| Research Article | ||
Open Vet. J.. 2025; 15(12): 6316-6328 Open Veterinary Journal, (2025), Vol. 15(12): 6316-6328 Research Article Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, IraqMohammed Jasim Mohammed Awed, Ayman Hani Taha* and Omar Hashim SheetDepartment of Veterinary Public Health, College of Veterinary Medicine, University of Mosul, Mosul, Iraq *Corresponding Author: Ayman Hani Taha. Department of Veterinary Public Health, College of Veterinary Medicine, University of Mosul, Mosul, Iraq. Email: dr_vetayman [at] uomosul.edu.iq- Submitted: 10/08/2025 Revised: 01/11/2025 Accepted: 09/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
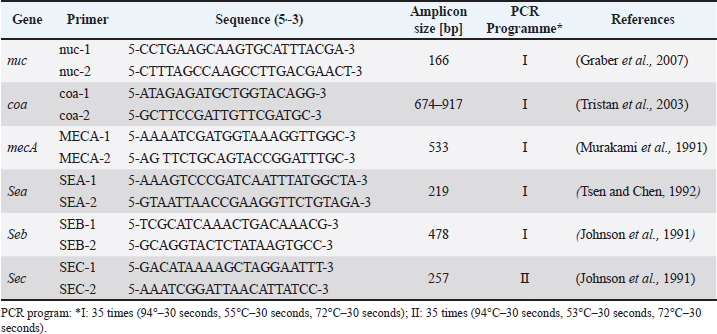
AbstractBackground: The World Health Organization has identified methicillin-resistant Staphylococcus aureus (MRSA) as a "priority bacterium" because of its significant worldwide incidence, resistance to several drugs, and difficult-to-treat infections in both community and clinical settings. Aim: The objectives of this study are to collect and discuss MRSA isolates from restaurants, investigate and contrast the genetic profiles of the MRSA strains found in this study with those found in other regions of the world, and use polymerase chain reaction (PCR) techniques to examine the isolates for the existence of the sea, seb, and sec enterotoxin genes. Methods: Three hundred fifty samples (knives, meat, tables, hands, and machinery) were gathered from 20 restaurants in Erbil city. The period of study was from August to November 2024. All samples were transferred to the Laboratory for examination. Results: The results of the research showed that 44.1% of the S. aureus isolates from restaurants were triggered by MRSA. Meat samples had the highest rate of MRSA (64.5%), subsequent by knife samples (62.1%), machine surfaces (50.0%), and table and hand swab samples (38.7%). Moreover, the results demonstrated that all MRSA isolates (100%) carried the nuc, coa, and mecA genes, as confirmed by PCR analysis. In terms of enterotoxin gene distribution, the sea gene was the most prevalent, detected in 95.5% (64/67) of the MRSA isolates, followed by the sec gene in 65.7%, and the seb gene in 25.4% of the isolates. Profile III—characterized by the presence of nuc, coa, mecA, sea, and sec—was the most predominant, accounting for 55.3%. The phylogenetic analysis revealed that the sequence similarity among the isolates obtained in this study ranged from 98.8% to 99.8%, indicating a high degree of genetic relatedness. The sequences obtained in this study exhibited a close genetic relationship with isolates from Norway and Poland, showing sequence similarities ranging from 98.6% to 99.8%. Conclusion: The prepared meat for eating in restaurants was contaminated with S. aureus. All equipment used in restaurants was contaminated with S. aureus, which contributes to spread it which leading to food poisoning. There is a closed relationship between S. aureus isolated with S. aureus isolated from various regions in worldwide. Keywords: MRSA, Enterotoxins genes, Phylogenetic analysis tree, Restaurants, Food contamination. IntroductionThe bacterium Staphylococcus aureus (S. aureus) is frequently discovered harmlessly on the skin and in the nasal passages and infects more than one-third of humans. In addition to being a harmless colonizer, S. aureus is a pathogenic bacterium linked to a number of illnesses. Staphylococcal food poisoning (SFP), which is brought on by enterotoxins and shows as acute gastrointestinal symptoms as cramps and vomiting, is one known example. Additionally, S. aureus leads potentially fatal diseases like toxic shock syndrome, which is associated with bacterial strains that produce toxins, as well as respiratory infections like pneumonia and sepsis, where bloodstream invasion can have significant global implications (Fisher et al., 2018). Methicillin-resistant Staphylococcus aureus (MRSA) has been identified as an important risk factor for infections linked to hospital environments since its identification in 1962 (HA-MRSA). The increasing worldwide concern of antibiotic resistance is largely caused by MRSA and other multidrug-resistant Enterococcus species (Rossolini et al., 2014). While healthcare-associated (HA) invasive MRSA infections have been moving downward, community-associated MRSA (CA-MRSA) incidences have been rising in the general population. Additionally, HA-MRSA, CA-MRSA, and livestock-associated MRSA strains have been recognized in food items and have been linked to other SFP cases (Sergelidis and Angelidis, 2017). When food with large concentrations of staphylococcal enterotoxins (SE) synthesized by S. aureus is consumed, it can lead to staphylococcal food poisoning (Le Loir et al., 2003). After consuming contaminated food, the most apparent clinical signs of SFP usually show two to eight hours later. Usually starting suddenly, these symptoms include extreme nausea and vomiting that are often described as violent or explosive. Abdominal cramping is also common, characterized by sharp or painful stomach discomfort. In many cases, diarrhea accompanies these symptoms, although its presence can vary. Other possible, though less frequent, symptoms include headache, weakness, and sometimes low-grade fever. In most cases, the condition resolves without medical intervention within 24–48 hours (Balaban and Rasooly, 2000; Murray, 2005) Staphylococcal enterotoxins are a diverse group of at least 23 exotoxins with low molecular weight, exhibiting significant similarities in both structure and function. Based on their ability to cause emesis, SEs are grouped into several classical enterotoxins—Staphylococcal enterotoxin (SE), (SEA, SEB, SEC, SED, SEE)—and a group of more recently discovered enterotoxin-like proteins, which share structural similarities but often lack confirmed emetic activity. These enterotoxins act as superantigens, bypassing normal antigen presentation pathways to hyperactivate T cells, leading to an overwhelming cytokine response and immune dysregulation (Fisher et al., 2018; Benkerroum, 2018). Staphylococcal enterotoxin genes are largely associated with mobile genetic elements, which contribute to the wide variability in the presence and distribution of these genes among S. aureus. Furthermore, the expression of SEs is ordered by multiple, often interconnected pathways, which can be modulated by a range of environmental factors (Fisher et al., 2018). Therefore, MRSA is considered a controversial foodborne pathogen because its capacity to cause SFP is not related to its antimicrobial resistance profile. Because MRSA is not highly prevalent in the food chain, foodborne illnesses related to it are generally not regarded as being particularly serious (Sergelidis and Angelidis, 2017). MRSA has been discovered in a variety of sources, including animals, wildlife, and different food items, according to multiple studies (González et al., 2024). Additionally, MRSA has been frequently discovered in individuals working in environments related to animal husbandry and food manufacturing. The prevalence of MRSA differs by region, species, and hygiene standards across different industry sectors due to its broad prevalence and significance for public health (Kraushaar and Fetsch, 2014; Lozano et al., 2016). MRSA has been isolated from many different kinds of sources, such as butcher shop supplies and minced beef (Sheet et al., 2023), meat products and workers in restaurants (Saber et al., 2025), as well as ready-to-sell meats such as poultry, pork, sausage, and minced meat (Sheet et al., 2025). In addition, MRSA has been recovered from diverse animal sources such as fish (Taha et al., 2024) and milk from bovines (Sheet et al., 2022), goats (Taha et al., 2025), and sheep (Alalaf et al., 2025) suffering from subclinical mastitis. The goals of the present study are threefold: (1) to isolate and identify MRSA from restaurants located in different areas of Erbil city; (2) to detect the sea, seb, and sec enterotoxin genes in S. aureus using polymerase chain reaction (PCR) assays; and (3) to investigate the genetic relationship between the MRSA strains isolated in this study and those reported in other regions worldwide. Materials and MethodsSamples collectionThree hundred fifty samples were gathered from 20 restaurants in Erbil city in the Kurdistan Region of Iraq. The samples comprised seventy each from knives, meat, tables, hands, and machinery. The period of the present study was conducted from August to November 2024. Samples were obtained using sterile swabs and immediately placed into sterile bottles under aseptic conditions. The swabs were immersed in peptone water for pre-enrichment, and they were subsequently incubated for the entire night at 37°C. MRSA isolation and characterizationIn order to examine the properties of MRSA colonies, isolates were initially cultivated utilizing traditional microbiological procedures (Quinn et al., 2002). Further biochemical tests included the catalase reaction, which differentiates Staphylococcus species (catalase-positive) from other Gram-positive cocci, and the coagulase test, which specifically identifies S. aureus by its ability to clot plasma. These combined methods allowed for accurate identification and characterization of MRSA isolates. Isolation of DNAMRSA isolates identified through initial screening were cultured on Mannitol Salt Agar and incubated at 37°C for 18–24 hours to promote robust growth, after which genomic DNA was extracted for further analysis. Genomic DNA was extracted using the DNeasy Blood and Tissue Kit (Addbio, Korea) according to the manufacturer’s instructions for Gram-positive organisms. The concentration of DNA was determined using a NanoDrop spectrophotometer (Jenway, UK). MRSA DNA concentrations ranged between 40 µg/μl and 80 µg/μl, and all samples were preserved at −20°C for subsequent analyses. Reaction of PCRThe detection of MRSA genes—including nuc, coa, mecA, sea, seb, and sec—was carried out using PCR assays. The expected amplicon sizes for each target gene were as follows: nuc—166 bp (Graber et al., 2007), coa—674 bp (Tristan et al., 2003), mecA—533 bp (Murakami et al., 1991), sea—219 bp (Tsen and Chen, 1992), seb—478 bp (Johnson et al., 1991), and sec—257 bp (Johnson et al., 1991) (Table 1). These gene targets were selected for their relevance in confirming species identity, antibiotic resistance, and enterotoxin production. The total volume of the PCR reaction was 25 μl, prepared in 200 μl PCR tubes (Biozym, Germany). The amplified PCR products were subsequently resolved by electrophoresis on a 2% agarose gel (Peqlab, Germany). A 100 bp DNA marker was utilized as a molecular size marker to estimate the fragment lengths of the resulting amplicons. The PCR reaction mixture consisted of 12.5 μl of GoTaq Master Mix (2×) (Addbio, Korea), 4 μl of MRSA genomic DNA, 1 μl of each primer (1 and 2), and 6.5 μl of DNeasy-free water (Promega Corporation, USA), resulting in a total reaction volume of 25 μl. Table 1. Primers and PCR conditions for the detection of MRSA genetic markers.
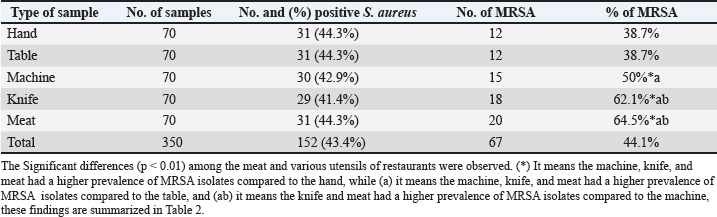
DNA sequencingThe PCR-amplified products corresponding to four target genes, previously confirmed as positive for MRSA by conventional PCR, were submitted to Macrogen Inc. (Seoul, South Korea), a commercial sequencing service provider, for purification and DNA sequencing analysis. Nucleotide sequencing was performed on the seb gene, and the resulting sequences were analyzed using BLASTn (NCBI, http://www.ncbi.nlm.nih.gov) to identify similarities with previously reported MRSA seb gene sequences archived in the GenBank database. For further analysis, multiple sequence alignment was conducted using the CLUSTALW algorithm implemented in the MegAlign software (DNASTAR, USA), allowing for detailed comparison of sequence similarity and variation among the isolates. Phylogenetic analysis was performed using the Neighbor-Joining method implemented via the CLUSTALW tool available at Genome Net. To enhance the robustness and accuracy of the tree topology, 500 duplicate sequences of the seb gene from MRSA isolates were incorporated as an outgroup. This comprehensive approach—including gene purification, sequencing, and subsequent bioinformatic analysis—was employed to elucidate the genetic relationships among MRSA isolates obtained from restaurant environments and to provide insights into their phylogenetic positioning. Statistical analysisStatistical analysis was performed using JMP® 16.1 software (JMP®, 1989-2021). The chi-square test showed a significant variation in the percentage of MRSA isolates across various streets in Erbil City, with a p-value less than 0.05. Ethical approvalThe Institutional Animal Care and Use Committee (IACUC) of the College of Veterinary Medicine, University of Mosul, approved all animal procedures under approval number UM.Vet.2025.007. The sample obtained was approved with informed consent from the animal owners, and all protocols adhered strictly to ethical standards Date: 12/2/2025. ResultsIn the current study, S. aureus colonies were identified by their characteristic golden-yellow pigmentation on Mannitol Salt Agar, indicative of mannitol fermentation. Further validation was achieved through standard biochemical assays, such as catalase and coagulase tests, both of which yielded positive results consistent with S. aureus identification. The concentration of genomic DNA extracted was from 40 µg/μl to 80 µg/μl, as measured by spectrophotometry. The results obtained through classical microbiological methods were further validated by PCR analysis, which confirmed that all S. aureus isolates were positive for the species-specific nuc gene, thereby supporting their molecular identification. According to the Table 2, the overall prevalence of S. aureus isolated from restaurants in Erbil city was 43.4%. The highest isolation rate was observed from hand, table, and meat samples, each accounting for 44.3% of the positive cases. In contrast, the lowest prevalence rates were detected in samples taken from knives (41.4%) and machines (42.9%), as detailed in Table 2. Furthermore, the study revealed that the occurrence of MRSA among S. aureus isolates from restaurant environments in Erbil city was 44.1% (67/152). The highest proportion of MRSA was detected in meat samples, with a prevalence rate of 64.5% (20/31). In comparison, MRSA was isolated from knives at a rate of 62.1% (18/29), machines at 50.0% (15/30), and both table and hand samples at 38.7% (12/31). The Significant differences (p < 0.01) among the meat and various utensils of restaurants were observed. (*) It means the machine, knife, and meat had a higher prevalence of MRSA isolates compared to the hand, while (a) it means the machine, knife, and meat had a higher prevalence of MRSA isolates compared to the table, and (ab) it means the knife and meat had a higher prevalence of MRSA isolates compared to the machine, these findings are summarized in Table 2. Table 2. Prevalence and distribution of Staphylococcus aureus and MRSA isolates from restaurant samples in Erbil City.
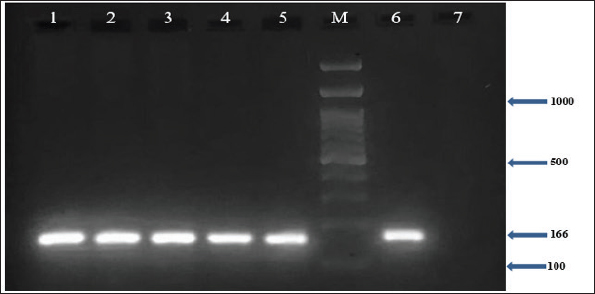
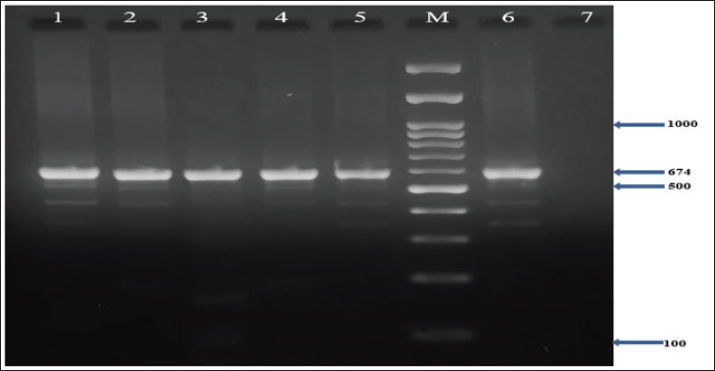
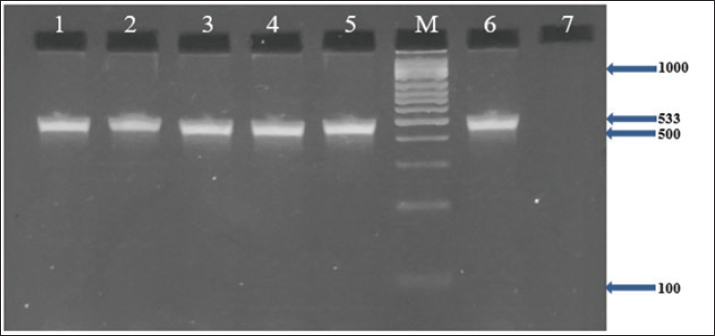
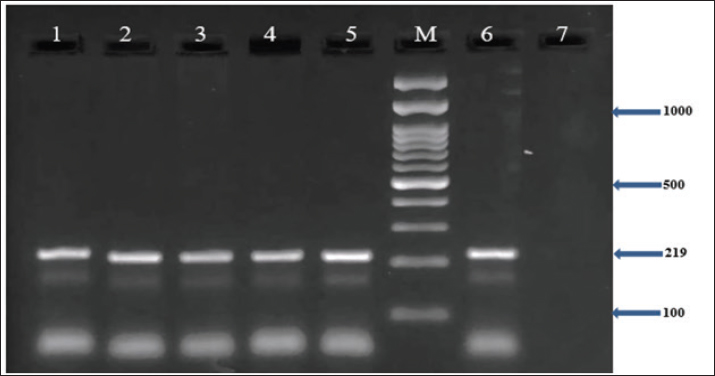
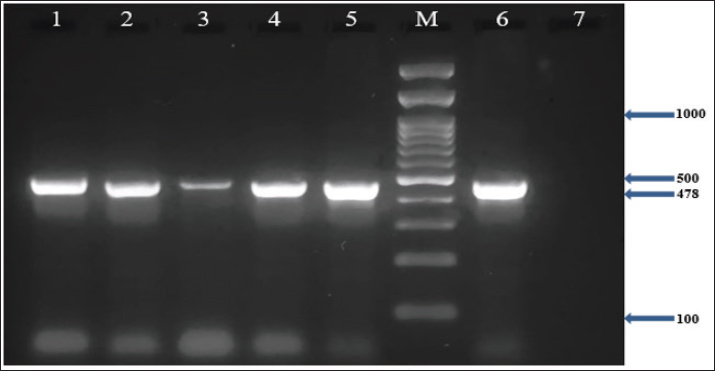
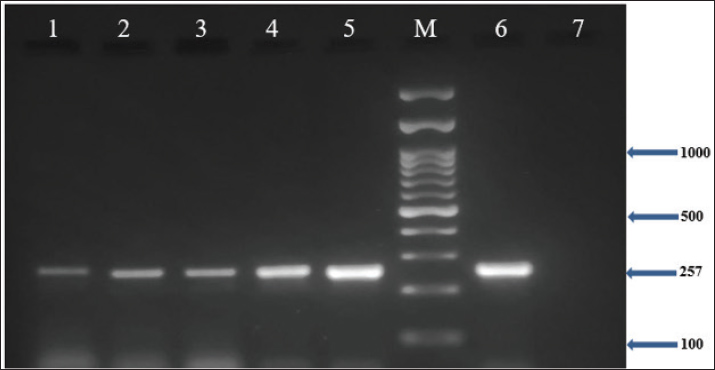
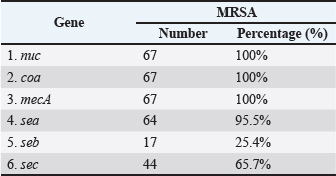
In addition, the results demonstrated that all MRSA isolates (100%) harbored the nuc, coa, and mecA genes, as confirmed by PCR (Figs. 1–3). Furthermore, the incidence of enterotoxin genes among the MRSA isolates was as follows: sea was detected in 95.5% (64/67), seb in 25.4% (17/67), and sec in 65.7% (44/67) of the isolates (Figs. 4–6). A detailed summary of these findings is presented in Table 3. The findings also showed that MRSA isolates exhibited five unique gene profiles (Table 4). The most prevalent profile was Profile III (nuc + coa + mecA + sea + sec), which accounted for 55.3% (37/67) of the total isolates. The most gene-rich profile, Profile I (nuc + coa + mecA + sea + seb + sec), was identified in 10.4% (7/67) of the isolates. Additionally, Profile II (nuc + coa + mecA + sea + seb), Profile IV (nuc + coa + mecA + sea), and Profile V (nuc + coa + mecA) were observed in 10% (10/67), 10% (10/67), and 4.5% (3/67) of the isolates, respectively (Table 4).
Fig. 1. Agarose gel electrophoresis (2%) showing the typical amplicon of the nuc gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is a negative control.
Fig. 2. Agarose gel electrophoresis (2%) showing the typical amplicon of the coa gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is a negative control.
Fig. 3. Agarose gel electrophoresis (2%) showing the typical amplicon of the mecA gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is a negative control.
Fig. 4. Agarose gel electrophoresis (2%) showing the typical amplicon of the sea gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is negative control.
Fig. 5. Agarose gel electrophoresis (2%) showing the typical amplicon of the seb gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is negative control.
Fig. 6. Agarose gel electrophoresis (2%) showing the typical amplicon of the sec gene product of MRSA isolates. The amplification of DNA appears as a ladder-like pattern. Lanes 1–5 represent positive isolates, Lane M is a DNA Marker 100 bp ladder (Biozym Diagnostic), Lane 6 is a positive control, and Lane 7 is negative control. Table 3. Frequency and prevalence of target genes identified in MRSA isolates.
Table 4. Genotypic profiles of MRSA strains isolated from the restaurant environment.
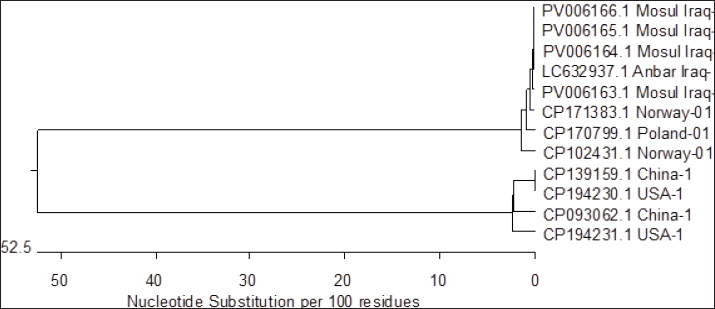
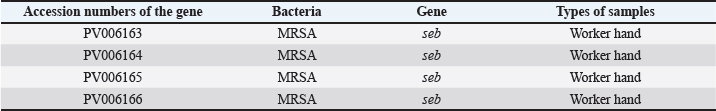
Individual sequence analysis using the BLASTn tool was conducted on four novel seb gene sequences obtained from MRSA isolated from a Worker hand in restaurants. These sequences were aligned with known sequences in the NCBI database to verify their identity and homology. As summarized in Table 5, the newly obtained sequences were deposited in the NCBI GenBank under the accession numbers PV006163, PV006164, PV006165, and PV006166. Furthermore, phylogenetic analysis using the CLUSTALW algorithm implemented in the MegAlign program demonstrated notable genetic divergence between the locally obtained gene sequences and reference sequences previously deposited in the GenBank database. As illustrated in Figure 7, the sequence similarity among the isolates from this study (accession numbers PV006163, PV006164, PV006165, and PV006166) ranged from 98.8% to 99.8%, as also detailed in Table 6. Furthermore, this study demonstrated a genetic relationship between the MRSA sequence isolated from Anbar city, Iraq (GenBank accession number LC632937.1), and the sequences obtained in this study (PV006163, PV006164, PV006165, and PV006166), showing sequence similarities of 100%, 96.4%, 98.0%, and 97.8%, respectively. This study demonstrated a close genetic relationship between the sequences PV006163 and PV006165 of this study with sequences with accession numbers CP102431.1 and CP171383.1 from Norway, as well as CP170799.1 from Poland, with all showing 100% sequence identity. The sequence PV006164 exhibited a high degree of genetic similarity to previously reported sequences CP102431.1 and CP171383.1 from Norway, and CP170799.1 from Poland, with a sequence identity of 98.6%. In addition, the sequence PV006166 showed a close genetic relationship to previously reported sequences CP102431.1 and CP171383.1 from Norway, and CP170799.1 from Poland, with a sequence identity of 99.8%.
Fig. 7. Phylogenetic tree showing clustering analysis of MRSA gene sequences retrieved from NCBI GenBank. Accession numbers are indicated in parentheses next to each sequence, representing their genetic relatedness based on multiple sequence alignment. Table 5. The NCBI GenBank accession numbers for the seb gene of Staphylococcus aureus sequences in the worker hand.
Table 6. Sequence identity and variation between MRSA isolates based on DNASTAR pairwise analysis.
DiscussionThe continued occurrence of foodborne disease outbreaks worldwide—driven by pathogenic bacteria and their associated toxins—underscores the essential role of food quality and safety in safeguarding public health (Chin et al., 2000). These pathogenic bacteria impose a substantial burden on community hygiene, causing widespread illness and mortality across all population groups, with over 600 million people affected globally each year by foodborne diseases (Faour and Todd 2020; Lamas et al., 2023). Furthermore, MRSA strains are capable of producing enterotoxins that can lead to staphylococcal food poisoning, particularly when conditions within the food process support bacterial proliferation and toxin production. This study found that the prevalence of MRSA in restaurants was 44.1%, which is higher than previously reported rates such as the prevalence of MRSA was 37% in Salvador, Brazil (Carvalho et al., 2020), 28.75% in Egypt (Ali and Abd-El-Aziz, 2011), and 25% in Benha City, Egypt (Saif et al., 2019). In contrast, the occurrence of MRSA was lower than that reported in previous studies, where MRSA was isolated from restaurants at a rate of 69.74% in Iran (Ranjbar et al., 2017) and 51.6% in Egypt (Ghanem et al., 2025). This variation may be attributed to several factors, including such as the health of food handlers, hygienic habits, educational attainment, the nation's socioeconomic development, and the implementation of food safety laws and inspections (Sezer et al., 2015). Furthermore, cross-contamination throughout the food production process from outside sources and kitchen personnel (Al-Amery et al., 2019). The results of this investigation revealed that the occurrence of MRSA in meat samples was 64.5%, which is significantly greater than the prevalence rates obtained from previous studies that found that the prevalence was only 0.6% in Northern Greece (Komodromos et al., 2022), 3.2% in China (Ou et al., 2017), and 16.9% in Algeria (Chaalal et al., 2018), and in Egypt was 28.75% (Fathi et al., 2011). The detected variation in incidence of MRSA could be explained by a number of variables, such as cross-contamination during animal transportation, extended contact between infected and non-infected animals at slaughterhouses, inadequate evisceration that results in contamination of feces, and poor hygiene throughout post-slaughter processing and storage (Marriott and Gravani, 2008). Food handlers at restaurants have an important role in preventing hand-to-meat cross-contamination throughout food preparation. The absence of glove use, poor hand hygiene, and wearing dirty or contaminated clothing are all contributing variables that may significantly increase the risk for microbial contamination (Yenealem et al., 2020). When meat that was undercooked spoils or when bacteria cross-contaminate ready-to-eat meals, the risk of infection increases dramatically (Wang et al., 2017). Furthermore, the current study's findings indicated that MRSA was identified on blades in 62.1% of cases, on machines in 50% of cases, and on both tables and hands in 38.7% of cases. In order to prevent contamination and control infection, screening and mitigation measures must be effective, and this involves a knowledge of the sources and transmission dynamics of MRSA (Chon et al., 2017). Food handlers’ hands play an important role in the transmission of MRSA to food. Previous studies reported a prevalence of 4.2% for MRSA isolated from slaughterhouse workers in Greece (Drougka et al., 2019), whereas no MRSA was detected on workers’ hands in Northern Greece (Komodromos et al., 2022). Workers in restaurants are largely responsible for the cross-contamination of meat and hands throughout handling. High bacterial contamination can be significantly exacerbated by behaviors like not wearing gloves, washing your hands frequently, and dressing in filthy clothing (Yenealem et al., 2020). However, by adequately cleaning and sanitizing instruments and equipment before and after they are utilized for handling meat, restaurants can lower the amount of harmful bacteria (Adesokan and Raji, 2014). Furthermore, MRSA and other pathogenic bacteria can infect restaurant equipment, such as cutting boards, knives, grinders, and counters, while meat is being prepared and cooked. Direct contact with contaminated equipment or incorrect handling and poor washing procedures might result in this contamination. All MRSA isolates (100%) were found to possess the nuc, coa, and mecA genes. This finding is consistent with a previous study from Egypt, which also reported universal detection of the mecA gene among MRSA isolates (Abolghait et al., 2020). Early studies conducted in Brazil and China reported that all MRSA isolates recovered from ready-to-eat foods harbored the mecA gene (Wu et al., 2019; Carvalho et al., 2020). Similarly, all MRSA isolates from fresh meat processing facilities in Northern Greece were found to carry the mecA gene (Komodromos et al., 2022). In Iraq, 100% of MRSA isolates from meat, employee hands, and equipment carried both the nuc and mecA genes (Sheet et al., 2023). Additionally, it became apparent that every MRSA isolate from fish meat, employee hands, and equipment included both the nuc and mecA genes (Taha et al., 2024). Likewise, the nuc and mecA genes were present in 100% of the MRSA isolates that have been obtained from Basturma (Sheet et al., 2024). Furthermore, the mecA gene has been determined to be present in all S. aureus strains correlated with goat subclinical mastitis (Taha et al., 2025). These results illustrate a variety of S. aureus strains' genetic makeup, which can be influenced by host species, geographic origin, or particular environmental and management circumstances. There is significant evidence that MRSA can transmit from animals to people, and tainted food items may be a major contributing factor (Oniciuc et al., 2017). Furthermore, among the MRSA isolates in this investigation, the incidence of enterotoxin genes was 65.7% for sec, 25.4% for seb, and 95.5% for sea. The previous research reported that 75% of the MRSA isolates carried the seb gene, while none were found to possess the sea or sec genes (Abolghait et al., 2020). The sea, seb, and sec genes have been discovered to be present in MRSA isolates from retail food samples in China at occurrence rates of 63.89%, 50.93%, and 75%, respectively (Wu et al., 2019). The sea and sec genes have been shown to be 72.6% and 57.53% prevalent, respectively, in MRSA isolates from seafood in Iran (Derke et al., 2025). Due to variances in strain variety, origin, and geographic location, there has been significant variance in the incidence of SE genes among MRSA isolates among studies. Environmental influences, genetic heterogeneity among strains, and inconsistent testing methods are other contributing factors. The findings also revealed that MRSA isolates carried one or more SE genes, highlighting the genetic diversity and varied virulence potential of the strains examined. Differences in genomic architecture, horizontal gene transfer through mobile genetic elements, and regulatory networks controlling toxin gene expression are possibly the primary causes of this variance in enterotoxin gene presence and incidence (Schelin et al., 2011). The clonal lineage of MRSA is intimately associated with the occurrence of its enterotoxin genes, with specific genetic backgrounds often exhibiting distinctive toxin gene profiles. This lineage-specific association plays a significant role in shaping regional and host-specific prevalence trends (Dicks et al., 2021). Four new sequences have been identified in this analysis (accession numbers PV006163–PV006166), and they showed a high degree of sequence similarity (98.6%–100%) to S. aureus entries that had already been submitted to GenBank from across the world. The significant degree of sequence similarity suggests that the newest discovered sequences are either universally circulating or evolutionarily conserved staphylococcal gene variations, indicating widespread transmission within MRSA populations or shared ancestral origins. This high level of sequence correspondence highlights the significance of the obtained sequences in clarifying the genetic variability and global distribution of MRSA virulence factors while also verifying the legitimacy of the found sequences (Jans et al., 2022). ConclusionThis study evaluated the sanitation practices of restaurants in Erbil, Iraq, covering an extensive variety of geographic areas. MRSA's isolation from meat samples increases the possibility that contamination occurred throughout a number of processes, such as handling, transit, storage, or slaughter, most probably in unsanitary settings. Additionally, the finding of MRSA on restaurant utensils emphasizes the possibility that tainted tools and equipment play an important part in the propagation and spread of MRSA, which can contaminate meat and other foods. Furthermore, the presence of MRSA on restaurant utensils and equipment suggests either ongoing reuse without proper sanitation or inappropriate procedures for washing, cleaning, and sterilization. These errors could result in meat being tainted with MRSA, which would be extremely hazardous to the customers' health. Furthermore, many restaurants might use unpackaged meat or improperly store it at non-refrigerated temperatures, in violation of standards for sanitation. These conditions could facilitate MRSA development and the process of multiplication, raising the risk of enterotoxin production and, eventually, food-related consumer illnesses. The MRSA strains gathered in this study were found to be genetically varied and contained a range of sequence types, as evidenced by the identification of numerous virulence-associated genes in the isolates. Furthermore, the genetic similarities between the MRSA isolates from this investigation and those obtained in other nations raise the possibility that they are globally spread or share evolutionary roots. AcknowledgmentOur heartfelt thanks go out to the University of Mosul's College of Veterinary Medicine for their helpfulness and for providing the facilities and materials required for performing this study. Conflict of interestThe manuscript's author verifies that there were no conflicts of interest associated with the study's publishing, data analysis, or authorship. FundingThe research is fully self-funded by the authors. Authors’ contributionOmar HS conceived the idea for this project. Ayman HT and Mohammed JM performed DNA isolation and PCR reactions. All authors contributed to writing the initial draft and collaboratively revised and organized the final version of the research. Data availabilityThe data generated in this study are included in the article. ReferencesAbolghait, S.K., Fathi, A.G., Youssef, F.M. and Algammal, A.M. 2020. Methicillin-resistant Staphylococcus aureus (MRSA) isolated from chicken meat and giblets often produces staphylococcal enterotoxin B (SEB) in non-refrigerated raw chicken livers. Int. J. Food. Microbiol. 328, 108669; doi:10.1016/j.ijfoodmicro.2020.108669 Adesokan, H.K. and Raji, A.O. 2014. Safe meat-handling knowledge, attitudes and practices of private and government meat processing plants' workers: implications for future policy. J. Prev. Med. Hygiene. 55(1), 10; https://pmc.ncbi.nlm.nih.gov/articles/PMC4718335/ Alalaf, A.S., Taha, A.H. and Sheet, O.H. 2025. Genotypic characteristics and phylogenetic tree analysis of Staphylococcus aureus isolates from sheep subclinical mastitis milk. Open Vet. J. 15(1), 289; doi: 10.5455/OVJ.2025.v15.i1.27 Al-Amery, K., Elhariri, M., Elsayed, A., El-Moghazy, G., Elhelw, R., El-Mahallawy, H., El Hariri, M. and Hamza, D. 2019. Vancomycin-resistant Staphylococcus aureus isolated from camel meat and slaughterhouse workers in Egypt. Antimicrob. Resist. &. InfectControl. 8(1), 129; doi:10.1186/s13756-019-0585-4 Ali, S.F.H. and Abd-El-Aziz, D.M. 2011. Incidence of enterotoxigenic Staphylococcus aureus in some ready-to-eat meat sandwiches in Assuit city with special reference to methicillin resistant Staphylococcus aureus strains. Assiut. Vet. Med. J. 57(129), 95–106; doi:10.21608/avmj.2011.173884 Balaban, N. and Rasooly, A. 2000. Staphylococcal enterotoxins. Int. J. Food Microbiol. 61(1), 1–10; doi:10.1016/S0168-1605(00)00377-9 Benkerroum, N. 2018. Staphylococcal enterotoxins and enterotoxin-like toxins with special reference to dairy products: an overview. Crit. Rev. Food Sci. Nutr. 58(12), 1943–1970; doi:10.1080/10408398.2017.1289149 Carvalho, J.S., Neto, A.F.L., Melo, I.M., Varjão, L.M., Andrade, C.A.D.N., Xavier, D.E., Leal, N.C. and De Castro Almeida, R.C. 2020. Occurrence of methicillin-resistant Staphylococcus aureus in ready-to-eat raw fish from Japanese cuisine restaurants in Salvador, Brazil. J. Food Prot. 83(6), 991–995; doi:10.4315/0362-028X.JFP-19-375 Chaalal, W., Chaalal, N., Bourafa, N., Kihal, M., Diene, S.M. and Rolain, J.M. 2018. Characterization of Staphylococcus aureus isolated from food products in Western Algeria. Foodborne Pathogens Dis. 15(6), 353–360; doi:10.1089/fpd.2017.23 Chin, J. 2000. Control of communicable diseases manual. https://www. mtpinnacle.com/pdfs/ccdm.pdf Chon, J., Sung, K. and Khan, S. 2017. Methicillin-resistant Staphylococcus aureus (MRSA) in food-producing and companion animals and food products. Frontiers in Staphylococcus aureus. doi:10.5772/66645 Derke, R.E., Rahimi, E., Shakerian, A. and Khamesipour, F. 2025. Prevalence, virulence factors, and antibiotic resistance of Staphylococcus aureus in seafood products. BMC. Infect. Dis. 25(1), 554; doi:10.1186/s12879-025-10870-1 Dicks, J., Turnbull, J.D., Russell, J., Parkhill, J. and Alexander, S. 2021. Genome sequencing of a historic Staphylococcus aureus collection reveals new enterotoxin genes and sheds light on the evolution and genomic organization of this key virulence gene family. J. Bacteriology 203(10), 10–1128; doi:10.1128/jb.00587-20 Drougka, E., Foka, A., Giormezis, N., Sergelidis, D., Militsopoulou, M., Jelastopulu, E., Komodromos, D., Sarrou, S., Anastassiou, E.D., Petinaki, E. and Spiliopoulou, I. 2019. Multidrug-resistant enterotoxigenic Staphylococcus aureus lineages isolated from animals, their carcasses, the personnel, and the environment of an abattoir in Greece. J. Food. Process. Preservation. 43(7), e13961; doi:10.1111/jfpp.13961 Faour-Klingbeil, D. and Todd, E. 2020. Prevention and control of foodborne diseases in Middle-East North African countries: review of national control systems. Int. J. Environ. Res. Public Health 17(1), 70; doi:10.3390/ijerph17010070 Fathi Hassan Ali, S.O.H.A.I.L.A. and Mohammed Abd-el-aziz, D.O.A.A. 2011. Incidence of enterotoxigenic staphylococcus aureus in some ready-to-eat meat sandwiches in Assuit city with special reference to methicillin resistant staphylococuus aureus strains. Assiut. Vet. Med. J. 57(129), 1–12; doi:10.21608/avmj.2011.173884 Fisher, E.L., Otto, M. and Cheung, G.Y.C. 2018. Basis of virulence in enterotoxin-mediated staphylococcal food poisoning. Front. Microbiol. 9, 436; doi:10.3389/fmicb.2018.00436 Ghanem, M.A., Idris, A.B.M., Abu El Roos, N. and Darwish, W.S. 2025. Prevalence of multidrug-resistant Staphylococcus aureus in meals served at hospitals. Egypt. J. Vet. Sci. 56(7), 1479–1488; doi:10.21608/EJVS.2024.282625.2001 González-Machado, C., Alonso-Calleja, C. and Capita, R. 2024. Methicillin-Resistant Staphylococcus aureus (MRSA) in different food groups and drinking water. Foods 13(17), 2686; doi:10.3390/foods13172686 Graber, H.U., Casey, M.G., Naskova, J., Steiner, A. and Schaeren, W. 2007. Development of a highly sensitive and specific assay to detect Staphylococcus aureus in bovine mastitic milk. J. Dairy. Sci. 90(10), 4661–4669; doi:10.3168/jds.2006-902 Jans, C., Wambui, J., Stevens, M.J.A. and Tasara, T. 2022. Comparative genomics of dairy-associated Staphylococcus aureus from selected sub-Saharan African regions reveals milk as reservoir for human-and animal-derived strains and identifies a putative animal-related clade with presumptive novel siderophore. Front. Microbiol. 13, 923080; doi:10.3389/fmicb.2022.923080 JMP®, Pro 16.1. 1989. SAS Institute Inc., Cary, NC, 1989–2021. Johnson, W.M., Tyler, S.D., Ewan, E.P., Ashton, F.E., Pollard, D.R. and Rozee, K.R. 1991. Detection of genes for enterotoxins, exfoliative toxins, and toxic shock syndrome toxin 1 in Staphylococcus aureus by the polymerase chain reaction. J. Clin. Microbiol. 29(3), 426–430; doi:10.1128/jcm.29.3.426-430.1991 Komodromos, D., Kotzamanidis, C., Giantzi, V., Pappa, S., Papa, A., Zdragas, A., Angelidis, A. and Sergelidis, D. 2022. Prevalence, infectious characteristics and genetic diversity of Staphylococcus aureus and methicillin-resistant Staphylococcus aureus (MRSA) in two raw-meat processing establishments in Northern Greece. Pathogens 11(11), 1370; doi:10.3390/pathogens11111370 Kraushaar, B. and Fetsch, A. 2014. First description of PVL-positive methicillin-resistant Staphylococcus aureus (MRSA) in wild boar meat. Int. J. Food. Microbiol. 186, 68–73; doi:10.1016/j.ijfoodmicro.2014.06.018 Lamas, A., Franco, C.M. and Regal, P., 2023. Foodborne Pathogens-Recent Advances in Control and Detection. London: Intechopen. doi:10.5772/intechopen.98159 Le Loir, Y., Baron, F. and Gautier, M., 2003. Staphylococcus aureus and Food Poisoning. Laboratoire de Microbiologie. Ecole Nationale Superieure Agronomique de Rennes, Institut Nationale de la Recherche Agronomique, Fransce Available via https://hal.science/hal-01123026/ Lozano, C., Gharsa, H., Ben Slama, K., Zarazaga, M. and Torres, C. 2016. Staphylococcus aureus in animals and food: methicillin resistance, prevalence and population structure. A review in the African continent. Microorganisms 4(1), 12; doi:10.3390/microorganisms4010012 Marriott, N.G. and Gravani, R.B. 2008. Principles of food sanitation. J. Environ. Health. 71(5), 42–43; doi:10.1007/978-3-319-67166-6_21 Murakami, K., Minamide, W., Wada, K., Nakamura, E., Teraoka, H. and Watanabe, S. 1991. Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J. Clin. Microbiol. 29(10), 2240–2244; doi:10.1128/jcm.29.10.2240-2244.1991 Murray, R.J. 2005. Recognition and management of Staphylococcus aureus toxin-mediated disease. Internal Med. J. 35, S106–S119; doi:10.1111/j.1444-0903.2005.00984.x Oniciuc, E.A., Nicolau, A.I., Hernández, M. and Rodríguez-Lázaro, D. 2017. Presence of methicillin-resistant Staphylococcus aureus in the food chain. Trends. Food. Sci. &. Technol. 61, 49–59; doi:10.1016/j.tifs.2016.12.002 Ou, Q., Peng, Y., Lin, D., Bai, C., Zhang, T., Lin, J., Ye, X. and Yao, Z. 2017. A meta-analysis of the global prevalence rates of Staphylococcus aureus and methicillin-resistant S. aureus contamination of different raw meat products. J. Food. Prot. 80(5), 763–774; doi:10.4315/0362-028X.JFP-16-355 Quinn, P. J., Bryan K. Markey, and M. E. Carter. "Veterinary microbiology and microbial disease." (2002): 113-118.Available: https://sid. ir/paper/657266/en Ranjbar, R., Shahreza, M.H.S., Rahimi, E. and Jonaidi-Jafari, N. 2017. Methicillin-resistant Staphylococcus aureus isolates from Iranian restaurant food samples: panton-Valentine Leukocidin, SCCmec phenotypes and antimicrobial resistance. Trop. J. Pharm. Res. 16(8), 1939–1949; doi:10.4314/tjpr.v16i8.26 Rossolini, G.M., Arena, F., Pecile, P. and Pollini, S. 2014. Update on the antibiotic resistance crisis. Curr. Opinion. Pharmacol. 18, 56–60; doi:10.1016/j.coph.2014.09.006 Saber, A.S. and Sheet, O.H. 2025. Molecular detection of the genes encoding virulence factors of coagulase-positive Staphylococcus aureus isolated from restaurants. Iraqi J. Vet. Sci. 39(2), 217–224; doi:10.33899/ijvs.2025.155576.4042 Saif, M., Saad, S., Hassanin, F., Shltout, F. and Zaghloul, M. 2019. Prevalence of methicillin-resistant Staphylococcus aureus in some ready-to-eat meat products. Benha Vet. Med. J. 37(1), 12–15; doi:10.21608/bvmj.2019.15641.1063 Schelin, J., Wallin-Carlquist, N., Thorup Cohn, M., Lindqvist, R. and Barker, G.C. 2011. The formation of Staphylococcus aureus enterotoxin in food environments and advances in risk assessment. Virulence 2(6), 580–592; doi:10.4161/viru.2.6.18122 Sergelidis, D. and Angelidis, A.S. 2017. Methicillin-resistant Staphylococcus aureus: a controversial food-borne pathogen. Lett. Appl. Microbiol. 64(6), 409–418; doi:10.1111/lam.12735 Sezer, C., Özgür, C., Aksem, A. and Leyla, V. 2015. Food handlers: a bridge in the journey of enterotoxigenic MRSA in food. J. Für. Verbraucherschutz. Und. Lebensmittelsicherheit. 10(2), 123–129; doi:10.1007/s00003-015-0939-7 Sheet, O.H. 2022. Molecular detection of mecA gene in methicillin-resistant Staphylococcus aureus isolated from dairy mastitis in Nineveh governorate, Iraq. Iraqi. J. Vet. Sci. 36(4), 939–943; doi:10.33899/ijvs.2022.132643.2115 Sheet, O.H., Al-Mahmood, O.A., Othamn, S.M., Al-Sanjary, R.A., Alsabawi, A.H. and Abdulhak, A.A. 2023. Detection of positive meca Staphylococcus aureus isolated from meat and butchers' shops by using PCR technique in Mosul city. doi:10.33899/ijvs.2023.136964.2632 Sheet, O.H., Al-Mahmood, O.A., Taha, A.H., Alsanjary, R.A., Plötz, M. and Abdulmawjood, A.A. 2025. Molecular Detection of Methicillin-resistant Staphylococcus aureus Isolated From Foods in Germany Using LAMP Assay. Egypt. J. Vet. Sci. 56(6), 1153–1160; doi:10.21608/EJVS.2024.281105.1981 Sheet, O.H., Al-Mahmood, Y.S., Taha, A.H. and Abdulmawjood, A.A. 2023. Detection the spa type of methicillin-resistant Staphylococcus aureus isolated from local Basturma in Mosul city, Iraq. Iraqi J. Vet. Sci. 38(4), 739–745; doi:10.33899/ijvs.2024.148254.3569 Taha, A.H., Al-Mahmood, O.A., Sheet, O.H., Hamed, A.A., Al-Sanjary, R.A. and Abdulmawjood, A.A. 2024. Molecular detection of methicillin resistant Staphylococcus aureus isolated from local fish in Mosul city. Iraqi J. Vet. Sci. 38(2), 437–441; doi:10.33899/ijvs.2023.142707.3191 Taha, A.H., Sheet, O.H. and Jwher, D.M. 2025. Molecular detection and phylogenetic diversity of Staphylococcus aureus isolated from goat subclinical mastitis in Nineveh governorate. Iraqi. J. Vet. Sci. 39(1), 71–79; doi:10.33899/ijvs.2024.152066.3791 Tristan, A., Ying, L., Bes, M., Etienne, J., Vandenesch, F. and Lina, G. 2003. Use of multiplex PCR to identify Staphylococcus aureus adhesins involved in human hematogenous infections. J. Clin. Microbiol. 41(9), 4465–4467; doi:10.1128/jcm.41.9.4465-4467.2003 Tsen, H.Y. and Chen, T.R. 1992. Use of the polymerase chain reaction for specific detection of type A, D and E enterotoxigenic Staphylococcus aureus in foods. Appl. Microbiol. Biotechnol. 37(5), 685–690; doi:10.1007/bf00182814 Wang, W., Baloch, Z., Jiang, T., Zhang, C., Peng, Z., Li, F., Fanning, S., Ma, A. and Xu, J. 2017. Enterotoxigenicity and antimicrobial resistance of Staphylococcus aureus isolated from retail food in China. Front. Microbiol. 8, 2256; doi:10.3389/fmicb.2017.02256 Wu, S., Huang, J., Zhang, F., Wu, Q., Zhang, J., Pang, R., Zeng, H., Yang, X., Chen, M., Wang, J., Dai, J., Xue, L., Lei, T. and Wei, X. 2019. Prevalence and characterization of food-related methicillin-resistant Staphylococcus aureus (MRSA) in China. Front. Microbiol. 10, 304; doi:10.3389/fmicb.2019.00304 Yenealem, D.G., Yallew, W.W. and Abdulmajid, S. 2020. Food Safety Practice and Associated Factors among Meat Handlers in Gondar Town: a Cross-Sectional Study. J. Environ. Public. Health. 1(1), 7421745; doi:10.1155/2020 | ||
| How to Cite this Article |
| Pubmed Style Awed MJM, Taha AH, Sheet OH. Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Vet. J.. 2025; 15(12): 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 Web Style Awed MJM, Taha AH, Sheet OH. Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. https://www.openveterinaryjournal.com/?mno=276487 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.15 AMA (American Medical Association) Style Awed MJM, Taha AH, Sheet OH. Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Vet. J.. 2025; 15(12): 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 Vancouver/ICMJE Style Awed MJM, Taha AH, Sheet OH. Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 Harvard Style Awed, M. J. M., Taha, . A. H. & Sheet, . O. H. (2025) Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Vet. J., 15 (12), 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 Turabian Style Awed, Mohammed Jasim Mohammed, Ayman Hani Taha, and Omar Hashim Sheet. 2025. Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Veterinary Journal, 15 (12), 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 Chicago Style Awed, Mohammed Jasim Mohammed, Ayman Hani Taha, and Omar Hashim Sheet. "Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq." Open Veterinary Journal 15 (2025), 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 MLA (The Modern Language Association) Style Awed, Mohammed Jasim Mohammed, Ayman Hani Taha, and Omar Hashim Sheet. "Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq." Open Veterinary Journal 15.12 (2025), 6316-6328. Print. doi:10.5455/OVJ.2025.v15.i12.15 APA (American Psychological Association) Style Awed, M. J. M., Taha, . A. H. & Sheet, . O. H. (2025) Molecular detection of the sea, seb, and sec genes and phylogenetic tree of methicillin-resistant Staphylococcus aureus isolated from restaurants in Erbil city, Iraq. Open Veterinary Journal, 15 (12), 6316-6328. doi:10.5455/OVJ.2025.v15.i12.15 |