., Hadi, A.H., Ali, A.A. 2018. Histopathological, Bacteriological and Molecular study of enzootic respiratory complex of Small Ruminants Slaughtered at Al Muthanna abattoir. Mirror Res. Vet. Sci. Anim. 1, 51. Tiwari, K., Cornish, C., Gamble, B., Thomas, D. and Sharma, R.N. 2016. Seroprevalence of bovine parainfluenza virus Type 3 (bPI-3V) in ruminants from Grenada. Open J. Vet. Med. 6(2), 23–27. Werid, G.M., Van, T.D., Miller, D., Hemmatzadeh, F., Fulton, R.W., Kirkwood, R. and Petrovski, K. 2024. Bovine Parainfluenza-3 Virus Detection Methods and Prevalence in Cattle: a Systematic Review and Meta-Analysis. Animals 14(3), 494. | ||
| How to Cite this Article |
| Pubmed Style Al-atabi AC, Al-anbagi NA, Abdulghafoor HA, Abbas M. Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Vet. J.. 2025; 15(12): 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 Web Style Al-atabi AC, Al-anbagi NA, Abdulghafoor HA, Abbas M. Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. https://www.openveterinaryjournal.com/?mno=277102 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.35 AMA (American Medical Association) Style Al-atabi AC, Al-anbagi NA, Abdulghafoor HA, Abbas M. Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Vet. J.. 2025; 15(12): 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 Vancouver/ICMJE Style Al-atabi AC, Al-anbagi NA, Abdulghafoor HA, Abbas M. Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 Harvard Style Al-atabi, A. C., Al-anbagi, . N. A., Abdulghafoor, . H. A. & Abbas, . M. (2025) Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Vet. J., 15 (12), 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 Turabian Style Al-atabi, Asaad Chasib, Nabeel Ahmed Al-anbagi, Hind A. Abdulghafoor, and Murtadha Abbas. 2025. Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Veterinary Journal, 15 (12), 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 Chicago Style Al-atabi, Asaad Chasib, Nabeel Ahmed Al-anbagi, Hind A. Abdulghafoor, and Murtadha Abbas. "Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq." Open Veterinary Journal 15 (2025), 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 MLA (The Modern Language Association) Style Al-atabi, Asaad Chasib, Nabeel Ahmed Al-anbagi, Hind A. Abdulghafoor, and Murtadha Abbas. "Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq." Open Veterinary Journal 15.12 (2025), 6506-6513. Print. doi:10.5455/OVJ.2025.v15.i12.35 APA (American Psychological Association) Style Al-atabi, A. C., Al-anbagi, . N. A., Abdulghafoor, . H. A. & Abbas, . M. (2025) Caprine parainfluenza virus 3 clinical, molecular detection, and phylogenetic tree analysis from goats of AL-Najaf, Iraq. Open Veterinary Journal, 15 (12), 6506-6513. doi:10.5455/OVJ.2025.v15.i12.35 |