Open Veterinary Journal, (2025), Vol. 15(12): 6342-6350
Research Article
10.5455/OVJ.2025.v15.i12.17
Buffalo rumen microbial cellulase as a candidate enzyme for hydrolysis of rice straw into glucose: A preliminary study
Agung Prabowo1, Slamet Hartanto2, Dini Dwi Ludfiani2, Bambang Haryanto1, Miranti Dian Pertiwi1, Abd. Azis Wasil1, Maya Larasati Donna Wardani1, Nandari Dyah Suretno2, Rini Nur Hayati2,
Forita Dyah Arianti1, Arif Dwi Santoso1, Megawati Megawati3, Widi Astuti3, Nuruliarizki Shinta Pandupuspitasari4*, Asih Kurniawati5 and Zaenal Bachruddin5
1Research Center for Sustainable Industrial and Manufacturing Systems, National Research and Innovation Agency, Banten, Indonesia
2Research Center for Animal Husbandry, National Research and Innovation Agency, Bogor Indonesia
3Chemical Engineering Department, Engineering Faculty, Universitas Negeri Semarang, Semarang, Indonesia
4Faculty of Animal and Agricultural Sciences, Universitas Diponegoro, Semarang, Indonesia
5Faculty of Animal Science, Gadjah Mada University, Yogyakarta, Indonesia
*Corresponding Author: Nuruliarizki Shinta Pandupuspitasari. Faculty of Animal and Agricultural Sciences, Universitas Diponegoro, Semarang, Indonesia. Email: shin_tse [at] yahoo.com
Submitted: 08/08/2025 Revised: 10/11/2025 Accepted: 20/11/2025 Published: 31/12/2025
© 2025 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Background: The buffalo rumen microbiome is enriched in functional genes, notably those encoding beta-glucosidase, which participate in polysaccharide breakdown. However, the characterization of enzymes and the application of buffalo rumen in biomass fermentation remain underdeveloped.
Aim: This study aimed to elucidate the enzymatic profile of buffalo rumen microbiota and assess the effect of heat of ammonia-based pretreatment combined with enzymatic hydrolysis isolated from buffalo ruminal fluid in rice straw to produce bioethanol.
Methods: A factorial experiment was performed with heat (H) and enzyme (E) combinations. The treatment was as follows: (1) no heat and commercial enzyme (H0E1), (2) heat and commercial enzyme (H1E1), (3) no heat and buffalo rumen enzyme (H0E2), and (4) heat and buffalo rumen enzyme (H1E2) with three replications. The enzyme activity, glucose, and ethanol levels were measured.
Results: Beta-glucosidase enzyme was the highest activity (64.23 ± 12.37 U/g) in buffalo rumen fluid. The activities of carboxymethyl cellulase, xylanase, and exoglucanase were 50.36 ± 8.05, 17.44 ± 5.95, and 0.20 ± 0.03 U/g, respectively. The interaction treatments impacted (p < 0.01) the glucose production. H1E1 treatment had the highest glucose content (193.94 ± 14.51 g/L) among all treatments. Moreover, the glucose level in the H0E1 group was higher than that in the H0E2 and H1E2 groups (102.96 ± 27.59 vs. 0.91 ± 0.15 and 1.32 ± 0.37 g/L, respectively). The combination treatments had no effect (p > 0.05) on ethanol production. Heat treatment did not (p > 0.05) influence the ethanol content. However, enzyme treatment affected (p < 0.01) the ethanol level. The ethanol production in the H0E1 and H1E1 groups (0.55% ± 0.13% and 0.99% ± 0.02% v/v) was higher than the ethanol content in the H0E2 and H1E2 treatments (0.19% ± 0.11% and 0.28% ± 0.10% v/v).
Conclusion: Buffalo rumen-derived cellulase can convert cellulose in rice straw into glucose. However, the glucose content in the buffalo rumen enzyme treatment was lower than that of the commercial enzyme.
Keywords: Bioethanol, Biomass, Buffalo rumen microbiome.
Introduction
Renewable energy is extensively studied as a significant strategy to mitigate global warming caused by fossil fuel use (Antar et al., 2021). Lignocellulosic biomass from agricultural waste is one of the putative sustainable energy sources to substitute fossil fuels because it is abundant and inexpensive (Inyang et al., 2022). They are then converted into bioethanol via fermentation (Fatma et al., 2018). However, to date, biomass is predominantly disposed of in processes such as burning, natural decomposition, and landfilling. They only account for 14% of energy consumption (Siwal et al., 2021).
Rice straw is an agricultural biomass containing more than 50% cellulose and hemicellulose; thus, it is a crucial feedstock for bioethanol production (Takano and Hoshino, 2018). Indonesia has a substantial supply of RSW as a potential feedstock for sustainable bioethanol production (Erdiwansyah et al., 2024). The total production of rice in Indonesia reached 53.14 million tons in 2024 (Badan Pusat Statistik, 2025), and it is estimated that 25%–30% of rice straw waste is produced, equivalent to 13.28–15.94 million tons (Morya et al., 2023). Previous research indicates that approximately 28.12% of rice straw remains unused in the field in Indonesia after harvest (Arianti et al., 2024). Therefore, there are potentially 3.74–4.48 million tons of rice straw to be used to produce biofuels in Indonesia.
One of the key steps to elevate cellulose and hemicellulose conversions into sugars in pretreated biomass is enzymatic hydrolysis (Vasić et al., 2021). β-Glucosidases, exoglucanases (cellobiohydrolases), and endoglucanases (cellulases) are the primary enzymes responsible for cellulose degradation (Basera et al., 2024). β-Glucosidase catalyzes the terminal step of cellulose degradation by converting cellobiose into glucose, thus completing the action of the cellulase system. Exoglucanase acts on β-1,4-glycosidic bonds from the ends of cellulose chains, primarily releasing cellobiose units. Together with endoglucanases, which cleave internal bonds, these enzymes cooperate in a synergistic cascade to convert cellulose into glucose (John et al., 2022). Moreover, xylanases participate in the breakdown of plant cell wall hemicelluloses and enhance cellulose hydrolysis through interactions with cellulase enzymes (Dias et al., 2022).
Rumen fluid contains diverse microbiota involved in cellulose degradation (Tamilselvan and Immanuel Selwynraj, 2024) via secreted enzymes (Liang et al., 2024). Rumen microorganisms enhance biofuel production through diverse enzymatic systems and synergistic metabolic interactions (Bhujbal et al., 2022). These microbiotas produce cellulase enzymes that potentially replace the high cost of commercial enzymes. The use of cow, goat, and sheep rumen fluid has significantly increased the digestion of lignocellulosic substrates in biofuel production (Sema et al., 2020). Furthermore, the microbiome of buffalo rumen is significantly enriched in functional genes, notably those encoding β-glucosidases. These enzymes belong to glycosyl hydrolase families 1 and 3 (GH1 and GH3), which participate in polysaccharide breakdown (Chen et al., 2012). The metagenomic study by Singh et al. (2014) also found that the gene encoding the GH3 enzyme was abundantly detected in the buffalo rumen microbiome, which facilitates efficient cellulose degradation. Therefore, the buffalo rumen can be used to enhance biofuel production.
Pretreatment and hydrolysis are essential processes for optimizing lignocellulosic biomass fermentation in ethanol production (Duque et al., 2021). These processes are the initial stages of polysaccharide depolymerization into monosaccharide units in bioethanol production (Kucharska et al., 2020). Alkali treatments enhance the raw material node by breaking the ester and glycosidic bonds in the lignocellulosic cell wall (Das et al., 2023). These methods improve cellulose and hemicellulose accessibility for the conversion of fermentable sugars. Efficient ethanol production from lignocellulosic biomass is significantly enhanced by a precise combination of pretreatment methods, including optimum temperature, reaction time length, and alkaline concentration (Tan et al., 2021). High temperature is a vital step in alkali-catalyzed pretreated biomass, as it increases lignin removal (Yang et al., 2019).
However, the characterization of enzymes and the application of buffalo rumen fluid in biomass fermentation, combined with pre-heating treatment, remain underdeveloped. Thus, this study aimed to elucidate the enzymatic profile of buffalo rumen microbiota and assess the effect of heat of ammonia-based pretreatment combined with enzymatic hydrolysis isolated from buffalo ruminal fluid in rice straw to produce bioethanol in Indonesia. This research provided novel insights regarding the applicability of buffalo rumen in the production of biofuels.
Materials and Methods
Data on microbial enzymes in buffalo rumen
Rumen microbes were isolated from the rumen contents of Indonesian swamp buffaloes collected from the slaughterhouse of Kudus Regency Agriculture and Food Service in Prambatan Kidul Village, Kaliwungu District, Kudus Regency, Central Java Province, Indonesia. The rumen contents were placed in a container, stored in a cooler box at 2°C–4°C, and transported to the laboratory. Then, the rumen fluid sample was kept at 2°C–4°C before further treatment. The rumen microbe was grown anaerobically in 0.50 g rice straw (Inpari 32 variety), 0.10 g (NH4)2SO4, 0.01 g MgSO4.7H2O, 0.20 g NaCl, 0.50 g Na2C03, 0.70 g K2HPO4, 0.15 ml resazurin (0.1%), 0.50 g yeast, 0.50 g cysteine HCl (3%), 30 ml clear rumen liquor, and 70 ml distilled water at 45°C and pH 7 for 7 days. The enzymatic activities of carboxymethyl cellulase, beta-glucosidase, exoglucanase, and xylanase were measured.
Research design
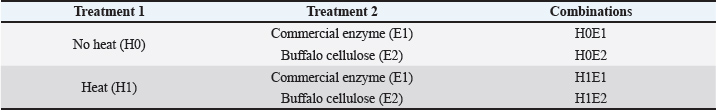
This study used a completely randomized design with a 2 × 2 factorial design with three replications (Table 1). The first treatment factor was heat treatment (H1) and heat treatment without heat (H0). Hydrolysis with the buffalo rumen microbial cellulase enzyme (E2) and commercial cellulase enzyme (E1) (Viscozyme Cassava® PL) was used as the second treatment. Therefore, the combination of treatment groups was (1) no heat and commercial enzyme (H0E1), (2) heat and commercial enzyme (H1E1), (3) no heat and buffalo rumen enzyme (H0E2), and (4) heat and buffalo rumen enzyme (H1E2). Figure 1 shows the research steps in the study.
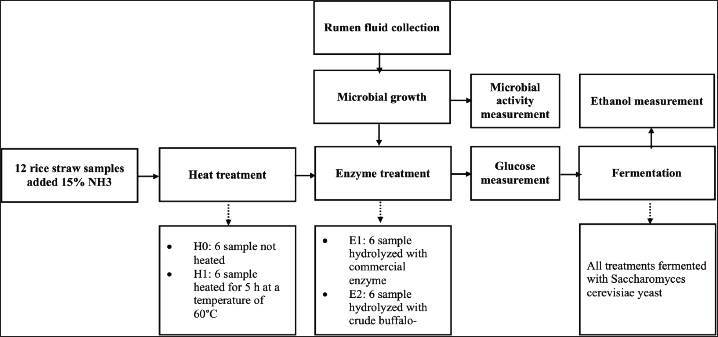
Fig. 1. Steps in the study.
Table 1. Experimental design.

Heating treatment
A total of 12 samples of 80 mesh dry milled straw (Inpari 32 variety), each 25 g, were placed in a two-neck flat-bottom flask, and 300 ml of 15% v/v ammonia solution was added. For the heating treatment (H1), 6 out of the 12 samples were heated by reflux for 5 hours at 60°C, after which the samples were cooled (Park and Kim, 2012). All samples were then filtered, and the residue was washed with distilled water, and 1 M H2SO4 was added until the pH was 7 (neutral). The filtered residue was dried at 50°C for 48 hours. The dried residue was weighed and used for enzymatic hydrolysis.
Hydrolysis of BM enzymes
A total of 12 samples (six heated samples and six non-heated samples), each containing 10 g of residue at the pretreatment stage, were placed in a 250 ml Erlenmeyer flask, and 100 ml of distilled water was added, followed by stirring until homogeneity was achieved. All samples were sterilized in an autoclave at 121°C for 15 minutes. One milliliter was taken for glucose testing. Cellulolytic enzymes (commercial or buffalo-derived) were added to heated and non-heated samples at 5% of the substrate volume, and the mixtures were stirred until they were evenly distributed. The sample was then adjusted to pH 5 by adding 0.1 N HCl. The Erlenmeyer flask was tightly closed using aluminum foil. The temperature of the shaker incubator was set to 39°C, and the material was incubated for 12 hours at 160 rpm for 12 hours. After 12 hours, the material was filtered through a filter paper after 12 hours. The filtrate of the obtained material was used for fermentation and glucose tests according to the method described by Nelson (1944).
Fermentation
A total of 12 samples (50 ml each of hydrolysis filtrate) were placed in an Erlenmeyer flask, and 1 mol/l H2SO4 solution was added until pH 4.6. The Erlenmeyer flask was then covered with aluminum foil. This process was repeated 12 times according to the number of samples collected. All samples were autoclaved at 121°C for 15 minutes. After sterilization, 4 g of Saccharomyces cerevisiae yeast and sorbitan monostearate (E491) (Firmipan, Lesaffre Inc., France), 0.15 g of urea, and 0.045 g of NPK fertilizer (Petrokimia Gresik Inc., Indonesia) containing 15% nitrogen (N), 15% phosphate (P2O5), 15% potash (K2O), and 0.1% zinc (Zn) were added to each sample solution and stirred until evenly distributed. Furthermore, the Erlenmeyer was tightly closed using an airlock. Fermentation was performed anaerobically for 5 days at 30°C in an incubator. Ethanol content was measured according to the method described by MacLeod (1949).
Statistical analysis
Descriptive analysis was employed to analyze enzyme activity in the rumen of buffalo. Data were presented in mean ± SE. The effect of heat pretreatment and the enzyme isolated from buffalo in hydrolysis was synthesized using a one-way analysis of variance. Duncan’s multiple range test was performed to detect the significant differences in the effects of treatments. All analyses were conducted in the R program (R Core Team, 2023).
Ethical approval
Not required for this study.
Results
Enzyme profile
The study found that the buffalo ruminal microbiota secreted the enzymes carboxymethyl cellulase, exoglucanase, xylanase, and beta-glucosidase (Fig. 2). The highest enzyme activity in the buffalo rumen was β-glucosidase (64.23 ± 12.37 U/g). Carboxymethyl cellulase, xylanase, and exoglucanase contents were 50.36 ± 8.05, 17.44 ± 5.95, and 0.20 ± 0.03 U/g, respectively.
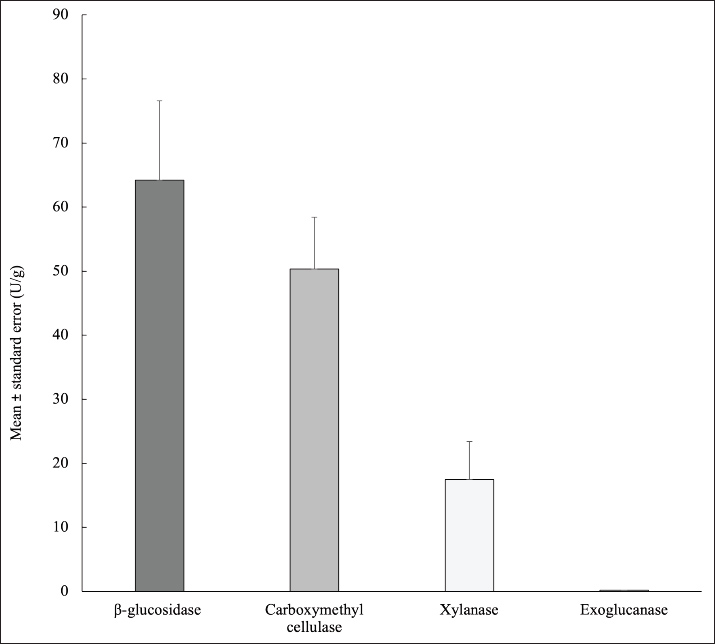
Fig. 2. Rumen enzyme activity in swamp buffalo in Indonesia.
Effect of treatments on the glucose and ethanol contents
The interaction treatments impacted (p < 0.01) the glucose production (Fig. 3). H1E1 treatment had the highest glucose content (193.94 ± 14.51 g/L) among all treatments. Moreover, the glucose level in the H0E1 group was higher than that in the H0E2 and H1E2 groups (102.96 ± 27.59 vs. 0.91 ± 0.15 and 1.32 ± 0.37 g/L, respectively). The combination treatments had no effect (p > 0.05) on ethanol production (Fig. 4).. Heat treatment did not (p > 0.05) influence the ethanol content. However, enzyme treatment affected (p < 0.01) the ethanol level. The ethanol production in the H0E1 and H1E1 groups (0.55% ± 0.13% and 0.99% ± 0.02% v/v) was higher than the ethanol content in the H0E2 and H1E2 treatments (0.19% ± 0.11% and 0.28% ± 0.10% v/v).
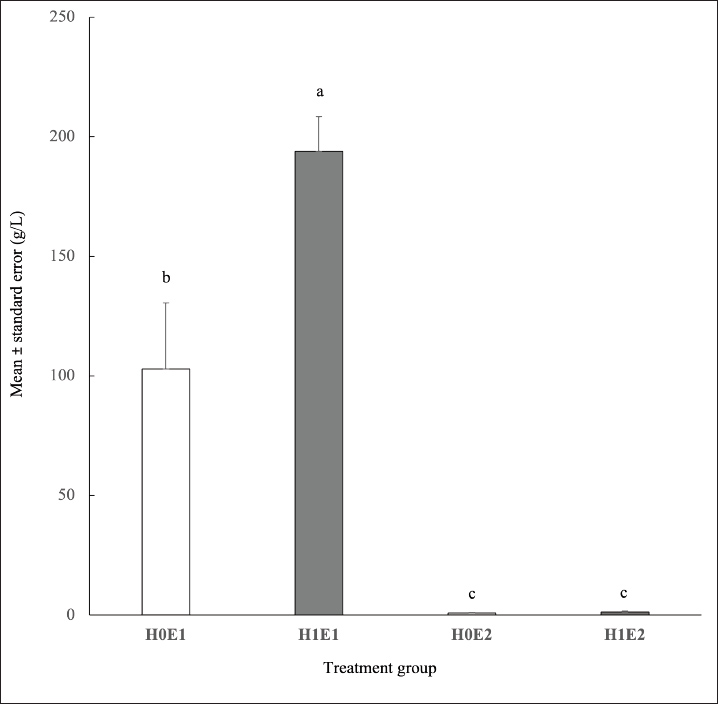
Fig. 3. Effect of treatments on glucose content (a, b denote statistically significant differences among interaction treatments, p < 0.02, F-value=8.44).
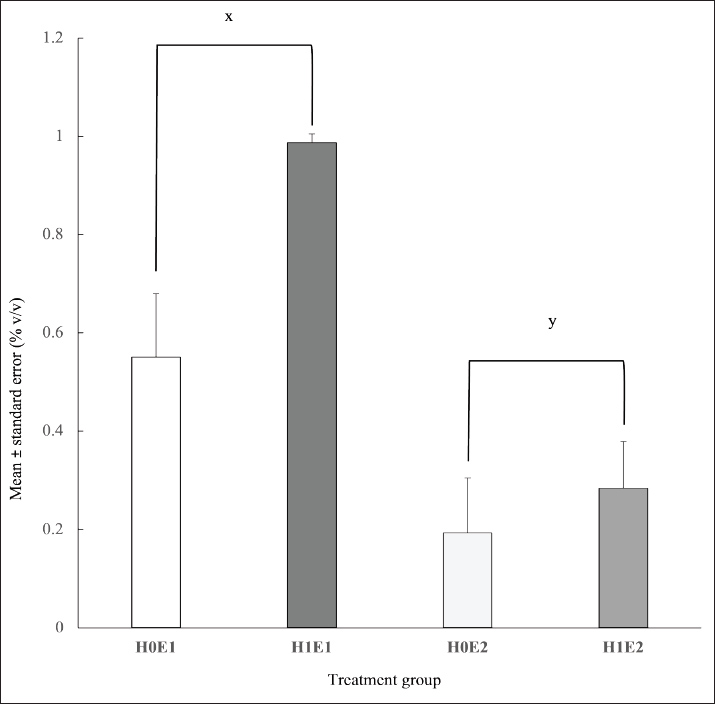
Fig. 4. Influence of treatments on ethanol production (x, y denote statistically significant differences among enzyme treatments, p < 0.003, F-value=15.09).
Discussion
The findings indicated that ruminal fluid contained cellulases that could degrade cellulose. Microbial rumens secrete cellulolytic enzymes to degrade cellulose into volatile fatty acids as the main nutrient source for ruminants (Weimer, 2022). Cellulose is broken down by a set of enzymes working together: first, endoglucanases cut internal links in the cellulose chain, then exoglucanases remove mainly cellobiose units from the ends, and finally β-glucosidase converts those cellobiose units into glucose (John et al., 2022). Furthermore, xylanases dismantle the hemicellulose and enhance cellulase enzymes' access and hydrolyze cellulose (Dias et al., 2022).
Metagenomic studies have shown an essential role in uncovering novel genes encoding cellulase and other lignocellulolytic enzymes from the rumen microbiome. The study characterized and cloned Cel-3.1 from buffalo rumen as a novel cellulase (Pabbathi et al., 2021). Moreover, the study found that Cel-3.1 had a robust capability to degrade and hydrolyze lignocellulose in rice and corn straw. Meng et al. (2023) also discovered a novel cellulase, CelXyn2, in buffalo rumen. Enzyme enhances lignocellulose hydrolysis and fermentation in vitro metagenomic studies have also found that the gene encoding GH3 enzyme was abundantly detected in the buffalo rumen microbiome, which contributes to cellulose digestion (Chen et al., 2012; Singh et al., 2014).
The study found that β-glucosidases activity was highest in the buffalo rumen. Interestingly, the enzyme profile in buffalo rumen in the study is in contrast with the enzyme activity in sheep and cattle ruminal fluids. Santra et al. (2007) reported that beta-glucosidase activity was lower than that of carboxymethyl cellulase and xylanase in the sheep rumen. Moreover, xylanase activity was higher than that of carboxymethyl cellulase in the rumen of pre- and post-weaning cattle (Hao et al., 2021). The discrepancy in enzyme profile between buffalo and cattle may be attributable to differences in laboratory methodology. It might also be influenced by diet composition and environmental rumen condition (Palmonari et al., 2024).
However, the finding confirmed a previous metagenomic study demonstrating that the reported buffalo rumen hosts various microbiota that secrete potent cellulolytic enzymes, particularly β-glucosidases. A metagenomic investigation revealed that genes encoding GH3 family β-glucosidases are abundantly represented within the buffalo rumen microbiome (Singh et al., 2014). These enzymes are involved in the production of biofuels from lignocellulosic biomass. They play a key role in enzyme mixtures used to degrade cellulose biomass (Santos et al., 2019). β-glucosidase facilitates the final stage of cellulose enzymatic hydrolysis by breaking β-1,4-glycosidic bonds in cellobiose to produce ethanol (Azevedo et al., 2025). This result is similar to that of Chen et al. (2012), who reported that the novel β-glucosidase NpaBGS isolated from buffalo rumen was a potential enzyme for simultaneous saccharification and fermentation. Thus, the enzyme profile identified in buffalo rumen fluid is a significant candidate for enzymatic hydrolysis application in biofuel production.
Pretreatment with 15% ammonia at 60 °C, followed by enzymatic hydrolysis using the enzyme cocktail (H1E1), had the highest glucose and ethanol contents. This finding is similar to that of Xu et al. (2023), who reported that pretreatment with 9% ammonia autoclaved at 60 °C effectively increased cellulose and decreased lignin content in corn straw. The results indicated that the combined pretreatment effectively degraded lignin to increase the cellulose content of rice straw. High temperature disintegrates the lignin structure and enhances the pore volume of the plant cell wall (Donohoe et al., 2008). Moreover, lignin degradation is associated with enhanced cellulose accessibility (Zheng et al., 2018). Thus, it increases the cellulose conversion rate.
This study discovered that the fermentation of all pretreated rice straw and hydrolysis using native enzymes isolated from buffalo ruminal (H1E2 and H0E2) successfully converted cellulose into glucose. However, these groups have lower concentrations than the commercial cocktail enzyme (H1E1 and H0E1) groups. The ruminal fluid possessed an elevated β-glucosidase level. β-glucosidases catalyze the hydrolysis of soluble cello-oligosaccharides into glucose. Their effective actions depend on the preliminary actions of cellobiohydrolase and endoglucanases to degrade insoluble cellulose fibers into soluble substrates accessible to β-glucosidases. Cellobiohydrolase and endoglucanase actions to attain enzymatic efficiency are impacted by β-glucosidase activity that converts cellobiose into glucose. The efficiency of biomass enzymatic hydrolysis is attained by a balanced enzyme formulation (Sørensen et al., 2013). The identification of the optimal level and combination of the best-performing mono-active enzymatic activities is key factor to achieve the conversion of cellulose to monomeric sugar units (Azevedo et al., 2025). Thus, identifying the optimal level and combination of enzymes isolated from buffalo rumen with other cellulose enzymes is essential to enhance the efficiency of cellulose degradation in biomass.
Conclusion
The study showed that the buffalo rumen contained cellulase activity, namely 64.23 ± 12.37 U/g β-glucosidases, 50.36 + 8.05 U/g carboxymethyl cellulase, 17.44 + 5.95 U/g xylanase, and 0.20 + 0.03 U/g exoglucanase. The buffalo rumen-derived cellulase can convert cellulose in rice straw into glucose. However, the glucose content in the buffalo rumen enzyme treatment was lower than that of the commercial enzyme. The optimal dosage and combination of enzymes isolated from the buffalo rumen with other cellulolytic enzymes are crucial for improving the efficiency of cellulose degradation in biomass.
Acknowledgments
The authors would like to thank the Energy and Manufacturing Research Organization, Research Center for Sustainable Industrial and Manufacturing Systems (RC SIMS). Riset dan Inovasi untuk Indonesia Maju, Indonesia Endowment Fund for Education (LPDP) of the Ministry of Finance, Republic of Indonesia, Gadjah Mada University, Diponegoro University, and Semarang State University.
Conflict of interest
The authors declare no potential conflicts of interest regarding the publication of this work.
Funding
This research was funded by the RIIM LPDP Grant and BRIN (grant number B 3837/II.7.5/FR.06.00/11/2023 and B-3977/III.3/FR.06.00/11/2023).
Authors’ contributions
Conceptualization, A.P.; S.H.; D.D.L.; M.D.P.; F.D.A.; A.D.S.; W.A.; and N.S.P.; Methodology, A.P.; S.H.; D.D.L.; B.H.; R.N.H.; F.D.A.; A.D.S.; M.M.; W.A.; N.S.P.; A.K.; and Z.B.; Software, D.D.L.; B.H.; M.D.P.; A.A.W.; M.L.D.M.; N.D.S.; and R.N.H.; Validation and investigation, A.P.; S.H.; B.H.; M.D.P.; M.L.D.M.; R.N.H.; F.D.A.; A.D.S.; and N.S.P.; Formal analysis and resources, A.P.; A.A.W.; M.L.D.M.; N.D.S.; A.D.S.; M.M.; W.A.; A.K.; and Z.B.; Recognized data and information, A.P.; S.H.; D.D.L.; B.H.; M.D.P.; A.A.W.; M.L.D.M.; N.D.S.; F.D.A.; A.D.S.; M.M.; W.A.; N.S.P.; A.K.; and Z.B.; Writing—original draft preparation, A.P.; S.H.; D.D.L.; B.H.; M.D.P.; A.A.W.; M.L.D.M.; N.D.S.; R.N.H.; F.D.A.; A.D.S.; M.M.; W.A.; A.K.; and Z.B.; Writing—review and editing, A.P.; S.H.; B.H.; M.D.P.; A.A.W.; M.L.D.M.; N.D.S.; R.N.H.; F.D.A.; M.M.; W.A.; N.S.P.; A.K.; Z.B.
Data availability
We will provide data on demand.
References
Antar, M., Lyu, D., Nazari, M., Shah, A., Zhou, X. and Smith, D.L. 2021. Biomass for a sustainable bioeconomy: an overview of world biomass production and utilization. Renew. Sustain. Energy. Rev. 139, 110691.
Arianti, F.D., Triastono, J., Pertiwi, M.D., Prabowo, A., Prasetyo, T., Haryanto, C., Astuti, M., Djarot, I.N., Santoso, A.D. and Wijayanti, S.P. 2024. Renewable energy potential of rice straw and paunch manure as bioethanol feedstocks in Central Java, Indonesia. Case. Stud. Chem. Environ. Eng. 9, 100677.
Azevedo, R.D.S., Santana, H., Seus, V.R., Camargo, A.D., Werhli, A.V., Machado, K.D.S., Cançado, L.J., Quirino, B.F. and Marins, L.F. 2025. Development of a β-glucosidase improved for glucose retroinhibition for cellulosic ethanol production: an integrated bioinformatics and genetic engineering approach. Biotechnol. Biofuels Bioprod. 18, 44.
Badan Pusat Statistik. 2025. Luas Panen dan Produksi Padi di Indonesia 2024 (Angka Tetap) (No. 15/02/Th. XXVIII). Badan Pusat Statistik, Jakarta.
Basera, P., Chakraborty, S. and Sharma, N. 2024. Lignocellulosic biomass: insights into enzymatic hydrolysis, influential factors, and economic viability. Discovery. Sustainability. 5, 311.
Bhujbal, S.K., Ghosh, P., Vijay, V.K., Rathour, R., Kumar, M., Singh, L. and Kapley, A. 2022. Biotechnological potential of rumen microbiota for sustainable bioconversion of lignocellulosic waste to biofuels and value-added products. Sci. Total Environ. 814, 152773.
Chen, H.L., Chen, Y.C., Lu, M.Y.J., Chang, J.J., Wang, H.T.C., Ke, H.M., Wang, T.Y., Ruan, S.K., Wang, T.Y., Hung, K.Y., Cho, H.Y., Lin, W.T., Shih, M.C. and Li, W.H. 2012. A highly efficient β-glucosidase from the buffalo rumen fungus Neocallimastix patriciarum W5. Biotechnol. Biofuels 5, 24.
Das, N., Jena, P.K., Padhi, D., Kumar Mohanty, M. and Sahoo, G. 2023. A comprehensive review of characterization, pretreatment and its applications on different lignocellulosic biomass for bioethanol production. Biomass. Convers. Biorefinery. 13, 1503–1527.
Dias, I.K.R., Siqueira, G.A. and Arantes, V. 2022. Xylanase increases the selectivity of the enzymatic hydrolysis with endoglucanase to produce cellulose nanocrystals with improved properties. Int. J. Biol. Macromol. 220, 589–600; doi:10.1016/j.ijbiomac.2022.08.047
Donohoe, B.S., Decker, S.R., Tucker, M.P., Himmel, M.E. and Vinzant, T.B. 2008. Visualizing lignin coalescence and migration through maize cell walls following thermochemical pretreatment. Biotechnol. Bioeng. 101, 913–925.
Duque, A., Álvarez, C., Doménech, P., Manzanares, P. and Moreno, A.D. 2021. Advanced Bioethanol Production: from Novel Raw Materials to Integrated Biorefineries. Processes 9(2), 206.
Fatma, S., Hameed, A., Noman, M., Ahmed, T., Shahid, M., Tariq, M., Sohail, I. and Tabassum, R. 2018. Lignocellulosic Biomass: a Sustainable Bioenergy Source for the Future. Protein. Pept. Lett. 25, 148–163.
Erdiwansyah, Gani, A., Mamat, R., Bahagia, Nizar, M., Yana, S., Mat Yasin, M.H., Muhibbuddin and Rosdi, S.M. 2024. Prospects for renewable energy sources from biomass waste in Indonesia. Case. Stud. Chem. Environ. Eng. 10, 100880.
Hao, Y., Guo, C., Gong, Y., Sun, X., Wang, W., Wang, Y., Yang, H., Cao, Z. and Li, S. 2021. Rumen Fermentation, Digestive Enzyme Activity, and Bacteria Composition between Pre-Weaning and Post-Weaning Dairy Calves. Animals 11, 2527.
Inyang, V., Laseinde, O.T. and Kanakana, G.M. 2022. Techniques and applications of lignocellulose biomass sources as transport fuels and other bioproducts. Int. J. Low-Carbon. Technol. 17, 900–909.
John J, A., Samuel, M.S., Govarthanan, M. and Selvarajan, E. 2022. A comprehensive review on strategic study of cellulase producing marine actinobacteria for biofuel applications. Environ. Res. 214(Pt 3), 114018.
Kucharska, K., Słupek, E., Cieśliński, H. and Kamiński, M. 2020. Advantageous conditions of saccharification of lignocellulosic biomass for biofuels generation via fermentation processes. Chem. Pap. 74, 1199–1209.
Liang, J., Zhang, R., Chang, J., Chen, L., Nabi, M., Zhang, H., Zhang, G. and Zhang, P. 2024. Rumen microbes, enzymes, metabolisms, and application in lignocellulosic waste conversion - A comprehensive review. Biotechnol. Adv. 71, 108308.
Macleod, L.D. 1949. Determination of alcohol by microdiffusion. J. Biol. Chem. 181, 323–331.
Meng, Z., Ma, J., Sun, Z., Yang, C., Leng, J., Zhu, W. and Cheng, Y. 2023. Characterization of a novel bifunctional enzyme from buffalo rumen metagenome and its effect on in vitro ruminal fermentation and microbial community composition. Anim. Nutr. 13, 137–149.
Morya, R., Andrianantenaina, F.H., Singh, S., Pandey, A.K., Kim, G.B., Verma, J.P., Kumar, G., Raj, T. and Kim, S.H. 2023. Exploring rice straw as substrate for hydrogen production: critical challenges and opportunities. Environ. Technol. & Innov. 31, 103153.
Nelson, N. 1944. A Photometric Adaptation of the Somogyi Method for the Determination of Glucose. J. Biol. Chem. 153, 375–380.
Pabbathi, N.P.P., Velidandi, A., Gandam, P.K., Koringa, P., Parcha, S.R. and Baadhe, R.R. 2021. Novel buffalo rumen metagenome derived acidic cellulase Cel-3.1 cloning, characterization, and its application in saccharifying rice straw and corncob biomass. Int. J. Biol. Macromol. 170, 239–250.
Palmonari, A., Federiconi, A. and Formigoni, A. 2024. Animal board invited review: the effect of diet on rumen microbial composition in dairy cows. Animal 18(10), 101319.
Park, Y.C. and Kim, J.S. 2012. Comparison of various alkaline pretreatment methods of lignocellulosic biomass. Energy 47(1), 31–35.
R Core Team. 2023. R: A Language and Environment for Statistical Computing version 4.2.2. Vienna, Austria: R Foundation for Statistical Computing.
Santos, C.A., Morais, M.A.B., Terrett, O.M., Lyczakowski, J.J., Zanphorlin, L.M., Ferreira-Filho, J.A., Tonoli, C.C.C., Murakami, M.T., Dupree, P. and Souza, A.P. 2019. An engineered GH1 β-glucosidase displays enhanced glucose tolerance and increased sugar release from lignocellulosic materials. Sci. Rep. 9, 4903.
Santra, A., Karim, S.A. and Chaturvedi, O.H. 2007. Rumen enzyme profile and fermentation characteristics in sheep as affected by treatment with sodium lauryl sulfate as defaunating agent and presence of ciliate protozoa. Small Ruminant Res. 67, 126–137.
Sema, Y.A., Urgessa, O.E. and Ayele, A.K. 2020. Evaluating Rumen Fluid-Inoculated Lignocellulosic Substrates for Biogas Production. East Afr. J. Sci. 14(2), 99–110.
Singh, K.M., Reddy, B., Patel, D., Patel, A.K., Parmar, N., Patel, A., Patel, J.B. and Joshi, C.G. 2014. High Potential Source for Biomass Degradation Enzyme Discovery and Environmental Aspects Revealed through Metagenomics of Indian Buffalo Rumen. Biomed. Res. Int. 1, 1–10.
Siwal, S.S., Zhang, Q., Devi, N., Saini, A.K., Saini, V., Pareek, B., Gaidukovs, S. and Thakur, V.K. 2021. Recovery processes of sustainable energy using different biomass and wastes. Renew. Sustain. Energy. Rev. 150, 111483.
Sørensen, A., Lübeck, M., Lübeck, P. and Ahring, B. 2013. Fungal Beta-Glucosidases: a Bottleneck in Industrial Use of Lignocellulosic Materials. Biomolecules 3, 612–631.
Takano, M. and Hoshino, K. 2018. Bioethanol production from rice straw by simultaneous saccharification and fermentation with statistical optimized cellulase cocktail and fermenting fungus. Bioresour. Bioprocess. 5, 16.
Tamilselvan, R. and Immanuel Selwynraj, A. 2024. Enhancing biogas generation from lignocellulosic biomass through biological pretreatment: exploring the role of ruminant microbes and anaerobic fungi. Anaerobe 85, 102815.
Tan, J., Li, Y., Tan, X., Wu, H., Li, H. and Yang, S. 2021. Advances in Pretreatment of Straw Biomass for Sugar Production. Front. Chem. 9, 696030.
Vasić, K., Knez, Z. and Leitgeb, M. 2021. Bioethanol Production by Enzymatic Hydrolysis from Different Lignocellulosic Sources. Molecules 26, 753.
Weimer, P.J. 2022. Degradation of Cellulose and Hemicellulose by Ruminal Microorganisms. Microorganisms 10, 2345.
Xu, Y., Xu, X., Su, X., Liu, W., Qu, J. and Sun, Y. 2023. Effect of Ammonia–Autoclave Pretreatment on the Performance of Corn Straw and Cow Manure Batch Anaerobic Digestion. Fermentation 9, 178.
Yang, H., Shi, Z., Xu, G., Qin, Y., Deng, J. and Yang, J. 2019. Bioethanol production from bamboo with alkali-catalyzed liquid hot water pretreatment. Bioresour. Technol. 274, 261–266.
Zheng, Q., Zhou, T., Wang, Y., Cao, X., Wu, S., Zhao, M., Wang, H., Xu, M., Zheng, B., Zheng, J. and Guan, X. 2018. Pretreatment of wheat straw leads to structural changes and improved enzymatic hydrolysis. Sci. Rep. 8, 1321.













