Open Veterinary Journal, (2026), Vol. 16(1): 26-40
Review Article
10.5455/OVJ.2026.v16.i1.3
Dietary responses of taste receptor genes to plant-based
diets in farmed fish: A literature review
Leonildo dos Anjo Viagem1,2,3,4*, Jean Nepomuscene Hakizimana2, Cyrus Rumisha1, Brunno da Silva Cerozi4 and Gerald Misinzo2,5
1Department of Animal, Aquaculture and Range Sciences, College of Agriculture, Sokoine University of Agriculture, Morogoro, Tanzania
2SACIDS Africa Centre of Excellence for Infectious Diseases, SACIDS Foundation for One Health, Sokoine University of Agriculture, Morogoro, Tanzania
3Institute of Natural Resources and Environment, Rovuma University, Cabo Delgado, Mozambique
4Department of Animal Science, College of Agriculture, University of São Paulo, São Paulo, Brazil
5Department of Microbiology, Parasitology and Biotechnology, College of Veterinary Medicine and Biomedical Sciences, Sokoine University of Agriculture, Morogoro, Tanzania
*Corresponding Author: Leonildo dos Anjo Viagem. Department of Animal, Aquaculture and Range Sciences, College of Agriculture, Sokoine University of Agriculture, Morogoro, Tanzania. Email: lviagem [at] unirovuma.ac.mz
Submitted: 03/07/2025 Revised: 15/11/2025 Accepted: 01/12/2025 Published: 31/01/2026
© 2025 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Taste receptor genes, particularly those in the bitter (tas2r) and umami/sweet (tas1r) families, are critical in regulating chemosensory perception and feeding behavior, influencing fish’s acceptance of plant-based diets. This review synthesizes the current research on how these genes respond to plant-based diets in farmed fish. A comprehensive search of scientific databases, including Web of Science, Scopus, PubMed, and Google Scholar, was conducted to identify relevant studies. Plant-based diets modulate gene expression, receptor activity, and signaling pathways, enabling fish to adapt to alternative feeds. The findings of the studies analyzed in this review show that plant ingredients affect fish taste receptor genes by changing gene expression, receptor activity, and signaling pathways, helping fish adapt to different feeds. Oral receptors detect feed palatability, which affects feed intake, growth, and health, whereas extraoral receptors in the gut, brain, and liver sense nutrients, regulate metabolism, and control appetite. Bitter receptors can detect antinutritional factors and trigger hormone responses that reduce feed intake, whereas umami receptors promote feeding through nutrient-related reward pathways. Functional assays, dose-response studies, and gene expression mapping are essential for identifying receptors that affect the acceptance of plant-based diets. Integrating taste receptor gene knowledge into selective breeding programs may increase feed intake, digestion, and nutrient utilization, thereby supporting sustainable aquaculture. Future research should investigate the relationships between taste receptor expression and behavior and physiology through dietary, genetic, and neurophysiological studies to clarify how interactions between diet, genetics, and neural responses influence feeding, growth, and adaptation to plant-based feeds.
Keywords: Feed acceptance, Fish nutrition, Fish physiology, Plant-based diet, Taste receptor.
Introduction
The global aquaculture sector is expanding rapidly to meet the growing demand for sustainable protein sources. Aquafeeds have relied heavily on fishmeal (FM) and fish oil because of their high nutritional value and palatability (Tacon and Metian, 2008). However, environmental sustainability concerns and the rising cost of FM have encouraged the adoption of plant-based protein alternatives (Gatlin et al., 2007; Naylor et al., 2009). Although many plant-derived proteins provide adequate nutrition, their inclusion in fish diets presents challenges related to taste perception, palatability, digestion, and feed acceptance.
Taste receptor genes, particularly those encoding tas1r (umami/sweet) and tas2r (bitter) receptors, play a central role in regulating these dietary responses by influencing feed intake behavior and nutrient detection (Morais, 2017; Heraud et al., 2022). Fish depend heavily on their chemosensory systems, including taste and olfaction, to assess feed quality and environmental cues. Plant-based diets often contain antinutritional factors (ANFs), such as tannins, saponins, and phytic acid, which can decrease palatability and limit nutrient absorption (Francis et al., 2001). For example, soybean meal, a common substitute for FM, impairs digestion and induces intestinal inflammation in carnivorous species, such as Atlantic salmon (Salmo salar) (Krogdahl et al., 2010). Similarly, behavioral studies have demonstrated that rainbow trout (Oncorhynchus mykiss) and gilthead seabream (Sparus aurata) reduce feed intake when fed high levels of plant proteins, highlighting the importance of sensory cues in feeding behavior (Kaushik et al., 1995; De Francesco et al., 2007).
Taste perception in fish is mediated by taste buds located in both oral and extraoral areas, including the lips, fins, and barbels (Kasumyan and Døving, 2003). These taste buds contain receptor proteins that detect amino acids, sugars, and bitter compounds. The tas1r family of genes (e.g., tas1r1, tas1r2, and tas1r3) responds primarily to amino acids and sweet substances, whereas the tas2r family of genes is specialized for detecting bitter and potentially toxic compounds (Shi and Zhang, 2006). The number and expression of these receptors vary across species, reflecting dietary specialization. Herbivorous and omnivorous fish, such as tilapia (Oreochromis niloticus), generally possess a greater diversity of taste receptor genes than carnivorous species, enabling them to utilize better plant-based feeds (Li and Zhang, 2014; Kong et al., 2018). Moreover, prolonged exposure to plant-based diets can induce changes in the expression of taste receptor genes, suggesting an adaptive mechanism to long-term dietary shifts (Martinat et al., 2025a,b).
Advances in transcriptomics have revealed how taste receptor genes respond to plant-based diets. For example, fish fed with soybean meal often exhibit altered expression of umami receptors (tas1r1, tas1r3) in tissues such as the tongue and gut. This phenomenon has been observed in species such as obscure pufferfish (Takifugu obscurus), indicating a potential mismatch between feed composition and receptor sensitivity (Yao et al., 2019). Additionally, exposure to ANFs can upregulate the expression of bitter taste receptor genes, likely as a protective mechanism against harmful compounds (Wooding et al., 2021).
Synthesizing current knowledge on the dietary responses of taste receptor genes to plant-based diets in farmed fish is essential for improving feed efficiency, enhancing growth performance, and advancing sustainable aquaculture. This review also helps identify research gaps, integrate genetic and nutritional insights, and guide the development of cost-effective, environmentally responsible feeding strategies that safeguard fish health, welfare, and consumer acceptance. Therefore, this literature review aims to review and synthesize existing studies on the dietary responses of taste receptor genes to plant-based diets in farmed fish.
Literature search approach
This literature review employed a structured approach to identify, select, and analyze studies examining the relationships between taste receptor genes and dietary responses to plant-based diets in farmed fish. A comprehensive search was conducted in scientific databases, including Web of Science, Scopus, PubMed, and Google Scholar, using a combination of keywords such as “taste receptor genes,” “fish nutrition,” “plant-based diets,” “aquaculture feed,” and “molecular responses.” Studies published between January 2000 and June 2025 were considered to capture contemporary findings in molecular nutrition and aquaculture practices. The inclusion criteria were as follows: peer-reviewed articles, reviews, and experimental studies that specifically investigated the molecular genetic or physiological responses of fish taste receptors to plant-derived feed ingredients. Studies that addressed only general growth performance without being linked to taste receptor mechanisms were excluded. The reference lists of the selected papers were further screened to capture additional relevant literature.
Fish responses to plant-based diets
Physiological and behavioral responses
In aquaculture, the transition from FM and fish oil to plant-based diets presents significant physiological and behavioral challenges for fish, which vary based on species, diet composition, and adaptation mechanisms (Colombo, 2020). Studies indicate that partially or completely replacing FM with plant-based sources, such as soybeans, seaweed, corn gluten meal, or rapeseed meal, can impact growth performance and feed utilization, while some studies have reported positive effects (Sharawy et al., 2016; Kari et al., 2022). Other studies have highlighted potential drawbacks depending on the plant protein source and the level of replacement, and certain fish species can maintain comparable growth rates on plant-based diets (Valente et al., 2016; Dhanasiri et al., 2020; Kari et al., 2023).
Diet composition, particularly the proportions of protein, carbohydrates, and lipids, can significantly influence digestive enzyme activity. Variations in dietary components may alter the types and quantities of enzymes produced, thereby affecting the efficiency of nutrient breakdown and absorption. For example, protease activity, which plays a key role in protein digestion, is modulated by changes in dietary protein levels (Pavasovic et al., 2006; Jayant et al., 2018; Wang et al., 2018). Long-term consumption of plant-based diets has been linked to metabolic changes, including alterations in key digestive enzymes (Véron et al., 2016). Carnivorous fish, which naturally adapt to animal-based proteins, often show reduced protease and lipase activity when they are transitioned to plant-based diets, potentially leading to inefficient nutrient digestion (Lima et al., 2021; Jiao et al., 2023). Conversely, some omnivorous species adapt more readily because of their ability to produce relatively high levels of cellulase and amylase enzymes (Kurniasih et al., 2024). A study on rainbow trout (Oncorhynchus mykiss), a carnivorous species, revealed that fish fed a plant-based diet without FM ingredients had reduced hepatic enzyme activities, such as glucokinase and pyruvate kinase, which are essential for carbohydrate metabolism. Additionally, an increase in glutamate dehydrogenase activity suggested an imbalance in amino acid metabolism. These enzymatic changes are associated with decreased growth performance and protein retention, indicating potential nutritional deficiencies in plant-based feeds (Véron et al., 2016).
Plant-based diets can also alter the composition of the gut microbiota, promoting beneficial bacteria that increase nutrient digestion by producing digestive enzymes such as amylases, proteases, and lipases. These bacteria contribute essential nutrients and support fish health (El-Sayed et al., 2025). Podell et al. (2023) studied the gut microbiomes of three Hawaiian reef fishes using deep metagenomic sequencing. They reported the dominance of Bacteroidota and Bacillota, microbial enzymes that specialize in the digestion of sulfated polysaccharides from red algae, and shifts in the microbiome composition throughout the gut, emphasizing the key role of gut microbes in marine herbivory and macroalgal digestion.
Additionally, plant-based diets contain various bioactive compounds, such as polysaccharides, polyphenols, and other phytochemicals, which can modulate immune responses by interacting with the fish’s immune system (Nhu et al., 2019; Firmino et al., 2021; Siddik et al., 2023). While some phytochemicals, such as flavonoids and saponins, have immunostimulatory effects, others act as ANFs, inducing stress responses. For example, high levels of soybean meal in fish diets can cause enteritis (intestinal inflammation) in some fish species, such as Atlantic salmon (Salmo salar) and chinook salmon (Oncorhynchus tshawytscha) (Booman et al., 2018); large yellow croaker (Larimichthys crocea) (Zhu et al., 2021); and zebrafish (Danio rerio) larvae (Hedrera et al., 2013), as ANFs affect intestinal morphology and increase susceptibility to infections (Hu et al., 2016; Zhu et al., 2021).
Switching fish to a plant-based diet can act as a physiological stressor, and the initial adaptation phase is often the most challenging, with elevated levels of stress biomarkers such as cortisol and heat shock proteins (Iwama et al., 1998; Varó et al., 2007; Fiocchi et al., 2020). Dietary composition affects fish behavior, which can serve as an indicator of fish welfare. For example, research on two isogenic lines of rainbow trout (Oncorhynchus mykiss) revealed that those fed a plant-based diet displayed reduced aggressive behaviors and stereotypies compared with those fed a fish-based diet. However, they also exhibited increased emotional reactivity when subjected to stress tests, such as isolation in a new environment, suggesting heightened stress sensitivity. This finding indicates that while some undesirable behaviors may decrease, overall welfare could be compromised due to increased stress responsiveness (Sadoul et al., 2016). Additionally, a study on European sea bass (Dicentrarchus labrax) revealed that early exposure to a plant-based diet had minimal effects on behaviors such as self-feeding, learning ability in a T-maze, and exploratory activities. The plant- and FM-fed groups exhibited similar swimming activities and learning capabilities. However, in response to acute stress, the plant-fed group had lower plasma cortisol levels, suggesting that diet composition may influence stress physiology (Benhaïm et al., 2013).
Taste perception and diet adaptation
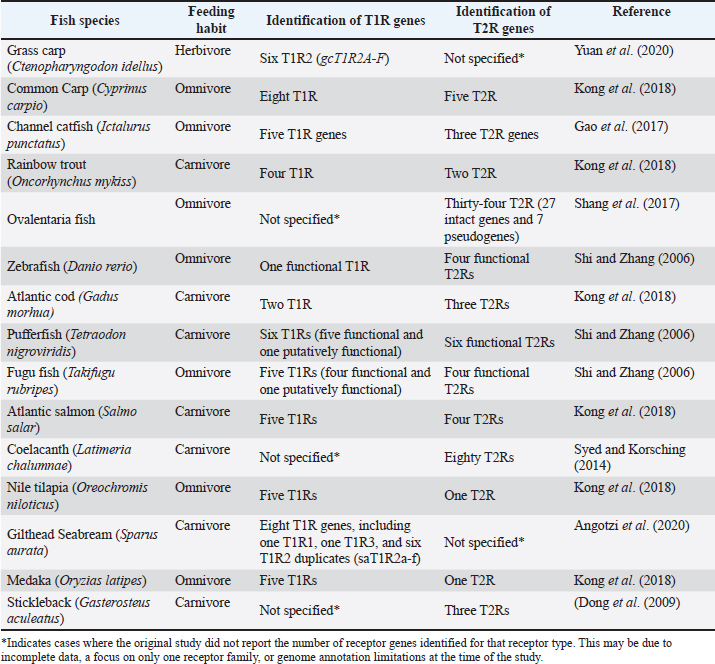
Taste perception in fish is a crucial sensory mechanism for feed detection and acceptance and is primarily mediated by two families of G protein-coupled receptors (GPCRs): type 1 (T1RS/TAS1R) and type 2 (T2RS/TAS2R), which are located in taste buds. Fish taste buds share structural similarities with those of mammals, comprising gustatory receptor cells (Taste/Gustatory Receptor Cells), supporting cells, and basal cells (Boudriot and Reutter, 2001; Ikenaga et al., 2023). Some species, such as the green swordtail (Xiphophorus helleri) and somalian cavefish (Phreatichthys andruzzii), exhibit morphologically diverse taste buds across their heads, reflecting adaptations to specific feeding strategies (Reutter et al., 1974). T1Rs are responsible for detecting amino acids (umami) and sugars (sweet taste), whereas T2Rs recognize bitter compounds, many of which are associated with ANFs in plant-based feeds (Chandrashekar et al., 2000; Morais, 2017; Cai et al., 2021). The diversity and specificity of these receptors vary among fish species, reflecting their dietary habits and ecological niches (Table 1).
Table 1. A comparative overview of identified T1R and T2R taste receptor genes in diverse fish species, highlighting variation in gene number in relation to feeding habits, and referencing the original studies is presented.
GPCRs are expressed not only in the oral cavity but also throughout the gastrointestinal tract, allowing fish to detect nutrients and secondary metabolites during both ingestion and digestion (Oike et al., 2007; Angotzi et al., 2020; Angotzi et al., 2022). TAS1Rs function as heterodimers, TAS1R1/TAS1R3 (umami) and TAS1R2/TAS1R3 (sweet taste in mammals), and are responsive to amino acids, such as L-glutamate and certain sugars. Their activation enhances the palatability of protein-rich diets and stimulates feed intake (Ishimaru et al., 2005; Calo et al., 2024; Miao et al., 2025). These receptors (tas1rs and tas2rs) initiate signaling cascades involving G proteins, phospholipases, and ion channels (Fig. 1), and their sensitivity can be influenced by dietary composition, ultimately affecting feeding behavior. For example, certain amino acids activate the TAS1R1/TAS1R3 complex in Chinese perch (Siniperca chuatsi), triggering the target of rapamycin pathway and neuropeptide Y, which are key appetite regulators (Wang et al., 2022).
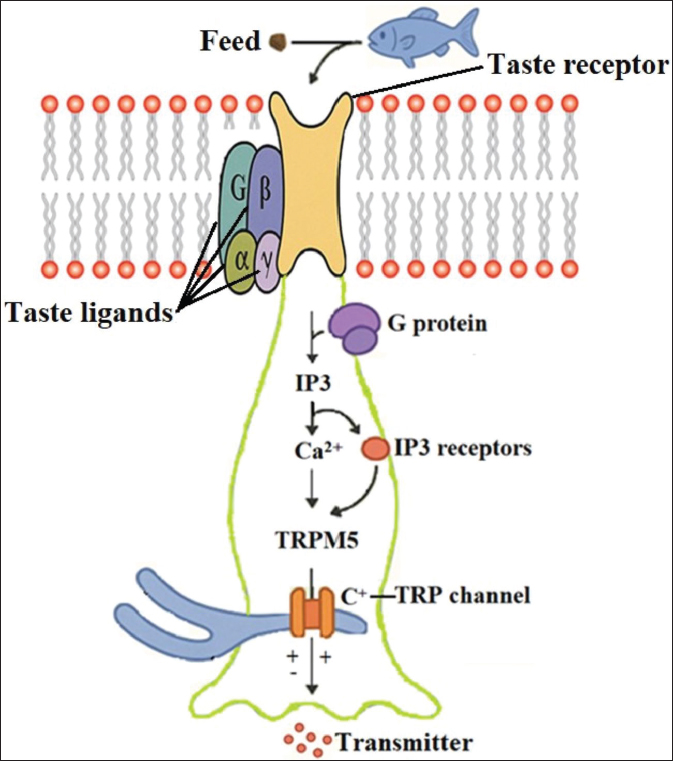
Fig. 1. Schematic representation of the taste transduction pathway in fish (adapted from Roper, 2014). Feed-derived taste ligands bind to membrane-bound taste receptors, activating associated G-protein subunits (G, α, β, γ). This triggers the production of inositol trisphosphate (IP3), leading to Ca2+ release via IP3 receptors. Elevated intracellular Ca2+ activates the TRPM5 channel, resulting in cation influx through the TRP channels and ultimately promoting the release of neurotransmitters to convey taste signals.
Studies in fish species, such as grass carp, zebrafish, and medaka, have shown species-specific responses of tas1r1/tas1r3 to amino acids, such as L-arginine, L-lysine, and L-proline, which influence dietary selection (Cai et al., 2018; Goda et al., 2023). Electrophysiological studies in Nile tilapia (Oreochromis niloticus) have revealed that amino acids such as L-histidine, L-arginine, L-serine, L-methionine, and L-glutamine elicit strong taste responses, whereas L-proline and betaine are less effective (Yacoob et al., 2001).
Some studies have indicated that specific amino acids can negatively affect feed palatability and intake. For example, in rainbow trout (Oncorhynchus mykiss), the oral sensing of L-proline and L-glutamic acid led to feed rejection, with proline having a stronger effect (Comesanã et al., 2020). Factors such as amino acid concentration, exposure duration, and receptor location within the body influence whether T1R-mediated signaling promotes or suppresses feeding.
tas2rs are activated by bitter compounds commonly found in plant-derived ingredients (Krogdahl et al., 2010; Behrens and Meyerhof, 2018; Behrens and Meyerhof, 2011). This activation can result in aversive feeding responses and reduced intake, presenting a significant challenge in developing sustainable aquafeeds with plant-based proteins (Glencross, 2020). Many studies have recommended the incorporation of palatability enhancers, such as amino acids, or bitter-masking agents, such as monosodium glutamate. Palatability enhancers, such as amino acids or bitter-masking agents like monosodium glutamate, have been repeatedly recommended to counteract these effects, improve voluntary intake, and support growth and metabolic performance (Lall and Dumas, 2015; Liu et al., 2025; Salamanca et al., 2025). Understanding the mechanisms by which these receptors are activated is essential for designing diets that either reduce bitterness or strategically modulate receptor signaling.
The tas2r gene encodes bitter taste receptors that detect plant secondary metabolites, a function strongly associated with toxin avoidance (Dong et al., 2009). Teleosts typically possess 3–5 tas2r genes, although some species, including cavefish, exhibit expanded or pseudogenized repertoires exceeding 20 variants, suggesting lineage-specific adaptation (Shiriagin and Korsching, 2019). These receptors are expressed in taste buds located not only in the oral cavity (mouth, tongue, and throat) but also on external body surfaces, where they assist in feed discrimination (Ishimaru et al., 2005; Yasuoka and Abe, 2009; Hardy and Hale, 2022). The tas2rs respond to diverse bitter compounds, including plant secondary metabolites, alkaloids, and antinutritional substances (Gopan et al., 2020; Wooding et al., 2021; Itoigawa et al., 2024) and their activation triggers neural pathways that reduce feed palatability and promote avoidance (Chandrashekar et al., 2000). In species zebrafish and common carp, tannins and saponins reliably elicit these responses through TAS2R activation (Behrens et al., 2021; Shimizu et al., 2021).
The variation in tas2r expression and ligand specificity among fish reflects ecological and evolutionary pressures. Herbivorous and omnivorous species often display reduced sensitivity to bitterness, a pattern consistent with adaptive shifts in gene expression or receptor binding affinity that allow tolerance of plant-rich diets (Kasumyan and Døving, 2003; Shiriagin and Korsching, 2019). Selective pressures from such diets may have driven the expansion and specialization of bitter taste receptors (Li and Zhang, 2014), supporting dietary flexibility in fluctuating environments (Heras et al., 2020).
Sensory dynamics also vary with developmental stage, nutritional state, and environment. Recent studies have proposed strategies to mitigate bitterness through feed additives, such as palatants, amino acids, or sweeteners that mask bitter tastes or modulate tas2r signaling (Shang et al., 2017; Salamanca et al., 2025). Although tas2r activation generally suppresses intake, its effects strongly depend on species-specific physiology and prior exposure to bitter compounds.
Studies have increasingly shown that fish taste relies on more than the canonical TAS1R GPCR pathways. The presence of ASIC and TRP channels, including ASIC4, TRPC2, TRPA1, TRPV1, and TRPV4, within adult taste buds in zebrafish indicates that ionotropic mechanisms contribute to chemical detection (Levanti et al., 2016). Similarly, electrophysiological work in channel catfish demonstrates that L-arginine can be detected through ligand-gated ion channels rather than TAS1R receptors (Grosvenor et al., 2004). Together, these findings imply that fish do not rely solely on TAS1R/TAS2R pathways for amino acid sensing; instead, their taste system integrates multiple receptor types, broadening the range and complexity of cues influencing feeding behavior.
Modulation of the expression of taste receptor genes
Dietary factors affect fish taste receptor genes through various mechanisms, including changes in gene expression, receptor protein activity, and downstream signaling pathways. These mechanisms allow fish to adapt their taste perception to different diets and environments, influencing feed choices and intake (Conde-Sieira and Soengas, 2017; Roy et al., 2022). The expression of taste receptor genes in both oral and extraoral tissues across different fish species has been demonstrated, providing insights into the molecular processes underlying dietary detection and adaptation.
The expression of oral taste receptor genes in fish is essential for detecting and responding to dietary components, especially when shifting from FM diets to plant-based proteins. The oral cavity, which contains taste buds and their associated receptor genes, is the first sensory interface for evaluating feed. This detection allows fish to assess feed palatability and nutritional value before ingestion (Bhatia et al., 2022). This mechanism influences feeding behavior by determining feed acceptance and preference, ultimately affecting the growth and health of aquaculture species. Taste receptor genes expressed in the oral region include receptors sensitive to amino acids, nucleotides, and bitter compounds, corresponding to umami, sweet, and bitter taste modalities, respectively (Zhang et al., 2003). Modulating these genes in response to plant-based diets is particularly important because many plant ingredients contain secondary metabolites and ANFs that can reduce palatability (Hussain et al., 2024). For example, soybean-based proteins commonly used in aquafeeds contain isoflavones and saponins that may activate bitter taste receptors (tas2r genes), potentially decreasing feed intake (Kumar et al., 2020). Conversely, plant proteins contain amino acids that can stimulate umami taste receptor genes (tas1r1 and tas1r3), improve palatability, and promote feeding (Cai et al., 2018).
The link between oral taste receptor gene expression and feed ingredients is reinforced by the differential gene regulation observed in fish fed with plant-based diets compared with those fed with FM diets. Exposure to plant-derived compounds can upregulate or downregulate specific taste receptor genes, indicating an adaptive response to novel dietary components (Balasubramanian et al., 2016). This plasticity in the gustatory system allows fish to adjust their taste perception based on feed chemistry. For example, increased expression of bitter receptor genes may signal increased sensitivity to deterrent compounds, leading to behavioral aversion or reduced consumption (Wang et al., 2022).
The dietary response of taste receptor genes to plant-based diets has been investigated in obscure pufferfish (Takifugu fasciatus), where it was found that increased dietary soybean antigenic proteins increased t1r1 mRNA expression in the tongue (oral) and intestine (extraoral), suggesting a link between taste receptor regulation and dietary composition (Yao et al., 2019). Baranek et al. (2024) studied the effects of prolonged plant-based feeding on the detection of fat in rainbow trout (Oncorhynchus mykiss). They reported that such diets alter (upregulate) the expression of free fatty acid receptor paralogues (ffar1 and ffar2) in the tongue, disrupt calcium signaling, and reduce serotonin levels, impairing nutrient detection without affecting hypothalamic neuropeptide expression. These findings highlight impaired hedonic sensation as a key factor in reduced feed efficiency and growth.
Conversely, the upregulation of umami and sweet receptors may reflect adaptation or improved acceptance due to the presence of palatable amino acids and sugars from plant ingredients. For example, a study on grass carp (Ctenopharyngodon idellus) compared individuals fed chironomid larvae as fish without a dietary transition to those fed duckweed (Lemna minor) following a transition to herbivory. The authors reported a significant upregulation of gcT1R2C, gcT1R2E, and gcT1R2F mRNA expression in the tongue, as well as gcT1R2C and gcT1R2E expression in the gut of duckweed-fed fish. The upregulation of tas1r2 during the dietary shift to plant material suggests that gene expansion and the selective expression of sweet taste receptors facilitate adaptation to herbivorous diets (Yuan et al., 2020).
Taste receptor genes in fish are not limited to the oral cavity but are also expressed in the intestine, brain, and liver, where they contribute to nutrient sensing and digestive regulation. For example, a study by Balasubramanian et al. (2016) explored the molecular mechanisms behind nutritional programming in rainbow trout (Oncorhynchus mykiss) to increase the acceptance of plant-based diets. Transcriptomic analysis revealed that early dietary exposure influenced the expression of genes related to sensory perception, cognition, and neuroendocrine regulation in the brain and affected intermediary metabolism, xenobiotic metabolism, and proteolysis in the liver. Another study in rainbow trout fed a blend of soybean meal, extruded peas, corn gluten meal, and whole wheat revealed developmental changes in gastrointestinal nutrient sensor gene expression. The expression of amino acid and fatty acid transporters, such as slc15a1 and fatp4, increased with age, whereas the expression of other transporters, such as lat4, decreased. Some receptors, such as tas1r2a, are diet-responsive; however, overall fatty acid receptor expression is age-dependent and not significantly affected by diet (Fernández-Maestú et al., 2025). These findings suggest the importance of age and early diet in shaping physiological and metabolic responses to plant-based diets.
Intestinal taste receptors detect chemical cues from digested feed, influencing gut motility, hormone secretion, and nutrient absorption. This gut chemosensory system forms part of a nutrient feedback loop that affects feeding behavior and metabolism (Conde-Sieira and Soengas, 2017; Behrens and Meyerhof, 2019). The expression patterns in the intestine often parallel those in the mouth, adapting according to the diet composition. For example, plant-based diets enriched with certain amino acids or carbohydrates can induce the expression of umami and sweet receptors in the gut, signaling nutrient availability and optimizing digestion (Calo et al., 2021; Chen et al., 2022). Amino acids, such as L-glutamate, and nucleotides, such as inosine monophosphate, are known attractants that stimulate umami receptor genes (tas1r1 and tas1r3) in fish. These compounds activate sensory pathways linked to reward and ingestion, thereby increasing feed intake and conversion efficiency (Hua et al., 2024; Roda et al., 2024; Belloir et al., 2025). Because plant-based diets often lack the attractants found in FM, this deficit may partly explain the lower feed intake of species that are exclusively fed with plant proteins (Liu et al., 2020).
However, a study conducted by Comesanã et al. (2020) showed that the oral detection of amino acids, especially L-proline, reduces feed intake in rainbow trout by triggering hypothalamic satiety pathways. Proline and glutamic acid downregulated taste signaling genes (plcb2 and trpm5), indicating that preabsorptive sensing relays nutrient signals to the brain, influencing feeding behavior via taste receptor pathways. These findings demonstrate that certain amino acids and nucleotides, such as L-proline, may increase feed intake by activating umami receptors, whereas others may reduce intake by triggering satiety pathways. These contrasting effects highlight the complexity of taste receptor signaling in fish and emphasize the need to balance attractants in plant-based diets.
Bitter taste receptors (tas2rs), which are traditionally associated with the oral cavity, are also expressed in the gastrointestinal tract. These receptors likely serve as chemosensory sentinels to detect harmful compounds, such as ANFs, which are commonly present in plant-based feeds (Shimizu et al., 2021; Itoigawa et al., 2024). Upon activation by bitter compounds, tas2rs initiates intracellular signaling cascades involving a gustducin-independent pathway, leading to the secretion of gut hormones such as cholecystokinin and glucagon-like peptide-1. These hormones modulate gastrointestinal motility and promote satiety by acting on the hypothalamus and vagus nerve, ultimately reducing feed intake (Jeon et al., 2011). Therefore, tas2rs in the fish gut not only function as taste receptors but also participate in nutrient sensing, appetite regulation, and immune-metabolic homeostasis, especially under plant-based dietary regimes rich in bitter compounds.
Implications for aquaculture
Integrating taste gene expression into selective breeding to improve feed efficiency
Selective breeding is a powerful tool for improving the use of plant-based dietary formulations in aquaculture. Desirable traits can be transmitted across generations by selecting fish that demonstrate superior growth, feed conversion efficiency, and overall health on plant-based diets, progressively increasing the population’s capacity to use these feeds effectively (Thodesen et al., 2011; Gjedrem et al., 2012). However, feed efficiency is influenced not only by the quality and type of plant ingredients but also by species-specific genetics, physiology, and behavior (Stone et al., 2024). For example, when tilapia are fed different seaweed species and soybean types, the average feed conversion rate (FCR) ranges from 1.1 to 3.0 and from 1.0 to 6.0, respectively. In other species, such as sea bream, the FCR ranges from 1.3 to 2.3; in catfish, it ranges from 1.0 to 3.0; in rainbow trout, it ranges from 0.7 to 2.0; in salmon, it ranges from 0.7 to 2.7; and in carp, it ranges from 1.1 to 3.5. Quantitative genetic studies have consistently reported moderate to high heritability for this trait, indicating a strong potential for genetic improvement (Kause et al., 2006; de Verdal et al., 2017; de Verdal et al., 2018). For example, de Verdal et al. (2017) estimated heritability values for FCR in Nile tilapia ranging from 0.15 to 0.39. Similar genotype diet interactions have been documented in other species, including rainbow trout and gilthead sea bream (Bestin et al., 2014).
Breeding programs targeting growth and feed intake under FM-based diets have shown positive carry-over effects when fish are switched to plant-based feeds. However, direct selection under plant-based diet conditions accelerates adaptation. For example, a selectively bred rainbow trout line thrived on a soybean-based diet without developing enteritis, a condition frequently observed in control lines, emphasizing the effectiveness of targeted selective breeding (Abernathy et al., 2019). Furthermore, salmonid trials comparing soybean meal and FM diets revealed that feed efficiency and daily gain remain strongly genetically correlated across diet types, with cross-diet correlations approaching unity. Additionally, selection for daily gain and daily feed intake on FM diets produced favorable responses in fish subsequently fed with soybean meal diets (Quinton et al., 2007). These findings suggest that selection for feed efficiency via FM diets can improve the performance of plant-based formulations, whereas direct selection under plant-based conditions may further accelerate adaptation.
Incorporating feed efficiency into breeding programs remains challenging because of the difficulty and cost of accurately measuring individual feed intake in aquatic environments. Molecular tools, such as marker-assisted selection and genomic selection, offer promising solutions by leveraging candidate genes associated with feed efficiency. Taste receptor genes have recently emerged as key targets.
Selective breeding for feed efficiency under plant-based diet conditions may not only improve growth and metabolic performance but also indirectly favor fish with increased sensory tolerance and plant-based ingredient acceptance. For example, transcriptomic analyses of isogenic trout lines revealed differential expression of genes related to sensory perception and feeding regulation, including tas1r2, which interacts with metabolic and immune pathways (Balasubramanian et al., 2016). These findings highlight the potential of integrating molecular genetics with selective breeding to develop strains that can efficiently utilize alternative feed ingredients.
Early exposure of fish fry to plant-based feeds can reprogram sensory pathways, later improving feed acceptance. When combined with selective breeding for fish with superior plant diet adaptation, the effects are synergetic: nutritional programming increases sensory gene expression, whereas genetics selects for optimal sensory and metabolic traits. This integration has produced fish lines with improved behavioral acceptance and efficient plant-based feed metabolism over generations. For example, selectively bred lines resulted in significantly greater voluntary intake of plant diets than unselected lines, whereas intake of commercial feeds remained unchanged, confirming diet-specific behavioral adaptation (Roques et al., 2023). Similarly, early exposure of rainbow trout (Oncorhynchus mykiss) to plant-based diets induced long-term upregulation of the taste receptor tas1r2 and other sensory-related genes (olfm, espn), correlating with improved acceptance of plant-rich feeds later in life (Balasubramanian et al., 2016). Domestication and long-term use of plant-based feeds also influence the expression of olfactory receptor genes. Multigenerational feed training in largemouth bass, for example, revealed selection signatures in genes linked to olfactory function, indicating evolutionary adaptation to artificial diets (Zhu et al., 2024).
Molecular and transcriptomic studies further highlight how genetic variation underlies adaptation to plant-based diets. Families better adapted to plant-based feeds presented more favorable hepatic gene expression profiles, particularly in pathways related to lipid, amino acid, and carbohydrate metabolism, immune response, and RNA processing (Geay et al., 2011). In grass carp (Ctenopharyngodon idellus), the evolutionary transition from carnivory to herbivory was associated with an expansion of sweet taste receptor genes (t1r2), which are expressed in taste organs and are responsive to sugars and amino acids, suggesting that gene duplication and elevated expression facilitated adaptation to plant-rich diets (Yuan et al., 2020). Similarly, in Atlantic salmon, plant protein diets change gene expression in the mid-intestine, liver, and muscle, impacting immune pathways, metabolism, and protein turnover; these diets even cause widespread transcriptomic changes and mild enteritis, whereas mixed plant ingredients help reduce these effects (Tacchi et al., 2012; Król et al., 2016). Although these studies did not directly examine taste receptor genes, diet-driven transcriptomic responses likely influence feed intake and palatability.
Collectively, these findings support a coherent model: (1) selective breeding improves the feed efficiency of both FM- and plant-based diets; (2) plant-based diets induce transcriptomic changes, including changes in sensory-related genes; (3) sensory genes play a role in diet adaptation through evolutionary and developmental mechanisms, such as t1r2 expansion in grass carp and nutritional programming in trout; and (4) reduced phenotypic variability in selectively bred lines implies underlying sensory pathway optimization. Collectively, these results suggest that selective breeding for feed efficiency indirectly promotes favorable sensory gene expression patterns, thereby enhancing the acceptance and use of plant-based diets in aquaculture species.
Progress, knowledge gaps, and future research directions
Recent studies on fish taste receptors have employed a wide range of complementary approaches, including comparative genomics and phylogenetics to map receptor repertoires across lineages (Angotzi et al., 2020); transcriptomics and ultrastructural analyses to examine tissue- and diet-specific expression; and functional studies, such as ligand characterization (Oike et al., 2007; He et al., 2021), CRISPR knockouts (e.g., zebrafish t1r1) (Cai et al., 2021), and behavioral trials linking receptor activity to physiological outcomes (Oike et al., 2007). Epigenetic analyses, particularly DNA methylation, and dietary manipulations with plant-based diets (Dhanasiri et al., 2020) or taste stimulants further expand the methodological toolkit, bridging evolutionary surveys with mechanistic insights.
A consensus is emerging that taste receptors have functions beyond the oral cavity, contributing to nutrient sensing and gut–brain signaling. However, contradictions persist, as expansions of bitter (Tas2R) genes in cavefish (Shiriagin et al., 2019) and sweet receptors in grass carp (Yuan et al., 2020) suggest adaptive roles, but functional confirmation is limited. Similarly, although extraoral Tas1Rs and Tas2Rs appear to influence feeding and digestion, most reviewed studies only report correlations. However, recent advances are beginning to fill these gaps. For example, zebrafish Tas1R1 knockouts (Cai et al., 2021) provide causal evidence of receptor roles in feeding regulation. Research on basal lineages, such as coelacanths (Behrens et al., 2021), has clarified the evolution of bitter receptors, while feeding trials using umami additives offer strategies to improve the acceptance of plant-based diets. Additionally, Dhanasiri et al. (2020) discussed epigenetic regulation, and Calo et al. (2021) discussed gastrointestinal amino acid sensing as promising areas of focus.
The strengths of the reviewed studies include their multidisciplinary approach, ecological framing, and relevance to aquaculture. Their weaknesses include reliance on bulk expression data, incomplete ligand–receptor mapping, inconsistent experimental designs, and limited phylogenetic coverage. Future research should include studies on microbiome–receptor interactions, single-cell mapping of gut taste cells, in vivo ligand dose–response dynamics, posttranslational regulation, population-level genetic variation, and organoid models for gut–brain signaling. Progress requires moving beyond correlative expression studies toward mechanistic and integrative analyses.
These insights support the strategic use of plant-based ingredients in aquaculture, considering species-specific taste and feeding adaptations. Early dietary exposure can increase acceptance and growth, whereas selective breeding, nutritional programming, and optimized feed formulation to improve palatability and digestibility can maximize feed efficiency and support sustainable aquaculture practices.
Conclusion
Research on replacing fish meal and fish oil with plant-based diets in aquaculture shows that the interaction between sensory perception, taste receptor function, and feeding behavior shapes dietary adaptation. Across species, adaptation involves both physiological and molecular mechanisms, including changes in taste receptor expression, genetic predisposition, and developmental plasticity, which together determine tolerance to plant-derived compounds. While some fish readily adjust through innate or induced modifications, others require targeted dietary formulations to increase palatability and digestibility. Early dietary exposure, selective breeding, and nutritional programming have emerged as key strategies for promoting plant-based ingredient acceptance and improving feed efficiency. In summary, the central role of taste receptors in both oral and intestinal tissues highlights their dual function in mediating nutrient detection and appetite regulation. Future research should link taste receptor expression to behavior and physiology through dietary, genetic, and neurophysiological approaches to clarify how diet, genetics, and neural responses shape feed acceptance and growth.
Acknowledgments
This study was funded by the Partnership for Applied Skills in Sciences, Engineering, and Technology-Regional Scholarship and Innovation Fund (PASET-Rsif) and the Carnegie Corporation of New York, which awarded funding to LAV (no B8501G30223) for Ph.D. studies at the SACIDS Africa Center of Excellence for Infectious Diseases, the SACIDS Foundation for One Health, Sokoine University of Agriculture, Morogoro, Tanzania. The funders played no role in the study’s design, data collection and analysis, decision to publish, or preparation of the manuscript. The findings and conclusions of this study are solely those of the authors and do not necessarily reflect the views of the funders.
Conflict of interest
The authors have no competing interests to declare.
Funding
None.
Authors' contributions
Leonildo dos Anjo Viagem: conceptualization and planning of the review article, collection and analysis of relevant publications, funding acquisition; writing, original draft, and review and editing; Jean Nepomuscene Hakizimana: writing, review, and editing; and Cyrus Rumisha, Brunno da Silva Cerozi, and Gerald Misinzo: supervision and writing, review, and editing.
Data availability
Data availability does not apply to this manuscript because no new data were generated or analyzed in this study.
References
Abernathy, J., Brezas, A., Snekvik, K.R., Hardy, R.W. and Overturf, K. 2019. Integrative functional analyses using rainbow trout selected for tolerance to plant diets reveal nutrigenomic signatures for soy utilization without the concurrence of enteritis. PLos One. 12(7), 1–30; doi:10.1371/journal.pone.0180972
Angotzi, A.R., Leal, E., Puchol, S., Cerdá-Reverter, J.M. and Morais, S. 2022. Exploring the potential for an evolutionarily conserved role of the taste 1 receptor gene family in gut sensing mechanisms of fish. Anim. Nutr. 11, 293–308; doi:10.1016/j.aninu.2022.08.010
Angotzi, A.R., Puchol, S., Cerdá-Reverter, J.M. and Morais, S. 2020. Insights into the function and evolution of taste 1 receptor gene family in the carnivore fish gilthead seabream (Sparus aurata). Int. J. Mol. Sci. 21(20), 1–20; doi:10.3390/ijms21207732
Balasubramanian, M.N., Panserat, S., Dupont-Nivet, M., Quillet, E., Montfort, J., Le Cam, A., Medale, F., Kaushik, S.J. and Geurden, I. 2016. Molecular pathways associated with the nutritional programming of plant-based diet acceptance in rainbow trout following an early feeding exposure. BMC. Genomics 17(1), 1–20; doi:10.1186/s12864-016-2804-1
Baranek, E., Heraud, C., Larroquet, L., Surget, A., Lanuque, A., Terrier, F., Skiba-Cassy, S. and Jérôme, R. 2024. Long-term regulation of fat sensing in rainbow trout (Oncorhynchus mykiss) fed a vegetable diet from the first feeding: focus on free fatty acid receptors and their signalling. Br. J. Nutr. 131(1), 1–16; doi:10.1017/S0007114523001599
Behrens, M. and Meyerhof, W. 2011. Gustatory and extragustatory functions of mammalian taste receptors. Physiol. Behav. 105(1), 4–13; doi:10.1016/j.physbeh.2011.02.010
Behrens, M. and Meyerhof, W. 2018. Vertebrate Bitter Taste Receptors: keys for Survival in Changing Environments. J. Agricult. Food. Chem. 66(10), 2204–2213; doi:10.1021/acs.jafc.6b04835
Behrens, M. and Meyerhof, W. 2019. A role for taste receptors in (neuro)endocrinology?. J. Neuroendocrinology 31(3), 1–12; doi:10.1111/jne.12691
Behrens, M., Di Pizio, A., Redel, U., Meyerhof, W. and Korsching, S.I. 2021. At the Root of T2R Gene Evolution: recognition Profiles of Coelacanth and Zebrafish Bitter Receptors. Genome Biol. Evol. 13(1), 1–17; doi:10.1093/gbe/evaa264
Belloir, C., Moitrier, L., Karolkowski, A., Poirier, N., Neiers, F. and Briand, L. 2025. Inosine-5′-monophosphate interacts with the TAS1R3 subunit to enhance sweet taste detection. Food Chem. Mol. Sci. 10(November 2024), S1–12; doi:10.1016/j.fochms.2025.100246
Benhaïm, D., Bégout, M.L., Péan, S., Manca, M., Prunet, P. and Chatain, B. 2013. Impact of a plant-based diet on behavioural and physiological traits in sea bass (Dicentrarchus labrax). Aquatic Living Resour. 26(2), 121–131; doi:10.1051/alr/2013049
Bestin, A., Dupont Nivet, M., Haffray, P., Médale, F., Quillet, E., Vandeputte, M., Cariou, S., Desgranges, A., Laureau, S. and Ricoux, R. Genotype by diet interactions on growth and processing traits in rainbow trout (Oncorhynchus mykiss), European sea bass (Dicentrarchus labrax), gilthead sea bream (Sparus aurata) and meagre (Argyrosomus regius) fed diets with almost complete substitution. Proceedings of the 10th World Congress on Genetics Applied to Livestock Production (WCGALP), 2014, 17–22, Vancouver, Canada.
Bhatia, V., De Jesus, V.C., Shaik, F.A., Jaggupilli, A., Singh, N., Chelikani, P. and Atukorallaya, D. 2022. Extraoral expression and characterization of bitter taste receptors in Astyanax mexicanus (Mexican tetra fish). FASEB. BioAdvances. 4(9), 574–584; doi:10.1096/fba.2022-00032
Booman, M., Forster, I., Vederas, J.C., Groman, D.B. and Jones, S.R.M. 2018. Soybean meal-induced enteritis in Atlantic salmon (Salmo salar) and Chinook salmon (Oncorhynchus tshawytscha) but not in pink salmon (O. gorbuscha). Aquaculture 483, 238–243; doi:10.1016/j.aquaculture.2017.10.025
Boudriot, F. and Reutter, K. 2001. Ultrastructure of the taste buds in the blind cave fish Astyanax jordani (‘Anoptichthys’) and the sighted river fish Astyanax mexicanus (Teleostei, Characidae). J. Comparative Neurol. 434(4), 428–444; doi:10.1002/cne.1185
Cai, W., He, S., Liang, X.F. and Yuan, X. 2018. DNA Methylation of T1R1 Gene in the Vegetarian Adaptation of Grass Carp Ctenopharyngodon idella. Scientific. Rep. 8(1), T1–15; doi:10.1038/s41598-018-25121-4
Cai, W.J., Li, J., Li, L., Chen, X., Wei, J.R., Yin, Z., He, S. and Liang, X.F. 2021. Knockout of t1r1 gene in zebrafish (Danio rerio) by CRISPR/Cas9 reveals its roles in regulating feeding behavior. Aquaculture 545, t1–t8; doi:10.1016/j.aquaculture.2021.737189
Calo, J., Blanco, A.M., Comesaña, S., Conde-Sieira, M., Morais, S. and Soengas, J.L. 2021. First evidence for the presence of amino acid sensing mechanisms in the fish gastrointestinal tract. Scientific Rep. 11(1), 1–14; doi:10.1038/s41598-021-84303-9
Calo, J., Comesaña, S., Fernández-Maestú, C., Blanco, A.M., Morais, S. and Soengas, J.L. 2024. Impact of feeding diets with enhanced vegetable protein content and presence of umami taste-stimulating additive on gastrointestinal amino acid sensing and feed intake regulation in rainbow trout. Aquaculture 579, 1–17; doi:10.1016/j.aquaculture.2023.740251
Chandrashekar, J., Mueller, K.L., Hoon, M.A., Adler, E., Feng, L., Guo, W., Zuker, C.S. and Ryba, N.J.P. 2000. T2Rs function as bitter taste receptors. Cell 100(6), 703–711; doi:10.1016/S0092-8674(00)80706-0
Chen, H.T., Li, L.L., Wang, L.H., Cheng, D.H., Ma, H., Sun, M.J., Yang, Y.O. and Yuan, X.C. 2022. Sweet taste receptors are the potential mediator involved in appetite regulation of grass carp in response to high digestible carbohydrates intake. Aquac. Rep. 27, 1–11; doi:10.1016/j.aqrep.2022.101386
Colombo, S.M. 2020. Physiological considerations in shifting carnivorous fishes to plant-based diets. In Fish Physiology: Aquaculture. Eds., Benfey, T.J., Farrell, A.P. and Brauner, C.J. Amsterdam, The Netherlands: Elsevier Inc, Vol. 38, pp: 53–82; doi: 10.1016/bs.fp.2020.09.002
Comesaña, S., Conde-Sieira, M., Velasco, C., Soengas, J.L. and Morais, S. 2020. Oral and pre-absorptive sensing of amino acids relates to hypothalamic control of food intake in rainbow trout. J. Exp. Biol. 223(17), 1–15; doi:10.1242/jeb.221721
Conde-Sieira, M. and Soengas, J.L. 2017. Nutrient sensing systems in fish: impact on food intake regulation and energy homeostasis. Front. Neurosci. 10, 1–21; doi:10.3389/fnins.2016.00603
De Francesco, M., Parisi, G., Pérez-Sánchez, J., Gómez-Réqueni, P., Médale, F., Kaushik, S.J., Mecatti, M. and Poli, B.M. 2007. Effect of high-level fish meal replacement by plant proteins in gilthead sea bream (Sparus aurata) on growth and body/fillet quality traits. Aquac. Nutr. 13(5), 361–372; doi:10.1111/j.1365-2095.2007.00485.x
De Verdal, H., Komen, H., Quillet, E., Chatain, B., Allal, F., Benzie, J.A.H. and Vandeputte, M. 2018. Improving feed efficiency in fish using selective breeding: a review. Rev. Aquac. 10(4), 833–851; doi:10.1111/raq.12202
De Verdal, H., Mekkawy, W., Lind, C.E., Vandeputte, M., Chatain, B. and Benzie, J.A.H. 2017. Measuring individual feed efficiency and its correlations with performance traits in Nile tilapia, Oreochromis niloticus. Aquaculture 468, 489–495; doi:10.1016/j.aquaculture.2016.11.015
Dhanasiri, A.K.S., Johny, A., Xue, X., Berge, G.M., Bogevik, A.S., Rise, M.L., Fæste, C.K. and Fernandes, J.M.O. 2020. Plant-Based Diets Induce Transcriptomic Changes in Muscle of Zebrafish and Atlantic Salmon. Front. Genet. 11, 1–15; doi:10.3389/fgene.2020.575237
Dong, D., Jones, G. and Zhang, S. 2009. Dynamic evolution of bitter taste receptor genes in vertebrates. BMC. Evol. Biol. 9(1), 1–9; doi:10.1186/1471-2148-9-12
El-Sayed, A.F.M., Fagnon, M.S., Hamdan, A.M., Chabrillat, T., Kerros, S. and Zeid, S.M.S. 2025. Dietary Plant-Based Mixture Improves Feed Efficiency, Gross Profit, Physiological Performance, Gene Expression and Gut Health of Nile Tilapia (Oreochromis niloticus). Biology 14(2), 1–22; doi:10.3390/biology14020186
Fernández-Maestú, C., Calo, J., Martinat, M., Soengas, J.L., Roy, J. and Blanco, A.M. 2025. Effects of a plant-based diet from first feeding on the intestinal expression of nutrient sensors in rainbow trout (Oncorhynchus mykiss). Aquaculture, 599(August 2024), 742093; doi: 10.1016/j.aquaculture.2024.742093
Fiocchi, E., Civettini, M., Carbonara, P., Zupa, W., Lembo, G. and Manfrin, A. 2020. Development of molecular and histological methods to evaluate stress oxidative biomarkers in sea bass (Dicentrarchus labrax). Fish Physiol. Biochem. 46(4), 1577–1588; doi:10.1007/s10695-020-00811-x
Firmino, J.P., Galindo-Villegas, J., Reyes-López, F.E. and Gisbert, E. 2021. Phytogenic bioactive compounds shape fish mucosal immunity. Front. Immunol. 12, 1–20. doi: 10.3389/fimmu.2021.695973
Francis, G., Makkar, H.P.S. and Becker, K. 2001. Antinutritional factors present in plant-derived alternate fish feed ingredients and their effects in fish. Aquaculture 199(3–4), 3; doi:10.1016/S0044-8486(01)00526-9
Gao, S., Liu, S., Yao, J., Zhou, T., Li, N., Li, Q., Dunham, R. and Liu, Z. 2017. Taste receptors and gustatory associated G proteins in channel catfish, Ictalurus punctatus. Comparative Biochem. Physiol. - Part D. Genomics Proteomics 21, 1–9; doi:10.1016/j.cbd.2016.10.002
Gatlin, D.M., Barrows, F.T., Brown, P., Dabrowski, K., Gaylord, T.G., Hardy, R.W., Herman, E., Hu, G., Krogdahl, A., Nelson, R., Overturf, K., Rust, M., Sealey, W., Skonberg, D., J Souza, E., Stone, D., Wilson, R. and Wurtele, E. 2007. Expanding the utilization of sustainable plant products in aquafeeds: a review. Aquac. Res. 38(6), 551–579; doi:10.1111/j.1365-2109.2007.01704.x
Geay, F., Ferraresso, S., Zambonino Infante, J.L., Bargelloni, L., Quentel, C., Vandeputte, M., Kaushik, S., Cahu, C.L. and Mazurais, D. 2011. Effects of the total replacement of fish meal and fish oil by plant sources on the hepatic transcriptome of European sea bass (Dicentrarchus labrax). BMC. Genomics. 12(522), 1–8.
Gjedrem, T., Robinson, N. and Rye, M. 2012. The importance of selective breeding in aquaculture to meet future demands for animal protein: a review. Aquaculture 350–353, 117–129; doi:10.1016/j.aquaculture.2012.04.008
Glencross, B.D. 2020. A feed is still only as good as its ingredients: an update on the nutritional research strategies for the optimal evaluation of ingredients for aquaculture feeds. Aquac. Nutr. 26(6), 1871–1883; doi:10.1111/anu.13138
Goda, R., Watanabe, S. and Misaka, T. 2023. Zebrafish and medaka T1R (taste receptor type 1) proteins mediate highly sensitive recognition of l-proline. FEBS. Open. Bio. 13(3), 468–477; doi:10.1002/2211-5463.13558
Gopan, A., Lalappan, S., Varghese, T., Kumar Maiti, M. and Peter, R.M. 2020. Anti-Nutritional Factors in Plant-Based Aquafeed Ingredients: effects on Fish and Amelioration Strategies. Bioscience. Biotechnol. Res. Commun. Special. Issue. 13(12), 1–9.
Grosvenor, W., Kaulin, Y., Spielman, A.I., Bayley, D.L., Kalinoski, D.L., Teeter, J.H. and Brand, J.G. 2004. Biochemical enrichment and biophysical characterization of a taste receptor for L-arginine from the catfish, Ictalurus puntatus. BMC. Neurosci. 5, 1–19; doi:10.1186/1471-2202-5-25
Hardy, A.R. and Hale, M.E. 2022. Extraoral Taste Buds on the Paired Fins of Damselfishes. Integr. Organismal Biol. 4(1), 643–708; doi:10.1093/iob/obac035
He, S., You, J.J., Liang, X.F., Zhang, Z.L. and Zhang, Y.P. 2021. Transcriptome sequencing and metabolome analysis of food habits domestication from live prey fish to artificial diets in mandarin fish (Siniperca chuatsi). BMC Genom. 22(1), 1–12; doi: 10.1186/s12864-021-07403-w
Hedrera, M.I., Galdames, J.A., Jimenez-Reyes, M.F., Reyes, A.E., Avendaño-Herrera, R., Romero, J. and Feijóo, C.G. 2013. Soybean Meal Induces Intestinal Inflammation in Zebrafish Larvae. PLos One. 8(7), 1–10; doi:10.1371/journal.pone.0069983
Heras, J., Chakraborty, M., Emerson, J.J. and German, D.P. 2020. Genomic and biochemical evidence of dietary adaptation in a marine herbivorous fish. Proc. Roy. Soc. B. Biol. Sci. 287(1921), 1–9; doi:10.1098/rspb.2019.2327
Heraud, C., Hirschinger, T., Baranek, E., Larroquet, L., Surget, A., Sandres, F., Lanuque, A., Terrier, F. and Roy, J. 2022. Detection and Modulation of Olfactory Sensing Receptors in Carnivorous Rainbow Trout (Oncorhynchus mykiss) Fed from First Feeding with Plant-Based Diet. Int. J. Mol. Sci. 23(4), 1–18; doi:10.3390/ijms23042123
Hu, H., Kortner, T.M., Gajardo, K., Chikwati, E., Tinsley, J. and Krogdahl, A. 2016. Intestinal fluid permeability in Atlantic salmon (Salmo salar L.) is affected by dietary protein source. PLos One 11(12), 1–18; doi:10.1371/journal.pone.0167515
Hua, L., Zhang, P., Liu, H., Xin, M., Zhang, Z., Han, D., Zhang, Z., Zhu, X., Jin, J., Yang, Y. and Xie, S. 2024. Effects of Dietary Inosine 5′-Monophosphate Supplementation on the Growth Performance and Salinity and Oxidative Stress Resistance of Gibel Carp (Carassius auratus gibelio). Antioxidants 13(4), 1–15; doi:10.3390/antiox13040487
Hussain, S.M., Bano, A.A., Ali, S., Rizwan, M., Adrees, M., Zahoor, A.F., Sarker, P.K., Hussain, M., Arsalan, M.Z.U.H., Yong, J.W.H. and Naeem, A. 2024. Substitution of fishmeal: highlights of potential plant protein sources for aquaculture sustainability. Heliyon 10(4), 1–29; doi:10.1016/j.heliyon.2024.e26573
Ikenaga, T., Nakamura, T., Tajiri, T., Tsuji, M., Kato, D.I., Ineno, T., Kobayashi, Y., Tsutsui, N. and Kiyohara, S. 2023. Diversity and evolution of serotonergic cells in taste buds of elasmobranchs and ancestral actinopterygian fish. Cell. Tissue. Res. 394(3), 431–439; doi:10.1007/s00441-023-03837-8
Ishimaru, Y., Okada, S., Naito, H., Nagai, T., Yasuoka, A., Matsumoto, I. and Abe, K. 2005. Two families of candidate taste receptors in fishes. Mech. Develop. 122(12), 1310–1321; doi:10.1016/j.mod.2005.07.005
Itoigawa, A., Nakagita, T. and Toda, Y. 2024. The Remarkable Diversity of Vertebrate Bitter Taste Receptors: recent Advances in Genomic and Functional Studies. Int. J. Mol. Sci. 25(23), 1–23; doi:10.3390/ijms252312654
Iwama, G.K., Thomas, P.T., Forsyth, R.B. and Vijayan, M.M. 1998. Heat shock protein expression in fish. Rev. Fish Biol. Fisheries 8(1), 35–56; doi:10.1023/A:1008812500650
Jayant, M., Muralidhar, A.P., Sahu, N.P., Jain, K.K., Pal, A.K. and Srivastava, P.P. 2018. Protein requirement of juvenile striped catfish, Pangasianodon hypophthalmus. Aquac. Int. 26(1), 375–389; doi:10.1007/s10499-017-0216-0
Jeon, T.I., Seo, Y.K. and Osborne, T.F. 2011. Gut bitter taste receptor signalling induces ABCB1 through a mechanism involving CCK. Biochem. J. 438(1), 33–37; doi:10.1042/BJ20110009.Gut
Jiao, F., Zhang, L., Limbu, S.M., Yin, H., Xie, Y., Yang, Z., Shang, Z., Kong, L. and Rong, H. 2023. A comparison of digestive strategies for fishes with different feeding habits: digestive enzyme activities, intestinal morphology, and gut microbiota. Ecol. Evol. 13(9), 1–14; doi:10.1002/ece3.10499
Kari, A.Z., Mat, K., Kabir, M.A., Amir Zal, W.A., Munir, M.B., Seong Wei, L. and Téllez-Isaías, G. 2023. Soybean by-product: as an alternative to fish meal as protein source for aquaculture industry. J. Sustainability. Sci. Manage. 18(5), 179–202; doi: 10.46754/jssm.2023.05.013
Kari, Z.A., Kabir, M.A., Dawood, M.A.O., Razab, M.K.A.A., Ariff, N.S.N.A., Sarkar, T., Pati, S., Edinur, H.A., Mat, K., Ismail, T.A. and Wei, L.S. 2022. Effect of fish meal substitution with fermented soy pulp on growth performance, digestive enzyme, amino acid profile, and immune-related gene expression of African catfish (Clarias gariepinus). Aquaculture 546 (2022) 737418; doi: 10.1016/j.aquaculture.2021.737418
Kasumyan, A.O. and Døving, K.B. 2003. Taste preferences in fishes. Fish. Fisheries. 4(4), 289–347; doi:10.1046/j.1467-2979.2003.00121.x
Kaushik, S.J., Cravedi, J.P., Lalles, J.P., and Sumpter, J., Fauconneau B. and Laroche M. 1995. Partial or total replacement of fish meal by soybean protein on growth, protein utilization, potential estrogenic or antigenic effects, cholesterolemia and flesh quality in rainbow trout, Oncorhynchus mykiss. Aquaculture 133 (1995) 257–274.
Kause, A., Tobin, D., Dobly, A., Houlihan, D., Martin, S., Mäntysaari, E.A., Ritola, O. and Ruohonen, K. 2006. Recording strategies and selection potential of feed intake measured using the X-ray method in rainbow trout. Genet. Selection. Evol. 38(4), 389–409; doi:10.1051/gse:2006011
Kong, S., Dong, C., Lv, H., Chen, L., Zhang, J., Pu, F., Li, X. and Xu, P. 2018. Genome wide identification of taste receptor genes in common carp (Cyprinus carpio) and phylogenetic analysis in teleost. Gene 678, 65–72; doi:10.1016/j.gene.2018.07.078
Krogdahl, A., Penn, M., Thorsen, J., Refstie, S. and Bakke, A.M. 2010. Important antinutrients in plant feedstuffs for aquaculture: an update on recent findings regarding responses in salmonids. Aquac. Res. 41(3), 333–344; doi:10.1111/j.1365-2109.2009.02426.x
Król, E., Douglas, A., Tocher, D.R., Crampton, V.O., Speakman, J.R., Secombes, C.J. and Martin, S.A.M. 2016. Differential responses of the gut transcriptome to plant protein diets in farmed Atlantic salmon. BMC. Genomics 17(1), 1–16; doi:10.1186/s12864-016-2473-0
Kumar, V., Sakhawat Hossain, M., Ragaza, J. and Rubio Benito, M. 2020. The potential impacts of soy protein on fish gut health. In Soybean for Human Consumption and Animal Feed. Ed., Sudarić, A. London, UK: IntechOpen, pp: 91–115; doi: 10.5772/intechopen.92695
Kurniasih, T., Zulkarnain, W., Wijaya, R., Lesmana, D., Mumpuni, F.S., Panigoro, M., Sutisna, E., Heptarina, E., Fitria, Y., Saptana, S. and Ermiati. 2024. Digestibility of plant-based feeds in omnivorous, carnivorous, and herbivorous fish: a review of Nile tilapia (Oreochromis niloticus (Linnaeus, 1758)), North African catfish (Clarias gariepinus (Burchell, 1822)), and grass carp (Ctenopharyngodon idella). AACL. Bioflux. 17(6), 2994–3014.
Lall, S.P. and Dumas, A. 2015. 3—Nutritional requirements of cultured fish: formulating nutritionally adequate feeds. In Woodhead Publishing Series in Food Science, Technology and Nutrition. Eds., Webster, F. and Davis, A. Cambridge, UK: Woodhead Publishing, pp: 53–109; doi: 10.1016/B978-0-08-100506-4.00003-9
Levanti, M., Randazzo, B., Viña, E., Montalbano, G., Garcia-Suarez, O., Germanà, A., Vega, J.A. and Abbate, F. 2016. Acid-sensing ion channels and transient-receptor potential ion channels in zebrafish taste buds. Ann. Anatomy. 207, 32–37; doi:10.1016/j.aanat.2016.06.006
Li, D. and Zhang, J. 2014. Diet shapes the evolution of the vertebrate bitter taste receptor gene repertoire. Mol. Biol. Evol. 31(2), 303–309; doi:10.1093/molbev/mst219
Lima, J.D.C., Schorer, M., Melo, J.F.B. and Braga, L.G.T. 2021. Effect of enzymatic complex in the diet of pirarucu, arapaima gigas juveniles. Acta. Amazonica. 51(3), 207–213; doi:10.1590/1809-4392202003151
Liu, R., Zhang, Y., Liang, X., Lou, B. and Zhu, J. 2025. Effects of Glutamate on Growth Performance, Gut Digestion and Antioxidant Capacity in Juvenile Little Yellow Croaker. Fishes 10(4), 1–19; doi:10.3390/fishes10040188
Liu, X., Han, B., Xu, J., Zhu, J., Hu, J., Wan, W. and Miao, S. 2020. Replacement of fishmeal with soybean meal affects the growth performance, digestive enzymes, intestinal microbiota and immunity of Carassius auratus gibelio♀ × Cyprinus carpio♂. Aquac. Rep. 18, 1–9; doi:10.1016/j.aqrep.2020.100472
Martinat, M., Lasserre, M., Baranek, E., Surget, A., Lanuque, A., Terrier, F., Heraud, C. and Roy, J. 2025. Early sensory responses to plant-based diets in rainbow trout (Oncorhynchus mykiss) alevins: impact on feeding behavior. Aquac. Rep. 43, 1–19; doi:10.1016/j.aqrep.2025.102943
Martinat, M., Varvarais, A., Heraud, C., Surget, A., Lanuque, A., Terrier, F. and Roy, J. 2025. Effects of a Plant-Based Diet During the First Month of Feeding on Alevin Rainbow Trout (Oncorhynchus mykiss) in the Development of Tongue Sensory System Regulating Feeding Behavior. Aquac. Nutr. 2025b(1), 1–23; doi:10.1155/anu/6690967
Miao, R., Gong, Y., Zhang, N., He, M., Li, D., Morais, S., Li, S., Chen, Z., Chen, N. and Li, S. 2025. Effects of dietary umami taste stimulator on appetite, growth, antioxidant capacity, intestinal histology and target of rapamycin pathway related genes’ expression in largemouth bass (Micropterus salmoides) fed a high plant protein diet. Aquaculture Fisheries 11(1), 1–11; doi: 10.1016/j.aaf.2025.02.001
Morais, S. 2017. The Physiology of Taste in Fish: potential Implications for Feeding Stimulation and Gut Chemical Sensing. Rev. Fisheries Sci. Aquac. 25(2), 133–149; doi:10.1080/23308249.2016.1249279
Naylor, R.L., Hardy, R.W., Bureau, D.P., Chiu, A., Elliott, M., Farrell, A.P., Forster, I., Gatlin, D.M., Goldburg, R.J., Hua, K. and Nichols, P.D. 2009. Feeding aquaculture in an era of finite resources. Proc. Nat. Acad. Sci. United States Amer. 106(36), 15103–15110; doi:10.1073/pnas.0905235106
Nhu, T.Q., Bich Hang, B.T., Bach, L.T., Buu Hue, B.T., Quetin-Leclercq, J., Scippo, M.L., Phuong, N.T. and Kestemont, P. 2019. Plant extract-based diets differently modulate immune responses and resistance to bacterial infection in striped catfish (Pangasianodon hypophthalmus). Fish. Shellfish. Immunol. 92, 913–924; doi:10.1016/j.fsi.2019.07.025
Oike, H., Nagai, T., Furuyama, A., Okada, S., Aihara, Y., Ishimaru, Y., Marui, T., Matsumoto, I., Misaka, T. and Abe, K. 2007. Characterization of ligands for fish taste receptors. J. Neurosci. 27(21), 5584–5592; doi:10.1523/JNEUROSCI.0651-07.2007
Pavasovic, A., Richardson, N.A., Mather, P.B. and Anderson, A.J. 2006. Influence of insoluble dietary cellulose on digestive enzyme activity, feed digestibility and survival in the red claw crayfish, Cherax quadricarinatus (von Martens). Aquac. Res. 37(1), 25–32; doi:10.1111/j.1365-2109.2005.01389.x
Podell, S., Oliver, A., Kelly, L.W., Sparagon, W.J., Plominsky, A.M., Nelson, R.S., Laurens, L.M.L., Augyte, S., Sims, N.A., Nelson, C.E. and Allen, E.E. 2023. Herbivorous Fish Microbiome Adaptations to Sulfated Dietary Polysaccharides. Appl. Environ. Microbiol. 89(5), 1–18; doi:10.1128/aem.02154-22
Quinton, C.D., Kause, A., Koskela, J. and Ritola, O. 2007. Breeding salmonids for feed efficiency in current fishmeal and future plant-based diet environments. Genet. Selection Evol. 39(4), 431–446; doi:10.1051/gse:2007013
Reutter, K., Breipohl, W. and Bijvank, G. 1974. Taste bud types in fishes - II. Scanning electron microscopical investigations on Xiphophorus helleri Heckel (Poeciliidae, Cyprinodontiformes, Teleostei). Cell. Tissue. Res. 153(2), 151–165; doi:10.1007/BF00226604
Roda, T.N., Farzad, R. and Allen, M.S. 2024. A Review of Feed Attractants as a Guide for Aquaculture Operations. Edis 6(6), 1–7; doi:10.32473/edis-fs462-2024
Roper, S.D. 2014. TRPs in taste and chemesthesis. In Handbook of Experimental Pharmacology. Eds., Nilius, B and Flockerzi, V. Berlin, Germany: Springer, Vol. 223, pp: 827–871; doi: 10.1007/978 3 319 05161 1_
Roques, S., Deborde, C., Skiba-Cassy, S., Médale, F., Dupont-Nivet, M., Lefevre, F., Bugeon, J., Labbé, L., Marchand, Y., Moing, A. and Fauconneau, B. 2023. New alternative ingredients and genetic selection are the next game changers in rainbow trout nutrition: a metabolomics appraisal. Scientific Rep. 13(1), 1–16; doi:10.1038/s41598-023-46809-2
Roy, J., Baranek, E., Mercier, Y., Larroquet, L., Surget, A., Ganot, A., Sandres, F., Lanuque, A., Terrier, F. and Briand, L. 2022. Involvement of Taste Receptors in the Oro-Sensory Perception of Nutrients in Rainbow Trout (Oncorhynchus Mykiss) Fed Diets with Different Fatty Acid Profiles. Aquac. Nutr. 2022, 1–20; doi:10.1155/2022/1152463
Sadoul, B., Foucard, A., Valotaire, C., Labbé, L., Goardon, L., Lecalvez, J.M., Médale, F., Quillet, E., Dupont-Nivet, M., Geurden, I., Prunet, P. and Colson, V. 2016. Adaptive capacities from survival to stress responses of two isogenic lines of rainbow trout fed a plant-based diet. Scientific Rep. 6, 1–11; doi:10.1038/srep35957
Salamanca, N., Herrera, M. and De La Roca, E. 2025. Amino Acids as Dietary Additives for Enhancing Fish Welfare in Aquaculture. Animals 15(9), 1–15; doi:10.3390/ani15091293
Shang, S., Zhang, H., Wu, X., Chen, J., Zhong, H., Wei, Q., Zhao, C., Yan, J., Chen, Y., Tang, X. and Zhang, H. 2017. The repertoire of bitter taste receptor genes in Ovalentaria fish. Environ. Biol. Fishes 100(11), 1489–1496; doi:10.1007/s10641-017-0659-1
Sharawy, Z., Goda, A.M.A.S. and Hassaan, M.S. 2016. Partial or total replacement of fish meal by solid state fermented soybean meal with Saccharomyces cerevisiae in diets for Indian prawn shrimp, Fenneropenaeus indicus, Postlarvae. Anim. Feed. Sci. Technol. 212, 90–99; doi:10.1016/j.anifeedsci.2015.12.009
Shi, P. and Zhang, J. 2006. Contrasting modes of evolution between vertebrate sweet/umami receptor genes and bitter receptor genes. Mol. Biol. Evol. 23(2), 292–300; doi:10.1093/molbev/msj028
Shimizu, T., Kubozono, T., Asaoka, R., Toda, Y. and Ishimaru, Y. 2021. Expression profiles and functional characterization of common carp (Cyprinus carpio) T2Rs. Biochem. Biophys. Rep. 28, 1–7; doi:10.1016/j.bbrep.2021.101123
Shiriagin, V. and Korsching, S.I. 2019. Massive expansion of bitter taste receptors in blind cavefish, Astyanax mexicanus. Chem. Senses. 44(1), 23–32.
Siddik, M.A.B., Francis, P., Rohani, M.F., Azam, M.S., Mock, T.S. and Francis, D.S. 2023. Seaweed and Seaweed-Based Functional Metabolites as Potential Modulators of Growth, Immune and Antioxidant Responses, and Gut Microbiota in Fish. Antioxidants 12(12), 1–34; doi:10.3390/antiox12122066
Stone, N.M., Engle, C.R., Kumar, G., Li, M.H., Hegde, S., Roy, L.A., Kelly, A.M., Dorman, L. and Recsetar, M.S. 2024. Factors affecting feed conversion ratios in US commercial catfish production ponds. J. World. Aquac. Soc. 55(3), 1–36; doi:10.1111/jwas.13053
Syed, A.S. and Korsching, S.I. 2014. Positive Darwinian selection in the singularly large taste receptor gene family of an ‘ancient’ fish, Latimeria chalumnae. BMC. Genomics. 15(1), 1–15; doi:10.1186/1471-2164-15-650
Tacchi, L., Secombes, C.J., Bickerdike, R., Adler, M.A., Venegas, C., Takle, H. and Martin, S.A. 2012. Transcriptomic and physiological responses to fishmeal substitution with plant proteins in formulated feed in farmed Atlantic salmon (Salmo salar). BMC. Genomics. 13(1), 1–21; doi:10.1186/1471-2164-13-363
Tacon, A.G.J. and Metian, M. 2008. Global overview on the use of fish meal and fish oil in industrially compounded aquafeeds: trends and future prospects. Aquaculture 285(1–4), 146–158; doi:10.1016/j.aquaculture.2008.08.015
Thodesen (da-yong Ma), J., Rye, M., Wang, Y.X., Yang, K.S., Bentsen, H.B. and Gjedrem, T. 2011. Genetic improvement of tilapias in China: genetic parameters and selection responses in growth of Nile tilapia (Oreochromis niloticus) after six generations of multi-trait selection for growth and fillet yield. Aquaculture 322–323, 51–64; doi:10.1016/j.aquaculture.2011.10.010
Valente, L.M.P., Cabral, E.M., Sousa, V., Cunha, L.M. and Fernandes, J.M.O. 2016. Plant protein blends in diets for Senegalese sole affect skeletal muscle growth, flesh texture and the expression of related genes. Aquaculture 453, 77–85; doi:10.1016/j.aquaculture.2015.11.034
Varó, I., Navarro, J.C., Nunes, B. and Guilhermino, L. 2007. Effects of dichlorvos aquaculture treatments on selected biomarkers of gilthead sea bream (Sparus aurata L.) fingerlings. Aquaculture 266(1–4), 87–96; doi:10.1016/j.aquaculture.2007.02.045
Véron, V., Panserat, S., Le Boucher, R., Labbé, L., Quillet, E., Dupont-Nivet, M. and Médale, F. 2016. Long-term feeding a plant-based diet devoid of marine ingredients strongly affects certain key metabolic enzymes in the rainbow trout liver. Fish Physiol. Biochem. 42(2), 771–785; doi:10.1007/s10695-015-0174-2
Wang, L., Hu, S., Lou, B., Chen, D., Zhan, W., Chen, R., Liu, F. and Xu, D. 2018. Effect of Different Dietary Protein and Lipid Levels on the Growth, Body Composition, and Intestinal Digestive Enzyme Activities of Juvenile Yellow Drum Nibea albiflora (Richardson). J. Ocean. Univ. China. 17(5), 1261–1267; doi:10.1007/s11802-018-3660-1
Wang, Q., Liang, X.F., Gao, J., Cai, W., He, S. and Zhuang, W. 2022. Lysine regulates TOR and NPY through taste receptor T1R1 in Chinese perch (Siniperca chuatsi). Aquaculture 559, 738445; doi:10.1016/j.aquaculture.2022.738445
Wooding, S.P., Ramirez, V.A. and Behrens, M. 2021. Bitter taste receptors, Genes, evolution and health. Evol. Med. Public Health 9(1), 431–447; doi:10.1093/emph/eoab031
Yacoob, S.Y., Anraku, K., Marui, T., Matsuoka, T., Kawamura, G. and Archdale, M.V. 2001. Gustatory sensitivity of the external taste buds of Oreochromis niloticus L. to amino acids. Aquac. Res. 32(3), 217–222; doi:10.1046/j.1365-2109.2001.00550.x
Yao, J.T., Kong, C., Hua, X.M., Yang, J.F., Liu, T., Wang, G., Shui, C., Feng, Y. and Shi, Y.H. 2019. T1R1 expression in obscure puffer (Takifugu fasciatus) is associated with effect of dietary soybean antigenic protein on intestinal health. Aquaculture 501, 202–212; doi:10.1016/j.aquaculture.2018.11.010
Yasuoka, A. and Abe, K. 2009. Gustation in fish: search for prototype of taste perception. Results. Problems. Cell. Differentiation. 47, 239–255; doi:10.1007/400_2008_6
Yuan, X.C., Liang, X.F., Cai, W.J., He, S., Guo, W.J. and Mai, K.S. 2020. Expansion of sweet taste receptor genes in grass carp (Ctenopharyngodon idellus) coincided with vegetarian adaptation. BMC Evol. Biol. 20(1), 1–15; doi:10.1186/s12862-020-1590-1
Zhang, Y., Hoon, M.A., Chandrashekar, J., Mueller, K.L., Cook, B., Wu, D., Zuker, C.S. and Ryba, N.J. 2003. Coding of sweet, bitter, and umami tastes: different receptor cells sharing similar signaling pathways. Cell 112(3), 293–301; doi:10.1016/S0092-8674(03)00071-0
Zhu, T., Du, J., Song, H., Lei, C., Cen, Y., Wang, C. and Li, S. 2024. Whole genome resequencing reveals the correlation between selection signatures and adaptability of Micropterus salmoides to artificial fed. Scientific Rep. 14(1), 1–11; doi:10.1038/s41598-024-80904-2
Zhu, W.Q., Yuan, X.Q., Luo, H.J., Shao, J.C. and Chen, X.H. 2021. High percentage of dietary soybean meal inhibited growth, impaired intestine healthy and induced inflammation by TLR-MAPK/NF-κB signaling pathway in large yellow croaker (Larimichthys crocea). Aquac. Rep. 20(2021), 1–8; doi:10.1016/j.aqrep.2021.100735










