| Research Article | ||
Open Vet. J.. 2026; 16(1): 626-634 Open Veterinary Journal, (2026), Vol. 16(1): 626-634 Research Article The potential of mollusc-associated fungi as anti-pathogensJosua Gabriel Lumban Gaol1, Delianis Pringgenies1*, Wilis Ari Setyati1, Sunelsya Surya1, Muhammad Zainuddin2 and Dafit Ariyanto31Department of Marine Science, Faculty of Fisheries and Marine Science, Diponegoro University, Semarang, Indonesia 2Aquaculture Study Program, Faculty of Science and Technology, Islamic University of Nahdlatul Ulama Jepara, Jepara, Indonesia 3Research Center for Ecology, National Research and Innovation Agency, Cibinong, Indonesia *Corresponding Author: Delianis Pringgenies. Department of Marine Science, Faculty of Fisheries and Marine Science, Diponegoro University, Semarang, Indonesia. Email: delianispringgenies [at] lecturer.undip.ac.id Submitted: 01/08/2025 Revised: 02/12/2025 Accepted: 23/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
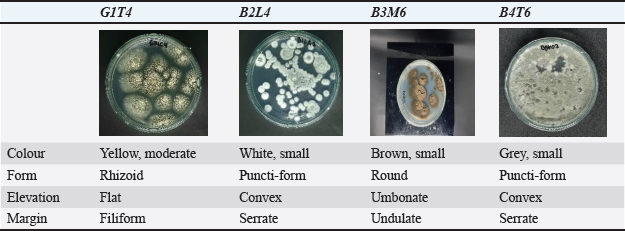
AbstractBackground: Molluscs are invertebrate organisms known to produce secondary metabolites that help them adapt to their environment. Aim: This study aims to explore the potential of fungi associated with molluscs and evaluate their secondary metabolites as potential antimicrobial agents against human pathogens. Methods: The fungi isolated from molluscs were examined both macroscopically and microscopically. Promising isolates were selected and cultured in bulk for phytochemical screening and thin-layer chromatography analysis. Extract from these fungal tested at a concentration 1,000 μg/disc. Observations were recorded every 12 hours over a 36 periods. Molecular identification of the selected isolated was performed using the Quick-DNA Magbead Plus Kit (Zymo Research, D4082). Statistical analysis was used to determine the significant differences isolates and control (p <0.05). Results: The extract from fungal isolate G1T4 demonstrated both antibacterial and antifungal activity against all four test pathogens, as evidenced by the formation of clear inhibition zones. Extracts from isolates B3M6, B4T6, and B2L4 showed antibacterial effect against Staphylococcus aureus, Escherichia coli, and the test fungus Candida albicans, but exhibited no activity against the fungal pathogen Trichoderma harzianum. The presence of inhibition zones suggests that these extracts contain active secondary metabolites or active compounds in the associated fungus extract. The result of statistical analysis showed significant differences (p < 0.05) between isolates and controls at all observation times (12–72 hours) against E. coli and S. aureus, C. albicans, and T. harzianum showed GT14 are consistently higher than other isolates (p < 0.05). Conclusion: Molecular analysis revealed that isolate G1T4 is closely related to Aspergillus subramanianii (accession number PQ325939), while isolate B3M6 is closely related to Cladosporium halotolerans (accession number PQ325940). Keywords: Human pathogens, Molluscs, Associated fungi, Secondary metabolites. IntroductionPathogens, including such as bacteria, viruses, fungi, and parasites, are microorganisms capable of causing diseases in human. Although antibiotics and antifungal agents have proven effective in treating many of these infections, the emergence of resistant strains continues to pose significant challenges. Mollusks possess robust defence mechanisms that enable their associated fungi to survive in extreme environments. These fungi, in turn, are potential sources of bioactive compounds with anti-pathogenic properties (Avila, 2020). Molluscs are known to produce secondary metabolites as part adaptive response of their adaptive response to environmental stressors. These compounds have promising applications in both the pharmaceutical and nutraceutical fields (Nurhikma. et al., 2021 ; Surya et al., 2025). Interestingly, fungi associated with mollusks also produce secondary metabolites. The interaction between molluscs and fungi and is biologically significant, as fungi may act as symbionts, food sources, or cohabitants (Ayer et al., 2018). These associated fungi have demonstrated the ability to synthesize pharmacologically valuable (Setyati et al., 2023). Previous study have shown that fungi living in symbiosis with other organisms can produce a wide range of antimicrobial compounds that are effective against human pathogens (Ayer et al., 2018; Varijakzhan et al., 2021). Pathogenic microbes are found in various environments, including the human body, and are responsible for numerous diseases. These illnesses often result from infections or toxins produced by bacteria and fungi (Panayidou et al., 2014). The term pathogenicity refers to a microorganism’s ability to cause disease in its host, leading to structural and one promising strategy for disease prevention involves harnessing secondary metabolites produced by fungi associated with other organisms. These compounds exhibit to have antibacterial and antifungal properties and have shown potential in treating infections microbes (Alam et al., 2021). Moreover, some of these metabolites may stimulate the human immune system, some of these metabolites enhancing resistance to infections (El-Bondkly et al., 2021). Fungal associations and interactions with host organisms can aid in developing more effective bioprospecting approaches for combating infectious diseases (Calvo and Cary, 2015). Despite this potential, research on fungi associated with molluscs in the waters of Lombok, West Nusa Tenggara, remains limited. This study seeks to address that gap by characterizing fungal isolates from molluscs, evaluating their potential anti-pathogenic activity, identifying the secondary metabolites, and assessing their potential as antibacterial and antifungal agents. Material and MethodsData collection and isolation of mollusc-associated fungiMollusc samples were collected from the three distinct marine ecosystems: Lombok, West Nusa Tenggara, mangrove forest, seagrass, and coral reef. The mollusk species sampled included Cypraea moneta, Spondylus sp., Atrina vexillum, and Spondylus squamosus. Fungal isolation was performed using sterilized Potato Dextrose Agar, prepared via autoclaving. The pour plate method, as described by Angelia (2020). This technique involved serial dilutions of the mollusc samples using sterile seawater, with dilution levels ranging from 10¹ to 106. Macroscopic and microscopic identification and mass culturePurification of the isolated fungi was carried out to obtain pure colonies. These isolates were then characterized both macroscopically and microscopically. Macroscopic observation included colony colour, size, shape, elevation, conidial structure, and the hyphal septation (Sharma et al., 2023). Selected fungal isolates were inoculated into 50 ml of Potato Dextrose Broth (PDB) medium in 250 ml Erlenmeyer flasks and incubated for 72 hours at 120 rpm. Subsequently, 1 ml of the initial culture was transferred into 250 ml of PDB medium in 1 l Erlenmeyer flasks. These cultures were incubated under 120 rpm for approximately 14 days (Tangapo et al., 2022). Sample extraction Thin layer chromatography (TLC)After 14 days of mass culture, the mycelia from each selected fungal isolate were filtered using Whatman No. 42 filter paper. The resulting supernatant was extracted with ethyl acetate in a 1:1 volume ration. The organic phase was then evaporated using a rotary evaporator at a controlled. The temperature of 40°C is used to preserve the integrity of the bioactive compounds and avoid exceeding the boiling point of ethyl acetate (Sibero et al., 2018). Thin layer chromatography was employed to separate and identify bioactive compounds present in the cruide extract. The mobile phase used was a mixture of ethyl acetate and chloroform in a 7:3 volume ratio. After development, the retention factor (RF) value of the compounds was calculated on the silica gel plates (Yulistyani et al., 2021). Phytochemical screening and molecular identificationPhytochemical screening was conducted to detect the presence of bioactive compounds in the fungal extracts (Pringgenies et al., 2021). This screening involved the use of specific reagents to test for flavonoids, alkaloids, saponins, and tannins. The crude fungal extracts were evaluated for antibacterial and antifungal activity using the agar diffusion method (Zainuddin et al., 2022). Extracts were prepared at a concentration of 1,000 µg per disc and tested against selected human for molecular identification, DNA was extracted using the Quick-DNA MagBead Plus Kit (Zymo Research, D4082). DNA amplification was conducted with the MyTaq HS Red Mix kit, 2X (Bioline, BIO-25048), following protocol B/7.2.1/IKP/003. The amplification products were analyzed via electrophoresis (B/7.2.1/IKP/005), and sequencing was carried out using the Sanger method with capillary electrophoresis. Statistical analysisStatistical analysis of the inhibition zones was performed using one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test, and results were expressed as mean ± standard deviation (SD), with significance set at p < 0.05 with R Studio software. Ethical approvalNot needed for this study. ResultsMacroscopic characterization and microscopic identificationFollowing purification, four fungal isolates associated with mollusks, were successfully obtained. These isolates were coded and characterized macroscopically based on colour, shape, size, elevation, and edge morphology. The results of this macroscopic characterization are summarized in Table 1. Table 1. Macroscopic identification results of fungal isolates associated with molluscs.
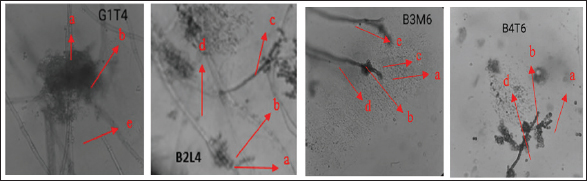
Microscopic identification was performed using a binocular microscope at 100x magnification, with lactophenol cotton blue used as a staining agent. The fungal structures were examined and identified based on key morphological features, including conidia, vesicles, phialides, conidiophore, and septate hyphae. Representative microscopic observations are shown in Figure 1.
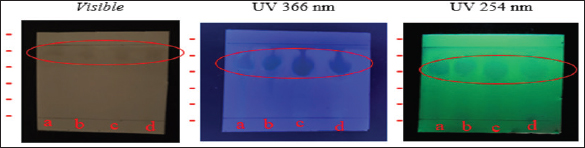
Fig. 1. Mollusc Association Fungi: a) conidia, b) vesicle, c) phialides, d) conidiophore, e) hyphae. Phytochemical test screening and thin-layer chromatographyPhytochemical screening was performed on extracts obtained from fungi associated with molluscs to qualitatively identify the types of secondary metabolites present. Using specific reagents, four fungal isolates, G1T4, B3M6, B4T6, and B2L4, were tested. The results revealed that the extract from isolate G1T4 contained flavonoids, alkaloids, and saponins. The B3M6 extract showed the presence of flavonoids and alkaloids. The B3M6 extract showed the presence of B4T6, and B2L4 extracts contained alkaloids only. To further analyze he bioactive compound, extracts from all four isolates were subjected to TLC. This technique was used to separate the compound fractions based on polarity. TLC plates were observed under ultraviolet (UV) light at 366 nm and 254 nm, and RF values were the results from the TLC analysis are presented in Figure 2.
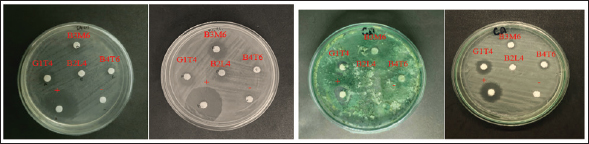
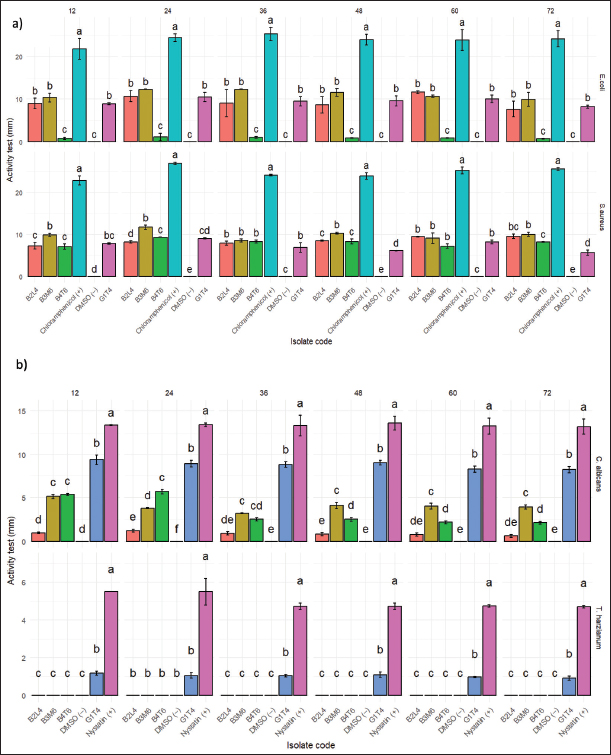
Fig. 2. TLC test results of mollusk association fungus isolate extract. Antibacterial and antifungal activity test against human pathogensSemi-quantitative antibacterial and antifungal assays were conducted to evaluate the bioactivity of fungal extracts derived from mollusc (Fig. 3). The extract from isolate G1T4 demonstrated strong antibacterial activity against Escherichia coli with a clear inhibition zone of 10.47 mm (±1.0). Against Staphylococcus aureus, the extract produced a zone of 9.05 mm (±0.14), categorized as moderate (Fig. 4a). In antifungal testing, the G1T4 showed strong activity against Candida albicans, with a zone of 9.43 mm (± 0.53). However, its activity against Trichoderma harzianum was minimal, with a clear zone of only 1.18 mm (±0.11), classified as very weak (Fig. 4b). The extract from isolate B3M6 exhibited strong antibacterial activity, producing a clear zone of 12.31 mm (±0.02) against E. coli and 11.75 mm (±0.49) against S. aureus (Table 2).
Fig. 3. Antibacterial (a & b) and antifungal (c & d) test results.
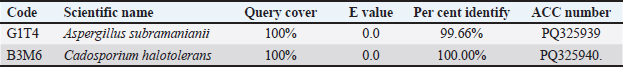
Fig. 4. (a) antibacterial and (b) antifungal activity tests that cause disease (mm). Table 2. Results of query cover and percent identify.
In antifungal testing, B3M6 showed weak activity against C. albicans with a zone of 5.13 mm (±0.25), and no detectable activity against T. harzianum (Fi. The extract from isolate B4T6 showed weak antibacterial activity against E. coli, with a clear zone of 1.13 mm (±0.87). It produced a zone of 9.35 mm (±0.04) categorized as moderate. In antifungal test B4T6 activity against C. albicans demonstrated activity with a clear zone of 5.68 mm (±0.25). Molecular identificationThe molecular identification of potential mollusc-associated fungal isolates G1T4 and B3M6 was conducted based on query coverage percentage identity. Phylogenetic analysis was performed using Penicilllium chrysogenum an outgroup, and the reconstructured phylogenetic tree is presented in Figure 5.
Fig. 5. Phylogenetic tree of potential isolates G1T4 and B3M6. DiscussionMacroscopic characterization and microscopic identificationSampling from Lombok Waters, West Nusa Tenggara, yielded four distinct mollusc species. One of the identified samples belonged to the gastropod class (Cypraea moneta), while the remaining samples were from the bivalvia class, including Spondylus sp., S. squamosus, and A. vexillum. Gastropod and bivalve mollusc exhibit unique morphological traits, underscoring the ecological diversity and the region’s marine habitats. Molluscs are known for their adaptability across a wide range of aquatic environments, which contributes to their ecological success. The fungal isolates (G1T4, B2L4, B3M6, and B4T6) were purified and characterized to distinguish their origin from the specific mollusc host (Table 1). The characterization revealed in colony color, edge morphology, and elevation. Fungal isolates associated with seagrass ecosystems predominantly exhibited green to greyish colony colors. The distinctive feature reflects the biodiversity of fungi associated with different marine habitats. Aligns with findings by Baron and Rigobelo (2022), who reported that fungi adapted to specific environments often display unique colony colors and morphologies, indicating habitat-driven diversity. According to Naranjo-Ortiz and Gabaldón (2019), such variations arise from physiological adaptations to environmental conditions. However, it is important to note that the colony size and colour may change over time as the fungal culture age (Sy. Pakaya et al., 2023). Microscopic identification of fungal isolates associated with molluscMicroscopic identification was conducted to examine the morphological characteristics of four potential fungal isolates using a binocular microscope at 100x magnification (Fig. 1). The objective of these observations was to assess the structure of hyphae, the presence of septa, and the shape of spores (Demeni et al., 2021). Microscopic examination of fungal cultures prepared on glass slides revealed that the following isolates G1T4 and B2L4 exhibited septate hyphae and well-defined conidiophores. Isolate B3M6 also displayed conidiophores, but its hyphae were non-septate. Isolate B4T6 showed the presence of conidiophores; however, hyphae and septa were not clearly visible under the microscope. Conidiophores, also known as conidial stalks, are specialized structures where conidia (asexual spores) are produced. Hyphae are the fundamental filamentous structures of fungi and can vary in size and appearance depending on the species and environmental conditions (Arifah et al., 2023). Phytochemical screening and TLC analysis of mollusc-associated fungal extractsMass cultures of potential mollusc-associated fungal isolates were extracted using ethyl acetate as the solvent. Ethyl acetate was selected due to because of its semi-polar nature, which enables efficient extraction of metabolites and bioactive compounds from the fungal biomass. It is a volatile, non-hygroscopic solvent with low toxicity (Minarni et al., 2017). The resulting crude extracts underwent phytochemical screening to identify the presence of secondary metabolites. The flavonoid test, conducted using 37% hydrochloric acid and magnesium powder, confirms the presence of flavonoids in G1T4 and B3M6. Alkaloid screening using Mayer’s, Wagner’s, and Dragendorff’s reagents indicated that all four fungal extracts (G1T4, B3M6, B4T6, and B2L4) tested positive for alkaloids. The saponin test revealed a result only for G1T4, while the other isolates showed no saponin content. Tests for steroids and tannins, using the Liebermann–Burchard reagent and 5% FeCl3, respectively, did not detect these compounds in any of the fungal extracts. The production of bioactive compounds by these fungi is believed to be influenced by their interaction with mollusc hosts. Studies by Photolo et al. (2020) and Singh and Kumar (2023) suggest that bioactive compounds may be transferred from the host to the associated fungi, enabling the fungi to synthesize secondary metabolites as a defence against pathogenic microbes. TLC was performed to further analyze the extracts. Each sample was applied to a silica gel plate (stationary phase), and a mixture of ethyl acetate: chloroform (7:3) served as the mobile phase (Fig. 2). Visual inspection revealed faint stains on plates, which became clearly visible under UV light at wavelengths of 366 nm and 254 nm. The UV observation showed that the stains had Rf values of 0.88 and 0.61 for the G1T4 extract, 0.96 and 0.60 for the B3M6 extract, 0.85 and 0.56 for the B4T6 extract, and 0.86 and 0.56 for the B2L4 extract (Table 2). Rf values indicate the identity of secondary metabolites present in the extracts (Khan et al., 2014). The provide insight into Rf values ranging from 0.85 to 0.96 are indicative of alkaloids (Sari et al., 2024), while values between 0.56 and 0.61 suggest the presence of flavonoids (Fathoni et al., 2021). Antibacterial and antifungal testing of fungal extractsAntibacterial and antifungal assays were performed to evaluate the bioactive of fungal extracts against selected human pathogens, including E. coli, S. aureus, C. albicans, and T. harzianum, all known to cause infections in human (Yagnik et al., 2021). Fungal derived from mollusc-associated isolated, demonstrated inhibitory extract from the fungus associated with molluks exhibited antibacterial and antifungal activity against these pathogens. A concentration of 1,000 μg per disc was used for all tests. This dosage was concentration was selected based on prior research, indicating that lower concentrations resulted in reduced antibacterial and antifungal activity (Mehta, 2020). Antibacterial and activity and molecular identification of mollusc-associated fungiThe results of antibacterial and antifungal testing demonstrated that the extract from fungal isolate G1T4 exhibited strong activity against the all four tested pathogens (S. aureus, E. coli, and C. albicans) as evidenced by the formation of prominent clear zones. Isolates B3M6, B4T6, and B2L4 also showed antibacterial activity against the pathogens S. aureus, E. coli, and C. albicans, but none exhibited antifungal activity against the test fungal pathogen T. harzianum (Fig. 3). The extracts from G1T4, which was isolated from the mollusc Cypraea moneta (Linnaeus, 1758), demonstrated notable had antibacterial and antifungal potential. The formation of a clear zone indicates the presence of secondary metabolites or active compounds capable of disrupting the cell walls of the bacteria and fungi (Aini, 2023). In comparison, research by Xu et al. (2015) showed that extracts from algae, sponge, and mangrove symbiont fungi exhibited activity against S. aureus and E. coli. In contrast, extracts from mollusc-associated fungi demonstrated more pronounced antibacterial effects, likely due to the presence of potential bioactive secondary metabolites (Hawar, 2022). Phytochemical and TLC screening confirm the presence of flavonoids, alkaloids, and saponins in these extract compounds, known for their antimicrobial properties. Statistical analysis of the inhibition zones was performed using one-way ANOVA followed by Tukey’s post hoc test, and results were expressed as mean ± SD, with significance set at p < 0.05 in the presented Figure 3a and Figure 4b. Flavonoids and their derivatives can inhibit microbial growth by disrupting cell wall integrity and interfering with cell division. This disruption leads to deformation of fungal cell walls, composed of β-glucan and chitin, resulting in cell death (Yun and Lee, 2017). Combining various secondary metabolites from fungal extracts is considered an effective strategy to combat microbial infections and resistance (Keita et al., 2022). Different types and concentrations of these compounds may act through multiple mechanisms (multi-targeting), helping to overcome relapses and resistance associated with the bacteriostatic/fungistatic and bactericidal/fungicidal effects. Molecular identification revealed that the potential isolate G1T4 corresponds to Aspergillus subramanianii strain PQ325939, while isolate B3M6 corresponds to Cladosporium halotolerans strain PQ325940. Phylogenetic tree reconstruction showed that G1T4 clustered within the genus Aspergillus, and B3M6 formed a clade with the genus Cladosporium, indicating that isolate G1T4 is closely related to the genus Cladosporium (Fig. 5). These relationships were supported by 100 % query coverage and identity in the NCBI database. Aspergillus subramanianii was isolated from C. moneta and C. halotolerans. These fungal species are known to inhabit marine ecosystems and are associated with marine organisms (Zhang et al., 2022). Previous studies have shown that the C. halotolerans and A. subramanianii rarely cause disease in humans (Maduri et al., 2015; Kaur et al., 2023). ConclusionThe isolation of fungi associated with molluscs from the waters of Lombok, West Nusa Tenggara, resulted in four promising fungal isolates, each exhibiting distinct macroscopic and microscopic characteristics. Among them, the extract from isolate G1T4 demonstrated strong antibacterial and antifungal activity against E. coli, S. aureus, C. albicans, and T. harzianum, highlighting its potential as a broad-spectrum antimicrobial agent. In comparison, extracts from isolates B3M6, B4T6, and B2L4 showed selective activity, being effective only against E. coli, S. aureus, and C. albicans. Phytochemical analysis confirmed the presence of secondary metabolites such as flavonoids, alkaloids, and saponins, in these fungal extracts compound known for their antimicrobial properties. These findings suggest that mollusc-associated fungi are a valuable source of bioactive compounds and can be used in the development of anti-pathogen treatments for humans. AcknowledgmentsThe authors would like to express their sincere gratitude to the Faculty of Fisheries and Marine Science, Diponegoro University, for supporting this research. Conflict of interestThe authors declare that there is no conflict of interest regarding the publication of this article. FundingNone. Authors’ contributionsJosua Gabriel Lumban Gaol: Writing—review & editing, Writing—original draft, Investigation. Delianis Pringgenies: Conceptualization, Writing—review & editing, Writing—original draft, Funding acquisition. Wilis Ari Setyati: Conceptualization, Writing—review & editing, Writing—original draft. Sunelsya Surya: Methodology, Investigation, Formal analysis, Data curation. Muhammad Zainuddin: Methodology, Investigation, Formal analysis, Data curation. Dafit Ariyanto: Writing—review & editing, Validation, submission. Data availabilityAll Data in the manuscript. ReferencesAini, M. 2023. The effect of modified media on the antibacterial activity of the sea sponge symbion fungi, Fusarium solani. J. BioTrop. 23(3), 546–553. Alam, B., Lǐ, J., Gě, Q., Khan, M.A., Gōng, J., Mehmood, S., Yuán, Y. and Gǒng, W. 2021. Endophytic fungi: from symbiosis to secondary metabolite communications or vice versa?. Front. Plant Sci. 12, 1–24. Angelia, I.O. 2020. Use of pour plate methods on microbial test on coconut flour. JASc 4(1), 43–51. Arifah, F., Aini, L.Q. and Muhibuddin, A. 2023. Molecular and morphological characterization of fungi isolated from nutmeg (Myristica fragrans) in North Sulawesi, Indonesia. Biodiversitas 24(1), 441–453. Avila, C. 2020. Terpenoids in marine heterobranch molluscs. Mar. Drugs 18(162), 1–38. Ayer, P.I.L., Sabdono, A. and Trianto, A. 2018. Activity of sponge symbiont fungi against Trichophyton sp. Fungi in biak island, Biak-Numfor Regency, Papua. J. Acropora Ilmu. Kelautan. Dan. Perikanan. Papua 4(1), 50–57. Baron, N.C. and Rigobelo, E.C. 2022. Endophytic fungi: a tool for plant growth promotion and sustainable agriculture. Mycol 13(1), 39–55. Calvo, A.M. and Cary, J.W. 2015. Association of fungal secondary metabolism and sclerotial biology. Front. Microbiol. 6, 1–16. Demeni, P.C.E., Betote, P.H.D., Kom, C.W., Tchamgoue, E.N., Ndedi Moni, E.D.F., Foumane Maniepi, J.S., Agbor, G.A. and Nyegue, M.A. 2021. Endophytic fungi from alstonia boonei de wild and greenwayodendron suaveolens (engl. and diels) verdc. subsp. suaveolens possess inhibitory activity against pneumonia causing bacteria. Evid. Based. Complement. Alternat. Med. 2021, 1. El-Bondkly, E.A.M., El-Bondkly, A.A.M. and El-Bondkly, A.A.M. 2021. Marine endophytic fungal metabolites: a whole new world of pharmaceutical therapy exploration. Heliyon 7(3), e06362. Fathoni, A., Hudiyono, S., Budianto, E., Cahyana, A. H. and Agusta, A. Metabolite detection and antibacterial activity of fungal endophytic extracts isolated from brotowali (Tinospora crispa) plants using TLC-bioautography assay. In IOP Conference Series: Materials Science and Engineering, (Vol. 1011, No. 1). IOP Publishing, Bristol, United Kingdom, 2021, p 1–10. Hawar, S.N. 2022. Extracellular enzyme of endophytic fungi isolated from Ziziphus spina leaves as medicinal plant. Int. J. Biomater. 2022, 1–9. Kaur, H., Gupta, P., Ahmad, H., Shankarnarayan, S.A., Srivastava, S., Sahu, S., Karuna, T., Narang, T., Gupta, S., Ghosh, A. and Rudramurthy, S.M. 2023. Cladosporium halotolerans: exploring an Unheeded Human Pathogen. Mycopathologia 188(6), 1027–1040. Keita, K., Darkoh, C. and Okafor, F. 2022. Secondary plant metabolites as potent drug candidates against antimicrobial-resistant pathogens. SN. Appl. Sci. 4(8), 1–10. Khan, H.A., Arif, I.A., Williams, J.B., Champagne, A.M. and Shobrak, M. 2014. Skin lipids from Saudi Arabian birds. Saudi J. Biol. Sci. 21(2), 173–177. Maduri, A., Patnayak, R., Verma, A., Mudgeti, N., Kalawat, U. and Asha, T. 2015. Subcutaneous infection by Cladosporium sphaerospermum-A rare case report. Indian J. Pathol. Microbiol. 58(3), 406–407. Mehta, R. 2020. A comparative study of antibacterial and antifungal activities of extracts from four indigenous plants. Bioinformation 16(3), 267–273. Minarni., Artika, I.M., Julistiono, H., Bermawie, N., Riyanti, E.I., Hasim. and Hasan, A.E.Z. 2017. Anticancer activity test of ethyl acetate extract of endophytic fungi isolated from soursop leaf (Annona muricata L.). Asian Pac. J. Trop. Med. 10(6), 566–571. Naranjo-Ortiz, M.A. and Gabaldón, T. 2019. Fungal evolution: major ecological adaptations and evolutionary transitions. Biol. Rev. 94(4), 1443–1476. Nurhikma.., Mirsa.. and Wulandari, D.A. 2021. Bioactive compound and antioxidant activity of balelo sea snail (Conomurex sp.). J. Pengolah. Has. Perikan. Indones. 24(1), 11–19. Panayidou, S., Ioannidou, E. and Apidianakis, Y. 2014. Human pathogenic bacteria, fungi, and viruses in Drosophila. Virulence 5(2), 253–269. Photolo, M.M., Mavumengwana, V., Sitole, L. and Tlou, M.G. 2020. Antimicrobial and antioxidant properties of a bacterial endophyte, Methylobacterium radiotolerans MAMP 4754, isolated from combretum erythrophyllum seeds. Int. J. Microbiol. 2020, 1–11. Pringgenies, D., Setyati, W.A., Djunaedi, A., Pramesti, R., Rudiyanti, S. and Ariyanto, D. 2021. Exploration of antimicrobial potency of mangrove symbiont against multi-drug resistant bacteria. J. Ilm. Perikan. Kelaut. 12(2), 222–232. Sari, A.I.P., Fitriana, F. and Amirah, S. 2024. Antibacterial activity of endophytic fungi isolated from bidara roots (Ziziphus mauritiana Lam.) against bacteria that cause skin infections using TLC-Bioutography. Microbiol. Sci. 4(1), 31–41. Setyati, W.A., Sedjati, S., Samudra, A. and Ariyanto, D. 2023. Investigation of seagrass-associated fungi as antifouling candidates with anti-bacterial properties. Jordan J. Biol. Sci. 16(2), 323–327. Sharma, N., Sharma, N., Sharma, S., Sharma, P. and Devi, B. 2023. Identification, morphological, biochemical, and genetic characterization of microorganisms. Basic Biotechniques for Bioprocess and Bioentrepreneurship, 47–84. Sibero, M.T., Radjasa, O.K., Sabdono, A., Trianto, A., Triningsih, D.W. and Hutagaol, I.D. 2018. Antibacterial activity of indonesian sponge associated fungi against clinical pathogenic multidrug resistant bacteria. J. Appl. Pharm. Sci. 8(2), 88–94. Singh, V.K. and Kumar, A. 2023. Secondary metabolites from endophytic fungi: production, methods of analysis, and diverse pharmaceutical potential. Symbiosis 90(2), 111–125. Surya, S., Pringgenies, D. and Sedjati, S. 2025. Antibacterial and enzymatic activities of symbiotic bacteria from gastropods and bivalves in marine skincare applications. Int. J. Design Nature Ecodynamics 20(1), 83–90. Sy. Pakaya, M., Ain Thomas, N., Hasan, H., H. Hutuba, A. and Mbae, G. 2023. Isolasi, karakterisasi, dan uji antioksidan fungi endofit dari tanaman batang kunyit (Curcuma domestica Val.,). J. Syifa. Sci. Clin. Res. 5(2), 220–231. Tangapo, A.M., Mambu, S.M., Kolondam, B., Pasappa, N. and Pelealu, J. 2022. Eksplorasi fungi endofit tumbuhan mangrove Avicennia marina sebagai penghasil senyawa antibakteri. Jurnal. Ilmu. Alam. Dan. Lingkungan. 13(2), 25–31. Varijakzhan, D., Loh, J.Y., Yap, W.S., Yusoff, K., Seboussi, R., Lim, S.H.E., Lai, K.S. and Chong, C.M. 2021. Bioactive compounds from marine sponges: fundamentals and applications. Mar Drugs.19(246), 1–38. Xu, L., Meng, W., Cao, C., Wang, J., Shan, W. and Wang, Q. 2015. Antibacterial and antifungal compounds from marine fungi. Mar. Drugs 13(6), 3479–3513. Yagnik, D., Ward, M. and Shah, A.J. 2021. Antibacterial apple cider vinegar eradicates methicillin resistant Staphylococcus aureus and resistant Escherichia coli. Sci. Rep. 11(1), 1–7. Yulistyani, I., Poernomo, A.T. and Isnaeni, I. 2021. KLT-bioautografi ekstrak etil asetat supernatan hasil fermentasi streptomyces g isolat tanah rumah kompos bratang Surabaya. J. Farm. Ilmu. Kefarmasian. Indones. 8(1), 1–9. Yun, D.G. and Lee, D.G. 2017. Silymarin exerts antifungal effects via membrane-targeted mode of action by increasing permeability and inducing oxidative stress. Biochimica. Et. Biophysica. Acta. 1859(3), 467–474. Zainuddin, M., Pringgenies, D., Radjasa, O.K. and Haeruddin. 2022. Natural bioactive compounds of sponge-associated fungi with three marine ecosystems in Karimunjawa Island, Indonesia. CMUJ. Nat. Sci. 21(1), e2022009. Zhang, K., Hu, J., Yang, S., Xu, W., Wang, Z., Zhuang, P., Grossart, H. and Luo, Z. 2022. Biodegradation of polyester polyurethane by the marine fungus Cladosporium halotolerans. J. Hazard. Mater. 437, 129406 | ||
| How to Cite this Article |
| Pubmed Style Gaol JGL, Pringgenies D, Setyati WA, Surya S, Zainuddin M, Ariyanto D. The potential of mollusc-associated fungi as anti-pathogens. Open Vet. J.. 2026; 16(1): 626-634. doi:10.5455/OVJ.2026.v16.i1.58 Web Style Gaol JGL, Pringgenies D, Setyati WA, Surya S, Zainuddin M, Ariyanto D. The potential of mollusc-associated fungi as anti-pathogens. https://www.openveterinaryjournal.com/?mno=274603 [Access: February 04, 2026]. doi:10.5455/OVJ.2026.v16.i1.58 AMA (American Medical Association) Style Gaol JGL, Pringgenies D, Setyati WA, Surya S, Zainuddin M, Ariyanto D. The potential of mollusc-associated fungi as anti-pathogens. Open Vet. J.. 2026; 16(1): 626-634. doi:10.5455/OVJ.2026.v16.i1.58 Vancouver/ICMJE Style Gaol JGL, Pringgenies D, Setyati WA, Surya S, Zainuddin M, Ariyanto D. The potential of mollusc-associated fungi as anti-pathogens. Open Vet. J.. (2026), [cited February 04, 2026]; 16(1): 626-634. doi:10.5455/OVJ.2026.v16.i1.58 Harvard Style Gaol, J. G. L., Pringgenies, . D., Setyati, . W. A., Surya, . S., Zainuddin, . M. & Ariyanto, . D. (2026) The potential of mollusc-associated fungi as anti-pathogens. Open Vet. J., 16 (1), 626-634. doi:10.5455/OVJ.2026.v16.i1.58 Turabian Style Gaol, Josua Gabriel Lumban, Delianis Pringgenies, Wilis Ari Setyati, Sunelsya Surya, Muhammad Zainuddin, and Dafit Ariyanto. 2026. The potential of mollusc-associated fungi as anti-pathogens. Open Veterinary Journal, 16 (1), 626-634. doi:10.5455/OVJ.2026.v16.i1.58 Chicago Style Gaol, Josua Gabriel Lumban, Delianis Pringgenies, Wilis Ari Setyati, Sunelsya Surya, Muhammad Zainuddin, and Dafit Ariyanto. "The potential of mollusc-associated fungi as anti-pathogens." Open Veterinary Journal 16 (2026), 626-634. doi:10.5455/OVJ.2026.v16.i1.58 MLA (The Modern Language Association) Style Gaol, Josua Gabriel Lumban, Delianis Pringgenies, Wilis Ari Setyati, Sunelsya Surya, Muhammad Zainuddin, and Dafit Ariyanto. "The potential of mollusc-associated fungi as anti-pathogens." Open Veterinary Journal 16.1 (2026), 626-634. Print. doi:10.5455/OVJ.2026.v16.i1.58 APA (American Psychological Association) Style Gaol, J. G. L., Pringgenies, . D., Setyati, . W. A., Surya, . S., Zainuddin, . M. & Ariyanto, . D. (2026) The potential of mollusc-associated fungi as anti-pathogens. Open Veterinary Journal, 16 (1), 626-634. doi:10.5455/OVJ.2026.v16.i1.58 |