| Research Article | ||
Open Vet. J.. 2025; 15(12): 6786-6800 Open Veterinary Journal, (2025), Vol. 15(12): 6786-6800 Research Article Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterizationNuo Xu1 , Hannuo Li1, Xinyu Wang1, Hui Su1, Yuhang Zheng1, Wanli Sha1,2,3, Baishuang Yin1,2,4 and Wenlong Dong1,2,3*1College of Animal Science and Technology, Ji.3Lin Agricultural Science and Technology College, Jilin, China 2Jilin Provincial Key Laboratory of Preventive Veterinary Medicine, Jilin, China 3Jilin Province Technology Innovation Center of Pig Ecological Breeding and Disease Prevention and Control, Jilin, China 4Jilin Province Cross Regional Cooperation Technology Innovation Center of Porcine Main Disease Prevention and Control, Jilin, China *Corresponding Author: Wenlong Dong. College of Animal Science and Technology, JiLin Agricultural Science and Technology College, Jilin, China. Email: dongwenlong888 [at] jlnku.edu.cn Submitted: 11/08/2025 Revised: 11/11/2025 Accepted: 18/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
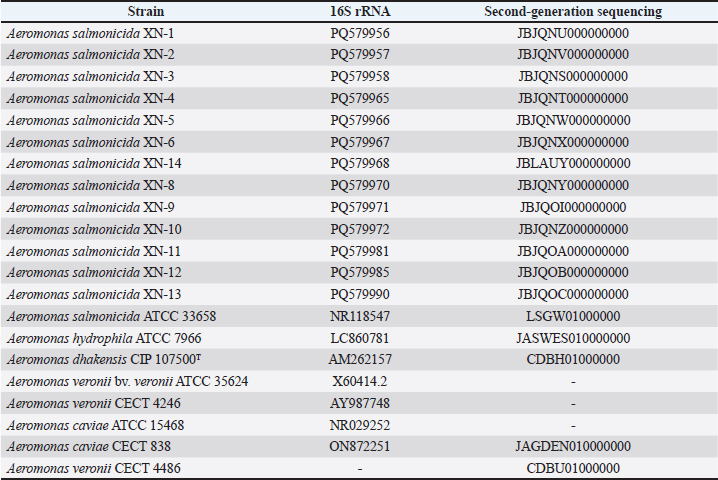
AbstractBackground: The emergence of Aeromonas salmonicida in swine respiratory disease represents a significant animal health concern with potential public health implications due to the risk of zoonotic antimicrobial resistance transmission. Aim: This study aimed to characterize the phenotypic and genotypic antimicrobial resistance profiles of A. salmonicida isolated from pigs with respiratory disease in Jilin Province, China, to assess its potential threat to animal and public health. Methods: Between January 2023 and December 2023, 786 pigs with mild respiratory disease were randomly sampled from 19 conveniently selected farms in Jilin Province, China. The presence of one or more mild clinical signs, such as nasal discharge, coughing, labored breathing, lethargy, or anorexia, defines this condition. From these pigs, 13 A. salmonicida strains were isolated from the upper respiratory tracts. The isolates’ taxonomic identification was systematically determined through a polyphasic approach. This included 16S rRNA gene sequencing, phylogenetic analyses based on the gyrB and rpoB housekeeping genes, and Average Nucleotide Identity analysis. Antimicrobial susceptibility testing against 21 agents was performed using the CLSI-standardized agar dilution minimum inhibitory concentration assay, with Escherichia coli ATCC 25922 as the quality control. Second-generation sequencing analysis was conducted, followed by a comprehensive screening of antibiotic resistance genes using the Comprehensive Antibiotic Resistance Database database (Resistance Gene Identifier module). Results: Antimicrobial susceptibility was interpreted according to the Clinical and Laboratory Standards Institute guidelines. All 13 isolates demonstrated complete resistance (13/13, 100.00%) to β-lactams (meropenem, aztreonam, cefazolin, and ampicillin) and sulfonamides (sulfamethoxazole and trimethoprim). In contrast, all isolates were susceptible to fosfomycin and rifampicin (13/13, 100%). Variable susceptibility to florfenicol was observed (9/13, 69.23%). Genomic analysis revealed the spread of resistance genes (mcr−3.16, cphA5, blaOXA-956) in A. salmonicida. The key mechanisms include enzymatic inactivation and efflux pumps. Biocide resistance genes (qacEdelta1) indicate disinfectant co-selection. No explicitly novel genes were reported; however, emerging variants (e.g., blaOXA-956) were identified. Conclusion: This study provides the first comprehensive antimicrobial resistance profile of porcine-associated A. salmonicida. Our results demonstrated that these bacteria were resistant to almost all commonly used antibiotics. The genes conferring this extensive resistance were located on mobile genetic elements, meaning they can spread to other bacteria, which poses a significant threat to both animal and public health. To address this, we recommend the following: (1) implementing antibiotic susceptibility testing in veterinary practice to guide treatment choices and avoid ineffective drugs; (2) prioritizing prevention through improved farm hygiene and biosecurity to reduce infection rates; and (3) establishing coordinated surveillance across human, animal, and environmental sectors to monitor the spread of these resistant bacteria. Keywords: Aeromonas salmonicida, Antimicrobial resistance, Pig, Resistance genes, Second-generation sequencing analysis. IntroductionAeromonas salmonicida (A. salmonicida) is a Gram-negative facultative anaerobe and one of the earliest documented bacterial pathogens in fish species (Vincent et al., 2022). The genus Aeromonas exhibits significant genomic diversity, and the 16S rRNA gene, while useful for genus-level identification, often lacks the resolution to reliably distinguish between closely related species due to its high conservation (Yáñez et al., 2003; Küpfer et al., 2006). Consequently, multilocus sequence analysis targeting housekeeping genes like gyrB and rpoB, average nucleotide identity (ANI), or phylogenomics are now considered essential for accurate species delimitation within this genus (Yáñez et al., 2003; Küpfer et al., 2006). The discovery of thermophilic A. salmonicida in pigs represents a notable shift from its traditional characterization as a psychrophilic fish pathogen. This study investigates the clinical and public health significance of thermophilic A. salmonicida strains isolated from pigs—a species in which infections may lead to economic losses and pose zoonotic risks through potential Antimicrobial Resistance transmission. Although A. salmonicida includes multiple psychrophilic subspecies (Pavan et al., 2000), we focused exclusively on thermophilic strains capable of thriving at mammalian body temperatures. To the best of our knowledge, this is the first study to conduct comprehensive antimicrobial resistance profiling of thermophilic A. salmonicida of porcine origin, providing urgently needed data to inform veterinary treatment protocols and antibiotic stewardship policies. Our work underscores the need to monitor these emerging pathogens to mitigate their impact on animal health and food safety. In recent years, numerous novel mesophilic A. salmonicida strains have been identified from diverse hosts, underscoring its expanding clinical relevance. Its recovery from human patients—including a 34-year-old woman with bacteremia and a 67-year-old immunocompetent man with a skin infection—highlights its zoonotic capacity and ability to cause opportunistic infections in humans (Tewari et al., 2014; Vincent et al., 2019). One Health significance is its isolation from livestock, such as from the spleen of a diseased pig (Long et al., 2023). Porcine A. salmonicida isolates are of particular concern as pigs may serve as reservoirs for the amplification and dissemination of AMR determinants at the critical human-animal-environment interface. The species A. salmonicida exhibits a notable capacity for AMR, encompassing both intrinsic and acquired mechanisms. Phenotypically, resistance to a range of antibiotics, including β-lactams, tetracyclines, sulfonamides, and quinolones, has been documented. Genotypically, the remarkable genomic plasticity of the genus Aeromonas, as evidenced by phylogenetic studies based on housekeeping genes like gyrB and rpoB, underpins its potential for AMR evolution (Yáñez et al., 2003; Küpfer et al., 2006). Although the potential acquisition and dissemination of resistance genes via mobile genetic elements is a recognized trait in Aeromonas, definitive resistance determinants are frequently encoded within the core chromosome. A key genotypic resistance mechanism involves the production of diverse β-lactamases (encompassing molecular classes A, B, C, and D), which confer resistance to β-lactam antibiotics. Furthermore, resistance to last-resort agents, such as colistin, has been reported in Aeromonas, involving both chromosomal mutations and plasmid-mediated genes (Goncalves-Pessoa et al., 2022). The presence of such multifaceted resistance profiles in porcine isolates underscores the potential role of this niche in the broader AMR landscape. Despite this recognized potential, there is a striking paucity of data exist on the multidrug resistance profiles and resistance gene content of porcine-derived A. salmonicida. Therefore, a comprehensive understanding of the mechanisms of AMR is essential for developing effective strategies to control infections in economically important livestock and for accurately assessing the full scope of zoonotic risk. This study aimed to isolate and characterize the phenotypic and genotypic antimicrobial resistance profiles of thermophilic A. salmonicida strains recovered from pigs in Jilin Province, China. We hypothesized that these strains harbor a diverse array of transferable multidrug-resistant genes, which may contribute to their persistence in swine production environments and pose a potential threat to public health. Materials and MethodsIsolation and identification of porcine-derived A. salmonicida strainsThis study reported the isolation of 13 A. salmonicida strains from the upper respiratory tracts of 786 randomly sampled pigs from 19 farms in Jilin Province, China. Farm selection was based on convenience and willingness to participate. Sampled pigs exhibited mild respiratory disease, defined by the presence of one or more early-stage clinical signs, such as nasal discharge, coughing, or increased respiratory rate. The nasopharyngeal swabs were collected and immediately placed into a sterile tube, stored at 4 °C, and processed within 12 hours. Samples were cultured on brain heart infusion (BHI) agar and incubated at 37°C for 12 hours to maximize the recovery of pathogens, a temperature selected for its physiological relevance to the porcine host. Genomic Deoxyribonucleic Acid was extracted from pure bacterial cultures using the Bacterial Genomic DNA Rapid Extraction Kit (Aidlab Biotechnologies, China), according to the manufacturer’s protocol for Gram-negative bacteria. The extraction procedure involved lysis of bacterial cells with a proprietary buffer (Solution GY) and Proteinase K at 70°C, followed by DNA binding to a silica membrane column in the presence of a binding buffer (Solution CB) and isopropanol. The bound DNA was then washed twice and eluted in a preheated EB. DNA concentration and purity (A260/A280 ratio) were measured using a NanoDrop One spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The integrity of the DNA was confirmed by 1% agarose gel electrophoresis. Polymerase Chain Reaction was performed in a 50 μl reaction mixture containing 25 μl of 2 × Taq Master Mix, 1 μl of each primer (forward: 5'-AGAGTTTGATCCTGGCTCAG-3'; reverse: 5'-GGTTACCTTGTTACGACTT-3'), 1 μl of template DNA, and 22 μl of sterile distilled water. The PCR amplification was performed under the following conditions: initial denaturation at 94°C for 5 minutes; 30 cycles of denaturation at 95°C for 1 minute, annealing at 55°C for 1 minute, and extension at 72°C for 1 minute; followed by a final extension at 72°C for 10 minutes. The PCR products were verified by 1% agarose gel electrophoresis prior to sequencing to confirm the amplification of a single band of the expected size (~1,500 bp). The 16S rRNA gene amplicons were generated with the universal primers 27F (5′-AGAGTTTGATCCTGGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′), which target the V3-V4 hypervariable regions (Moreno et al., 2002). These amplicons were purified and bidirectionally sequenced by General Biol (Anhui, China) on a Thermo Fisher Scientific 3730XL DNA Analyzer. The chromatograms obtained from both sequencing directions were assembled and manually curated to obtain the final consensus sequence for strain identification. Phylogenetic and genomic analysesA comprehensive analysis was conducted at both the single-gene and genome levels to elucidate the phylogenetic of our isolates. First, a phylogenetic tree was constructed based on the 16S rRNA gene sequences. The dataset comprised 66 strains, including 13 experimental A. salmonicida isolates, the standard strain A. salmonicida ATCC 33568, and 52 reference strains. The reference strains were retrieved from the National Center for Biotechnology Information. The reference strains were selected based on the following criteria: (i) host origin (encompassing a wide range of species such as fish, pigs, and humans), (ii) geographical diversity (to represent global distribution), and (iii) high 16S rRNA sequence identity and sequence length comparable to our isolates. Phylogenetic reconstruction was performed using the Neighbor-Joining algorithm in Molecular Evolutionary Genetics Analysis (MEGA) version 11. The resulting tree was visualized and annotated using tvBOT with the following parameters: circular layout; modified layers displaying leaf names, color blocks for leaf labels, and added symbols; and specific text in italics. The final tree was exported in the Portable Network Graphics format. To further investigate the relationships between our experimental strains and closely related species, we conducted phylogenetic analyses using the gyrB and rpoB genes to achieve higher resolution. This targeted analysis included a dataset of 22 strains: our 13 experimental isolates, the A. salmonicida ATCC 33568 standard strain, and 8 closely related species. The corresponding gene sequences for these strains were retrieved from the National Center for Biotechnology Information (NCBI) database. Then, we constructed neighbor-joining trees using MEGA11, and the resulting trees were visualized and annotated using tvBOT with the same set of parameters as before. Second-generation sequencing data from 18 strains were used to calculate the ANI. The dataset included 13 experimental A. salmonicida strains, the standard strain A. salmonicida ATCC 33568, and four closely related QC strains: Aeromonas hydrophila (A. hydrophila) ATCC 7966, Aeromonas dhakensis (A. dhakensis) CIP 107500T, Aeromonas caviae (A. caviae) CECT 838, and Aeromonas veronii (A. veronii) CECT 4486. Bacterial identification was performed using the online calculator provided by CJ Bioscience (Yoon et al., 2017). The ANI matrix was visualized using Origin 2024. The genome sequences and associated raw reads have been deposited in the NCBI database, with detailed accession numbers for each strain provided in Table 1. Table 1. NCBI accession numbers for 16S rRNA sequences and second-generation sequencing data of A. salmonicida isolates and QC strains.
Determination of the susceptibility of antimicrobial agentsThe purity and identity of all 13 A. salmonicida isolates were confirmed before antimicrobial susceptibility testing. Each isolate was subcultured onto fresh BHI agar plates and incubated at 37°C for 12 hours. Purity was assessed by examining individual colonies’ morphology, size, and coloration. Subsequently, the taxonomic identity of each purified isolate was reconfirmed by amplifying and sequencing the 16S rRNA from single colonies, using the primers and protocols described in the section on strain identification. The minimum inhibitory concentration (MIC) of the A. salmonicida isolates against 21 antimicrobial agents was determined by the agar dilution method on Mueller–Hinton Agar (MHA) plate. Following inoculation, the plates were incubated at 37°C for 12 hours before reading the MIC results. The antimicrobial panel included aztreonam, sulfamethoxazole, meropenem, cefazolin, tetracycline, levofloxacin, chloramphenicol, polymyxin B, trimethoprim, gentamicin, azithromycin, florfenicol, amikacin, ciprofloxacin, ceftazidime, tobramycin, ceftriaxone, cefotaxime, fosfomycin, ampicillin, and rifampicin. Escherichia coli (E. coli) ATCC 25922 was used as the quality control strain and was included in every agar dilution plate batch. The measured MIC values for the control strain for all antimicrobial agents were within the acceptable quality control ranges specified by the CLSI M100 (2023) guidelines, validating the test conditions. To ensure reproducibility, all tests were performed in triplicate using biologically independent cultures. The MIC interpretive criteria (breakpoints) were applied according to the CLSI guidelines (CLSI M45, 2016; CLSI M100, 2023; CLSI M42, 2011), as detailed in the annotations of Figure 4. The breakpoints for Enterobacterales (CLSI M100, 2023) were applied for antimicrobials lacking specific Aeromonas spp. breakpoints (polymyxin B, azithromycin, tobramycin, ampicillin, cefazolin, and fosfomycin), as explicitly recommended and justified by the derivation principles for Aeromonas spp. in CLSI M45 (2016). Data visualization and statistical analyses were conducted using the Origin software (version 2024; OriginLab Corporation, Northampton, MA, USA).
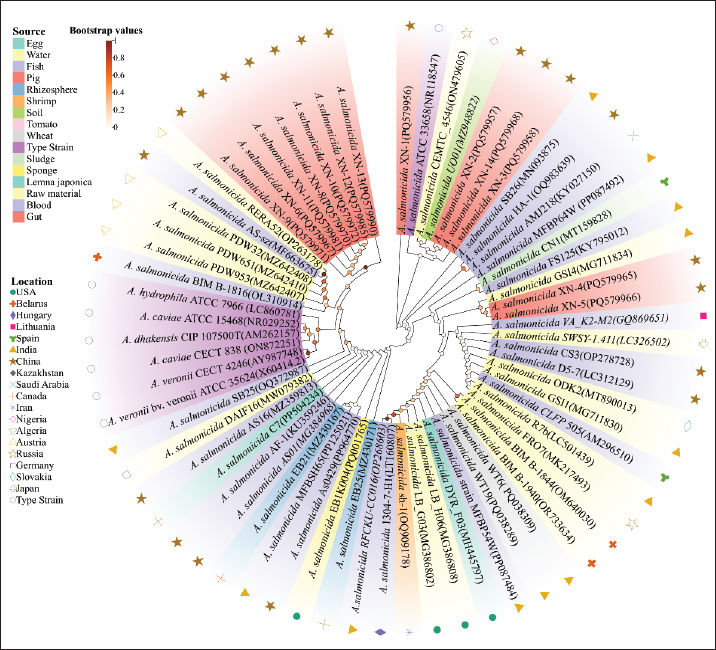
Fig. 1. Phylogenetic tree based on the 16S rRNA sequences of the isolates and selected species. The phylogenetic tree comprised 60 strains of A. salmonicida and 6 closely related quality control (QC) strains. Among the A. salmonicida strains, 13 were experimental isolates obtained from Jilin Province, China, one was the reference strain A. salmonicida ATCC 33658, and the remaining 46 were derived from 16S rRNA sequences of A. salmonicida isolated from various species in disparate areas, sourced from the NCBI database. The leaf color indicates the isolated species of the strains. The circles on the branches are colored to show the bootstrap values. The symbols in the outermost circle indicate the region from which the strains were isolated.
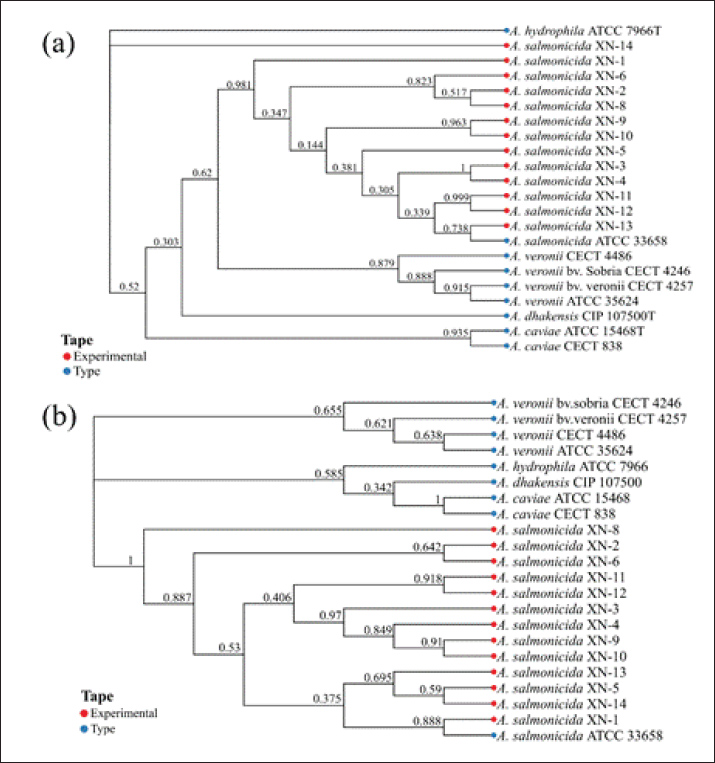
Fig. 2. Phylogenetic analysis of A. salmonicida and closely related species based on housekeeping genes. (a) Neighbor-joining tree reconstructed from gyrB gene sequences. (b) Neighbor-joining tree reconstructed from rpoB gene sequences. Both trees included 13 experimental strains (red symbols), the A. salmonicida QC strain ATCC 33658 (blue symbol), and six closely related species (blue symbols). The numbers at the nodes represent bootstrap values. The symbols preceding the strain names denote the strain type: experimental (red) and quality control (QC)/reference (blue) strains.
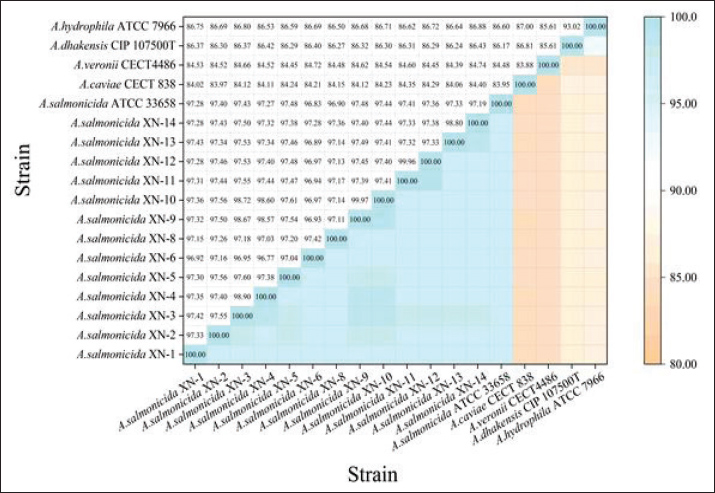
Fig. 3. ANI heatmap of the Aeromonas strains. This matrix illustrates the pairwise OrthoANI values among 13 experimental A. salmonicida strains, the A. salmonicida QC strain ATCC 33568, and four closely related QC strains. The color intensity in the lower-right heatmap corresponds to the ANI value, which is also numerically displayed in the upper-left quadrant. The genomic data used for this analysis were sourced from the NCBI database (for accession numbers, Table 1). The OrthoANI algorithm was used for the analysis, and the visualization was generated with Origin 2024 software.
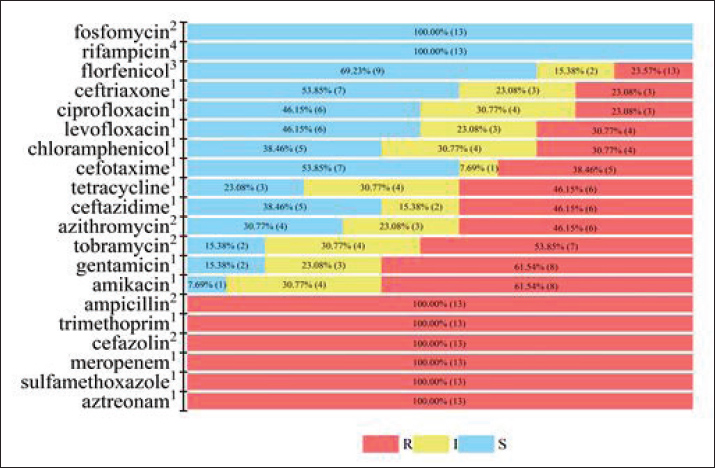
Fig. 4. Resistance rates of A. salmonicida to 20 antibiotics This graph presents the resistance profile of 13 strains of A. salmonicida, isolated from pig farms in Jilin Province, China, to 20 antibiotics. Before performing the MIC test, each isolate was subcultured in fresh BHI broth and incubated at 37°C for 12 hours. Antimicrobial susceptibility testing was performed using the agar dilution method on MHA. The QC strain was E. coli ATCC 25922. Red, resistance; yellow, intermediary; and blue, sensitive. The numerical value within the brackets represents the specific region identifier. The following corner markers illustrate the methodology for identifying the MIC inflection point: 1: M45 (CLSI, 2016, Aeromonas spp.); 2: M100 (CLSI, 2023, Enterobacterales); 3: M42 (CLSI, 2011, Aeromonas salmonicida); 4: (Esteve et al., 2012). To further screen for carbapenemase production phenotypically, the modified Carbapenem Inactivation Method (mCIM) was performed on all 13 experimental A. salmonicida isolates, following the guidelines outlined in CLSI M100 (33th ed., 2023). Briefly, a 1 μl loopful of bacteria from an overnight culture on sheep blood agar was emulsified in 2 ml of tryptic soy broth for each isolate. A 10 μg meropenem disk was then immersed in the bacterial suspension. After incubation at 37°C for 4 hours, the disk was retrieved and placed onto an MHA plate seeded with the meropenem-susceptible E. coli ATCC 25922 indicator strain. The inhibition zone around the disk was measured after an additional 18 hours of incubation at 37°C. All tests were performed in triplicate to ensure reproducibility. Second-generation sequencing analysisBacterial cultures were maintained under controlled incubation conditions until the logarithmic growth phase was achieved. We subjected the cell culture media to centrifugation at 7,000 rpm for 15 minutes to prepare them. The quality of the extracted DNA was verified using a NanoDrop spectrophotometer (A260/A280 ratios between 1.8 and 2.0) and 1% agarose gel electrophoresis. Qualified DNA samples were subjected to library preparation using the TrueLib DNA Library Rapid Prep Kit for Illumina (Yitaike Biotech Co., Ltd.) and subsequently submitted to Bemac Biotechnology Co. (Qingdao, China) for Illumina sequencing, which generated an average sequencing depth of 100× per genome. Raw sequencing reads were processed using fastp (version 0.23.2) to remove adapters and low-quality bases with the following parameters: -Q -y -g -Y 10 -l 100 -b 150 -B 150 (Haibin et al., 2012). Subsequently, reads were further filtered using fastq_filter_by_Qxx to retain only high-quality sequences with a minimum Q30 score threshold of 85% over a 30-bp sliding window (-q 0.85 -w 30). For genome assembly, the filtered reads were assembled using the Spades v3.6.2 software (Prjibelski et al., 2020). The assembly quality was assessed based on the contig N50 values. Furthermore, antibiotic resistance genes were identified using the Resistance Gene Identifier tool (Web portal version 6.0.5) against the Comprehensive Antibiotic Resistance Database (CARD, version 4.0.1). The analysis involved the following steps and criteria: Open Reading Frames were first predicted from the input DNA sequences using Prodigal; homolog detection was then conducted using DIAMOND. The results were filtered to include only "Perfect" and "Strict" ('Perfect' requires all amino acids to match the CARD reference and “Strict” requires a bitscore of ≥500) hits. The presence/absence of Antibiotic Resistance Genes was summarized into a binary matrix and used to construct a dendrogram illustrating strain clustering based on ARG profile similarity. Clustering analysis and visualization were performed in Origin 2024 using the UPGMA and Euclidean distance, with both circular and radial dendrogram layouts applied to visualize natural grouping patterns. Ethical approvalVerbal consent was obtained from the farm manager prior to sample collection, nasal swab samples were collected from pigs exhibiting mild respiratory symptoms by veterinarians. The sampling method was non-invasive and strictly adhered to animal welfare guidelines, ensuring that any potential distress to the animals was minimized. The study was approved by the Institutional Animal Ethics Committee of Jilin Agricultural Science and Technology College (Reference number: LLSC202301055), with an approved project period from January 2023 to December 2024. All procedures performed on the animals followed the ethical standards of the institution. ResultsIdentification and phylogenetic analysis of isolates of A. salmonicidaSequencing data were analyzed using the Basic Local Alignment Search Tool (BLAST) on the NCBI platform. The initial taxonomic identification of all 13 strains was confirmed by 16S rRNA gene sequencing, which revealed >99.00% sequence identity with A. salmonicida reference sequences. To further validate this identification, we performed multilocus sequence typing (MLST) based on the second-generation sequencing data of the 13 experimental strains using the PubMLST database. All strains were unequivocally assigned as A. salmonicida using the MLST scheme, thereby consolidating the initial BLAST-based identification. Phylogenetic analysis based on 16S rRNA gene sequences was conducted using a dataset of 66 strains, including 13 porcine-derived A. salmonicida experimental isolates, the standard strain A. salmonicida ATCC 33568, 46 A. salmonicida isolates from diverse geographical and host origins, and 6 standard strains of closely related species. The phylogenetic tree revealed distinct intraspecific clustering of the experimental strains into three major groups and one independent branch (Fig. 1; experimental isolates highlighted in red). Cluster I, comprising XN-6, XN-8, XN-9, XN-10, XN-11, XN-12, and XN-13 isolates, was strongly supported with a bootstrap value of 0.99. Cluster II, which consists of XN-4 and XN-5, received a moderate bootstrap value of 0.5. Cluster III, which included XN-2, XN-3, and XN-14, exhibited weak support with a bootstrap value of 0.2. In contrast, XN-1 formed an independent branch with no significant bootstrap support. Notably, all 13 experimental isolates clustered distantly from the other referenced A. salmonicida strain, indicating considerable phylogenetic divergence. For interspecific relationships, the closely related species formed a separate, well-supported clade, which was clearly distinct from the clade encompassing all A. salmonicida strains, further highlighting the evolutionary distance between the experimental isolates and related species. Phylogenetic analysis based on the gyrB housekeeping gene revealed distinct clustering patterns among the experimental strains (Fig. 2a). Twelve isolates (XN-1, 2, 3, 4, 5, 6, 8, 9, 10, 11, 12, and 13) formed a well-supported clade with the A. salmonicida standard strain ATCC 33568, exhibiting a high bootstrap value of 0.981. In contrast, the phylogenetic association between this A. salmonicida clade and the A. veronii species cluster received significantly lower support (bootstrap value 0.62), whereas all other relationships with closely related species were weakly supported (bootstrap values < 0.50). The rpoB gene analysis further confirmed these findings (Fig. 2b). All 13 experimental strains clustered robustly with A. salmonicida ATCC 33568, forming a monophyletic group with maximum bootstrap values of 1.00, and demonstrated clear phylogenetic divergence from other closely related species. ANI analysis was performed on the basis of whole-genome sequences to further elucidate the genetic relationships of the strains. As shown in the heatmap (Fig. 3), the OrthoANI values between all 13 experimental strains and the A. salmonicida standard strain ATCC 33658 were >96%, all significantly above the established species delineation threshold of 95.00% (Figueras et al., 2014), thus confirming their conspecificity. In contrast, the OrthoANI values between the 13 experimental strains and closely related species were approximately 85%, well below this species boundary, clearly supporting their distinct species status. In conclusion, our findings demonstrated that the 13 porcine isolates were unequivocally A. salmonicida, as confirmed by a combination of 16S rRNA gene sequencing, MLST, and core-genome phylogenies. Notably, these strains form a phylogenetically distinct cluster within the species, separate from other documented isolates. The successful growth of these porcine-derived strains at 37°C is a key phenotypic characteristic, which contrasts with the typical psychrophilic profile of many classical A. salmonicida strains and may reflect adaptation to a mammalian host. Aeromonas salmonicida multidrug resistance phenotypesThe phenotypic AMR profiles of 13 A. salmonicida strains isolated from porcine hosts are summarized in Figure 4. All tested strains demonstrated complete resistance (13/13, 100.00%) to six antimicrobials: meropenem, sulfamethoxazole, aztreonam, trimethoprim, cefazolin, and ampicillin. Additionally, high resistance rates were observed for amikacin (8/13, 61.54%), gentamicin (8/13, 61.54%), tobramycin (7/13, 53.85%), azithromycin (6/13, 46.15%), ceftazidime (6/13, 46.15%), and tetracycline (6/13, 46.15%). In contrast, all strains (13/13, 100%) were susceptible to fosfomycin and rifampicin. According to the CLSI interpretive criteria (CLSI M45, 2016; CLSI M100, 2023; CLSI M42, 2011), intermediate sensitivity was observed for florfenicol (9/13, 69.23%), suggesting that florfenicol represents a potential therapeutic alternative for infections caused by multidrug-resistant (MDR) strains. For polymyxin B, for which no CLSI "Susceptible" breakpoint is established in M100 (33th ed., 2023), the MIC distribution of the 13 strains was as follows: 11 isolates (84.62%) exhibited MICs of ≤1 µg/ml. According to the available breakpoints, one isolate (7.69%) was classified as intermediate (MIC=2 µg/ml), and one isolate (7.69%) was classified as resistant (MIC=4 µg/ml). The mCIM was performed for phenotypic verification of carbapenemase production to confirm the accuracy of carbapenem resistance. For the 13 experimental strains, the mean inhibition zones from triplicate experiments ranged from 8.74 to 10.48 mm. These results fall within the 6–15 mm range specified by the CLSI M100 (33th ed., 2023) guidelines for a positive carbapenemase test, confirming that all 13 strains are carbapenemase-positive. This indicates that the isolates produced carbapenemases capable of hydrolyzing meropenem from the susceptibility disks, thereby partially inhibiting the growth of the E. coli ATCC 25922 indicator strain. Antibiotic resistance genotypes in A. salmonicidaSecond-generation sequencing analysis was performed on 13 A. salmonicida strains, revealing 30 distinct AMR genes (Fig. 5). The most prevalent resistance genes were cphA5 (13/13, 100.00%), blaOXA-956 (13/13, 100.00%), adeF (12/13, 92.31%), and arnT (12/13, 92.31%). The other frequently detected genes were blaFOX-20 (9/13, 69.23%) and tet(E) (4/13, 30.77%). Gene such as blaFOX-18 was detected at lower frequencies (3/13, 23.08%). No novel resistance genes were identified; however, the combination and high prevalence of specific genes, such as cphA5 and blaOXA-956 are notable. A clear correlation was observed between resistance genotypes and phenotypes, particularly for β-lactam, aminoglycoside, and tetracycline resistance, where the phenotypic resistance patterns were consistently aligned with the presence of corresponding resistance genes. Table 2 shows the specific percentage identity and coverage for each resistance gene.
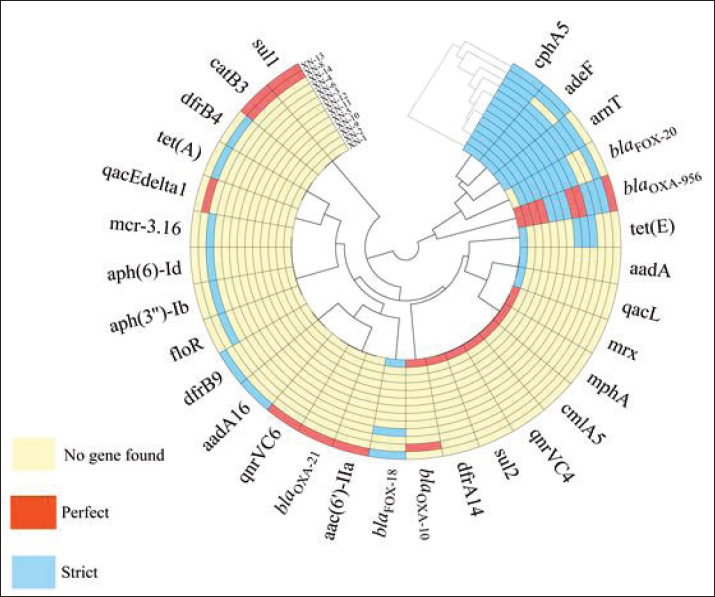
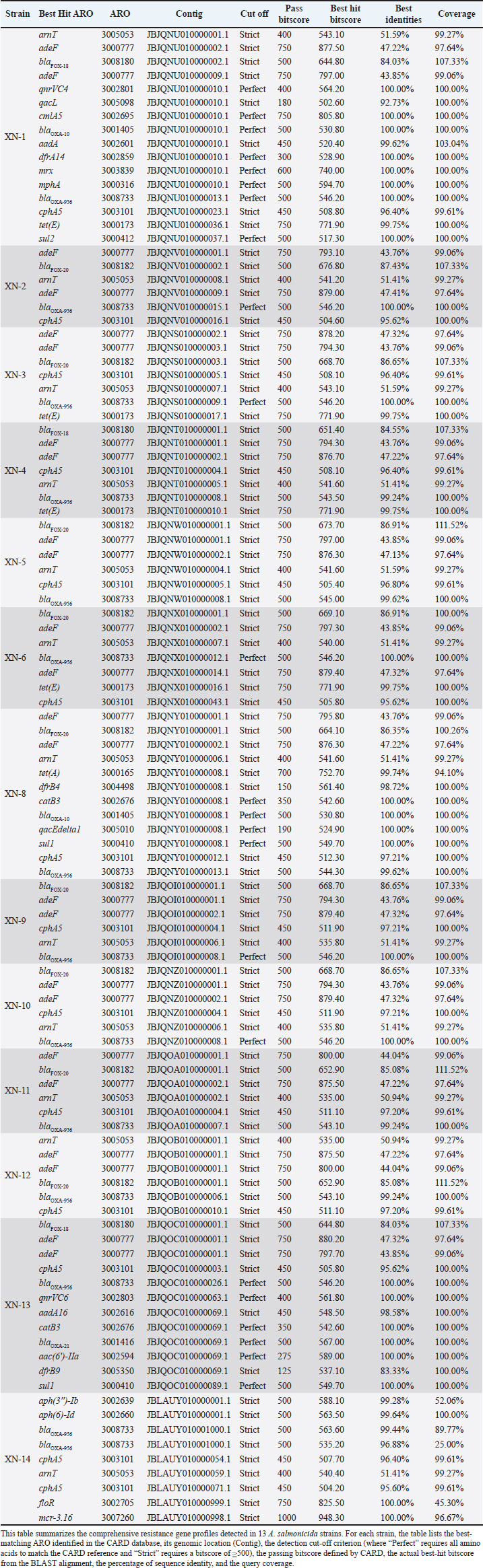
Fig. 5. Resistance genes of A. salmonicida in 13 isolates This figure shows the carriage of drug resistance genes in 13 A. salmonicida strains isolated from pigs in Jilin Province, China. Red, perfect resistance genes; blue, strict resistance genes; yellow, absence of resistance genes. The sequence number in parentheses following the strain number is the accession number of corresponding strain’s second-generation sequencing data. The angular branches were clustered using a circular dendrogram with the group average clustering method, Euclidean distance type, and a single cluster. The clustering along the radius is configured to utilize radial dendrograms, with the group average clustering method, the Euclidean distance type, and a single cluster. Table 2. Resistance gene profiles in 13 A. salmonicida strains detected using the CARD database.
DiscussionOur study’s robust phylogenetic and genomic evidence unequivocally supports the classification of all 13 porcine isolates as A. salmonicida. This conclusion is supported by a conclusive polyphasic approach, which integrated high 16S rRNA sequence identity, ANI values significantly above the species threshold, and definitive assignment via PubMLST analysis. Therefore, the possibility of misidentification as a closely related species can be dismissed confidently. The phylogenetic reconstruction based on the gyrB gene was particularly insightful. As proposed by Yáñez et al. (2003), gyrB serves as an excellent molecular chronometer for phylogenetic studies of Aeromonas and is a useful target for the simultaneous identification of species and strains due to its sequence diversity at the intraspecies level. Our gyrB-based analysis confirmed the specific identity of our isolates while also revealing a distinct clustering that separates them from other referenced A. salmonicida strains of diverse origins. Notably, these isolates formed a phylogenetically distinct cluster within A. salmonicida, suggesting that they may represent a specific sub-lineage. The successful cultivation of all isolates at 37°C is a pivotal and consistent finding that warrants emphasis. This mesophilic trait contrasts with the typical psychrophilic profile (optimal growth at 22°C–28°C) historically associated with A. salmonicida. Thus, our results challenge this rigid phenotypic classification and indicate the emergence of variants capable of thriving at mammalian body temperatures. This temperature tolerance may have implications for the ecological range and pathogenic potential of this species. Although human infections with A. salmonicida remain rare, cases such as the documented endocarditis in a 20-year-old man indicate the potential of A. salmonicida as an opportunistic pathogen in mammals (Salehi et al., 2019). The recovery of porcine strains capable of robust growth at 37°C, coupled with such clinical reports, suggests that certain lineages, possibly like the distinct sub-lineage identified here, may possess adaptive traits that enable warm-blooded host colonization. We hypothesize that the ability to grow at 37°C may be a critical phenotypic marker for these less common, potentially host-adapted variants, which could be underestimated in clinical settings due to the entrenched reputation of A. salmonicida as a cold water organism. Several resistance genes exhibited a detection rate of 15.38% (2/13), including those conferring resistance to phenolic compounds (catB3), β-lactams (blaOXA-10), and sulfonamides (sul1). In contrast, a distinct group of resistance determinants was detected at a lower frequency (1/13, 7.69%): qacEdelta1, floR, qnrVC4, qnrVC6, qacL, cmlA5, dfrA14, mrx, mphA, sul2, aph(3'')-Ib, aph(6)-Id, mcr-3.16, dfrB4, dfrB9, tet(A), aac(6')-IIa, aadA16, and blaOXA-21. The presence of these resistance genes—supports the possibility that MDR Aeromonas spp. from aquatic environments and food production chains could act as potential vectors for AMR dissemination (Lee et al., 2023). Evidence from the well-documented habitat overlap between Aeromonas and human activity, including contamination of water sources, aquaculture systems, and fresh produce, supports this ecological plausibility. Although direct transmission to humans via food or water remains an area requiring further validation, the genetic mobility of these determinants highlights a plausible route for the spread of AMR. Our genomic analysis revealed a concerning AMR profile in porcine-derived A. salmonicida strains. Several high-prevalence genes were clinically relevant, including the carbapenemase gene cphA5 (13/13, 100%) and the extended-spectrum β-lactamase blaOXA-956 (13/13, 100%). Notably, we also identified the colistin resistance gene mcr−3.16 (1/13, 7.69%), a finding of critical public health concern due to its role in conferring resistance to polymyxins, a last-resort class of antibiotics. The identification of the mcr−3.16 gene, which was robustly confirmed by its high-quality match in the CARD database, provides a valuable case to discuss the epidemiological implications of resistance gene localization. The genomic context of such a critical resistance determinant—whether chromosomal or plasmid-borne—fundamentally shapes its dissemination pathway. If on a plasmid, an mcr gene poses a significant threat because of its high mobility and potential for rapid horizontal transfer across diverse bacterial populations. In contrast, a chromosomally encoded mcr gene would exhibit restricted immediate transferability but would likely be stably maintained through vertical inheritance, becoming a persistent trait within a specific bacterial lineage. This distinction is particularly relevant for our A. salmonicida strain, which concomitantly exhibits growth at 37°C. We are not merely observing a transient vessel for a mobile resistance gene but a potential bacterial host that may have stably incorporated colistin resistance as a core genomic characteristic. Although A. salmonicida is a well-documented aquaculture pathogen, its recovery from a swine reservoir carrying antimicrobial resistance underscores its ecological expansion into terrestrial hosts (Vincent et al., 2022). Despite the 2017 ban on colistin in Chinese agriculture, the persistent detection of mcr−3.16 indicates its ongoing circulation. This aligns with previous findings that Aeromonas can act as a reservoir for colistin resistance, potentially facilitating its transfer within ecosystems and amplifying zoonotic risks (Shen et al., 2018). Regardless of its genetic location, the presence of mcr−3.16 in a strain adapted to both aquatic and terrestrial hosts highlights an unrecognized public health risk that warrants enhanced surveillance at the swine–human–environment interface. The genomic landscape of porcine-derived A. salmonicida isolates reveals a sophisticated and high-risk resistance scaffold. The ubiquitous presence of the carbapenemase gene cphA5 (100.00%) serves as a prime example, with phenotypic confirmation of this genotypic prediction. The mCIM demonstrated that all 13 strains are functional carbapenemase producers, with mean inhibition zones (8.74–10.48 mm) definitively falling within the positive range per CLSI M100 guidelines. This perfect genotype-phenotype concordance underscores a mechanism of carbapenem resistance. This core resistance is synergistically augmented by the universal penicillinase blaOXA-956 (100.00%) and prevalent efflux pumps (adeF, arnT; 92.30% each), collectively enabling resistance to β-lactams, aminoglycosides, and fluoroquinolones through enzymatic inactivation, active efflux, and membrane impermeability. This robust scaffold facilitates the accumulation of secondary resistance genes (e.g., tet(E): 30.77%, aadA: 7.69%, and sul1: 15.38%) under multidrug pressure. Beyond the core mechanisms of antibiotic resistance, a concerning link to agricultural practices emerges from the co-occurrence of biocide resistance genes (qacEdelta1: 7.69%) with other resistance determinants. This genetic association suggests that the use of quaternary ammonium compounds in farm sanitization could drive cross-resistance co-selection. The qacEdelta1 gene enhances bacterial survival in sanitized environments, thereby promoting the maintenance and dissemination of MDR reservoirs. This phenomenon highlights the urgent need to re-evaluate biocide usage policies in animal production to mitigate the unintended amplification of AMR threats. Despite these findings, critical knowledge gaps persist. The absence of farm-specific antibiotic usage data is a key limitation of this study, which precludes direct correlation between observed resistance profiles and local antimicrobial selection pressures. Furthermore, the genomic analysis in this study relied on second-generation sequencing. Although this method accurately identified the presence of resistance genes, its short-read nature limits the ability to confidently determine whether genes such as mcr−3.16 are located on the chromosome or on plasmids—a critical distinction for assessing their mobility potential and transmission risk. Additionally, although stringent bioinformatic filters were applied, the possibility of low-level contamination or misassembled sequences, which may have influenced the reported gene profiles, cannot be entirely ruled out. Future studies incorporating long-read sequencing and advanced assembly methods will be essential for resolving genetic contexts and verifying genomic integrity. Finally, the phenotypic contribution and regulatory mechanisms of low-frequency resistance genes remain unverified and warrant functional validation through transcriptomic and proteomic approaches to confirm their expression and activity under relevant conditions. Future studies should incorporate detailed documentation of on-farm antibiotic and disinfectant usage alongside genomic surveillance to address these gaps. Additionally, the experimental validation of resistance genes via gene knockout/complementation assays and dose-response profiling to biocides would help clarify their role in resistance development. Finally, integrating metagenomic profiling of swine and farm environmental microbiomes with in vitro or in vivo models of colonization and transmission would significantly enhance our understanding of zoonotic AMR dissemination pathways and potential virulence trade-offs in A. salmonicida and other emerging pathogens. This study revealed MDR in porcine A. salmonicida isolates, demonstrating complete resistance (13/13, 100%) to β-lactams, sulfamethoxazole, and trimethoprim. Notably, universal resistance was observed against β-lactams (including carbapenems) and sulfonamides—agents commonly employed as empirical first-line therapeutics for swine respiratory disease. These findings demonstrate that the empirical use of these antimicrobial classes is ineffective against A. salmonicida infections and underscore the urgent need to revise antimicrobial stewardship policies and clinical treatment guidelines in swine production systems. Genomic analysis further identified a diverse repertoire of 30 resistance genes that strongly correlated with the observed phenotypic resistance. The transition of MDR A. salmonicida from its conventional aquatic ecological niche into terrestrial livestock production systems is of particular concern, demonstrate successful crosshost adaptation. This shift poses substantial zoonotic risks, potentially enabling transmission to humans through direct animal contact, consumption of contaminated food products, or environmental exposure. Consequently, these findings highlight the urgent necessity of implementing an integrated One Health surveillance framework that encompasses human, animal, and environmental sectors to monitor dissemination pathways and mitigate the further spread of AMR. Moreover, the co-occurrence of biocide and antimicrobial resistance genes implies that disinfectant use in farming environments may coselect for multidrug-resistant strains, further complicating infection control efforts. This reinforces the importance of adopting a holistic approach in managing AMR that incorporates reasonable antibiotic use, targeted biocide application, and continuous genomic surveillance. AcknowledgmentsWe extend our appreciation to the Jilin Province Science and Technology Innovation Platform Subsidy Project and the JiLin Agricultural Science and Technology College staff. Conflict of interestThe authors declare that they have no conflicts of interest to the work reported in this paper. FundingThe Jilin Province Science and Technology Innovation Platform Subsidy Project (Grant number YDZJ202302CXJD038) funded this research. Authors' contributionsNuo Xu: Writing the original draft. Hannuo Li: Visualization. Xinyu Wang: Data analysis. Hui Su: Methodology. Yuhang Zheng: Formal analysis. Wanli Sha: Validation. Baishuang Yin: Supervision. Wenlong Dong: Writing, review, and editing. All authors have reviewed and approved the final version of the manuscript. Data availabilityThe 16S rRNA gene sequences and second-generation sequencing data of all 13 A. salmonicida isolates have been deposited in the NCBI database. The corresponding accession numbers are provided in Table 1. Relevant data supporting the findings are available upon reasonable request from the corresponding author. ReferencesClinical and Laboratory Standards Institute. 2011. Methods for Antimicrobial Disk Susceptibility Testing of Bacteria Isolated From Aquatic Animals. CLSI guideline M42. Clinical and Laboratory Standards Institute. 2016. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria, 3rd ed. CLSI guideline M45. Clinical and Laboratory Standards Institute. 2023. Performance Standards for Antimicrobial Susceptibility Testing, 33rd ed. CLSI guideline M100. Esteve, C., Alcaide, E. and Blasco, M. D. 2012. Aeromonas hydrophila subsp. dhakensis isolated from feces, water and fish in Mediterranean Spain. Microbes Environ. 27(4), 367–373. Figueras, M. J., Beaz-Hidalgo, R., Hossain, M. J. and Liles, M. R. 2014. Taxonomic affiliation of new genomes should be verified using average nucleotide identity and multilocus phylogenetic analysis. Genome Announc. 2(6), e00927-14. Haibin, X., Xiang, L., Jun, Q., Xiaohui, P., Jingyuan, S. and Guangrui, Q., et al. 2012. Fastuniq: a fast de novo duplicates removal tool for paired short reads. Plos One. 7(12), e52249. Küpfer, M., Kuhnert, P., Korczak, B. M., Peduzzi, R. and Demarta, A. 2006. Genetic relationships of Aeromonas strains inferred from 16S rRNA, gyrB and rpoB gene sequences. Int J Syst Evol Micr. 56(12), 2743–2751. Lee, H. J., Storesund, J. E., Lunestad, B. T., Hoel, S., Lerfall, J. and Jakobsen, A. N. 2023. Whole genome sequence analysis of Aeromonas spp. isolated from ready-to-eat seafood: antimicrobial resistance and virulence factors. Front Microbiol. 14, 1175304. Long, M., Fan, H., Gan, Z., Jiang, Z., Tang, S., Xia, H. and Lu, Y. 2023. Comparative genomic analysis provides insights into taxonomy and temperature adaption of Aeromonas salmonicida. J Fish Dis. 46(5), 545-561. Moreno, C., Romero, J. and Espejo, R. T. 2002. Polymorphism in repeated 16S rRNA genes is a common property of type strains and environmental isolates of the genus Vibrio. Microbiology(Reading). 148(4), 1233-1239. Pavan, M. E., Abbott, S. L., Zorzópulos, J. and Janda, J. M. 2000. Aeromonas salmonicida subsp. pectinolytica subsp. nov., a new pectinase-positive subspecies isolated from a heavily polluted river. Int J Syst Evol Microbiol. 50(3), 1119-1124. Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A. and Korobeynikov, A. 2020. Using SPAdes De Novo Assembler. Curr. Protoc. Bioinform. 70(1), e102. Salehi, M., Shadvar, S., Sadeghian, M., Doomanlou, M., Abdollahi, A., Manshadi, S. A. D., et al. 2019. Endocarditis with Aeromonas salmonicida. IDCases 18:e00625. Shen, Y., Xu, C., Sun, Q., Schwarz, S., Ou, Y., Yang, L., Huang, Z., Eichhorn, I., Walsh, T. R., Wang, Y., Zhang, R. and Shen, J. 2018. Prevalence and Genetic Analysis of mcr-3-Positive Aeromonas Species from Humans, Retail Meat, and Environmental Water Samples. Antimicrob agents ch. 62(9), e00404-18. Tewari, R., Dudeja, M., Nandy, S. and Das, A. K. 2014. Isolation of Aeromonas salmonicida from human blood sample: a case report. J Clin Diagn Res. 8(2), 139-140. Vincent, A. T. and Charette, S. J. 2022. To be or not to be mesophilic, that is the question for Aeromonas salmonicida. Microorganisms. 10(2), 240. Vincent, A. T., Fernández-Bravo, A., Sanchis, M., Mayayo, E., Figueras, M. J. and Charette, S. J. 2019. Investigation of the virulence and genomics of Aeromonas salmonicida strains isolated from human patients. Infect Genet Evol. 68, 1-9. Yañez, M. A., Catalán, V., Apráiz, D., Figueras, M.J. & Martínez-Murcia, A.J. 2003. Phylogenetic analysis of members of the genus Aeromonas based on GyrB gene sequences. Int J Syst Evol Microbiol. 53, 875-883. Yoon, S. H., Ha, S. M., Lim, J., Kwon, S. and Chun, J. 2017. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek. 110(10), 1281-1286. | ||
| How to Cite this Article |
| Pubmed Style Xu N, Li H, Wang X, Su H, Zheng Y, Sha W, Yin B, Dong W. Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Vet. J.. 2025; 15(12): 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 Web Style Xu N, Li H, Wang X, Su H, Zheng Y, Sha W, Yin B, Dong W. Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. https://www.openveterinaryjournal.com/?mno=276503 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.57 AMA (American Medical Association) Style Xu N, Li H, Wang X, Su H, Zheng Y, Sha W, Yin B, Dong W. Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Vet. J.. 2025; 15(12): 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 Vancouver/ICMJE Style Xu N, Li H, Wang X, Su H, Zheng Y, Sha W, Yin B, Dong W. Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 Harvard Style Xu, N., Li, . H., Wang, . X., Su, . H., Zheng, . Y., Sha, . W., Yin, . B. & Dong, . W. (2025) Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Vet. J., 15 (12), 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 Turabian Style Xu, Nuo, Hannuo Li, Xinyu Wang, Hui Su, Yuhang Zheng, Wanli Sha, Baishuang Yin, and Wenlong Dong. 2025. Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Veterinary Journal, 15 (12), 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 Chicago Style Xu, Nuo, Hannuo Li, Xinyu Wang, Hui Su, Yuhang Zheng, Wanli Sha, Baishuang Yin, and Wenlong Dong. "Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization." Open Veterinary Journal 15 (2025), 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 MLA (The Modern Language Association) Style Xu, Nuo, Hannuo Li, Xinyu Wang, Hui Su, Yuhang Zheng, Wanli Sha, Baishuang Yin, and Wenlong Dong. "Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization." Open Veterinary Journal 15.12 (2025), 6786-6800. Print. doi:10.5455/OVJ.2025.v15.i12.57 APA (American Psychological Association) Style Xu, N., Li, . H., Wang, . X., Su, . H., Zheng, . Y., Sha, . W., Yin, . B. & Dong, . W. (2025) Aeromonas salmonicida isolated from pigs: Genomic and phenotypic antimicrobial resistance characterization. Open Veterinary Journal, 15 (12), 6786-6800. doi:10.5455/OVJ.2025.v15.i12.57 |