Open Veterinary Journal, (2025), Vol. 15(12): 6470-6479
Research Article
10.5455/OVJ.2025.v15.i12.31
Exploring the matrilineal ancestry of swamp buffaloes (Bubalus bubalis) in Sijunjung Regency, West Sumatra, through partial cytochrome-b sequencing
Muhammad Ridho1, Muhammad Cahyadi1*, Ahmad Pramono1, Widya Pintaka Bayu Putra2,
Hartati Hartati3, Eko Handiwirawan3, Aprisal Aprisal4,5, Afdal Yulius6, Rasiravathanahalli Kaveriyappan Govindarajan7 and Muhammad Indra Maulana8
1Department of Animal Science, Universitas Sebelas Maret, Surakarta, Indonesia
2Research Center for Applied Zoology, National Research and Innovation Agency, Cibinong, Indonesia
3Research Center for Animal Husbandry, National Research and Innovation Agency, Cibinong, Indonesia
4Department of Animal Husbandry and Veterinary Medicine, West Sumatera Province, Padang, Indonesia
5Department of Animal Production Technology, Andalas University, Padang, Indonesia
6Department of Procurement, Agency for Regional Development of Sijunjung Regency, Sijunjung, Indonesia
7Department of Biotechnology, Karpagam Academy of Higher Education, Coimbatore, India
8Department of Biology, Faculty of Science, Institut Teknologi Sumatra, Lampung, Indonesia
*Corresponding Author: Muhammad Cahyadi. Faculty of Animal Science, Universitas Sebelas Maret, Surakarta, Indonesia. Email: mcahyadi [at] staff.uns.ac.id
Submitted: 10/07/2025 Revised: 27/10/2025 Accepted: 08/11/2025 Published: 31/12/2025
© 2025 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Background: Buffaloes (Bubalus bubalis) are a significant livestock in Indonesia, with approximately 36% (201,060 heads) of their breeding populations located on Sumatera Island, including Sijunjung Regency.
Aim: This study aimed to ascertain the matrilineal lineage of swamp buffaloes in Sijunjung Regency, Indonesia, using the mitochondrial cytochrome b (Cyt-b) region.
Methods: Blood samples were obtained from 23 adult female swamp buffaloes for DNA extraction and analysis. A total of 906 bp of bubaline Cyt-b amplicons was amplified for forward sequencing. Six bioinformatics programs (BioEdit, DNAsp, MEGA, STRUCTURE, BEAST, and R) were used for sequence alignment, haplotype diversity estimation, phylogenetic tree construction, pairwise genetic distance calculations, and genetic admixture analysis.
Results: Seventeen mutation sites were identified in the Cyt-b region. Consequently, four haplotypes (H) of Cyt-b were identified: H1 (56%), H2 (22%), H3 (17%), and H4 (5%). The haplotype diversity (Hd) and nucleotide diversity (pi) for the partial Cyt-b analyzed were 0.645 ± 0.081 and 0.003 ± 0.001, respectively. Phylogenetic tree analysis indicated that the swamp buffaloes in Sijunjung Regency were classified within Haplogroup 1 and exhibited close genetic ties to buffaloes from South Asia (Bangladesh), Southeast Asia (Thailand, Laos, Vietnam, Philippines), and East Asia (Taiwan, China). No evidence of genetic introgression from Indian and Pakistani buffaloes was found in the studied population.
Conclusion: The matrilineal origin of buffaloes in Sijunjung Regency may be traced back to the southeastern Chinese buffalo (Lineage A) via two migration routes: the land-bridge (China - IndoChina - Malaya Peninsula - Sumatra) and the insular pathway (China - Taiwan - Philippines - Borneo - Sumatra).
Keywords: Buffaloes, Cyt-b region, Matrilineal, Sequencing, Sijunjung Regency.
Introduction
Buffalo (Bubalus bubalis) is a significant livestock species in Indonesia, which is used for both draught work and meat and milk production. In 2024, the buffalo population in Indonesia was recorded at 556,794 heads, with a total carcass production of 17,361.50 tons (Kementan, 2024). Currently, the Indonesian Ministry of Agriculture has officially registered nine swamp buffalo breeds as AnGR, namely, Gayo, Simeulue, Kuntu, Pampangan, South Kalimantan Kalang, East Kalimantan Kalang, Toraya, Moa, and Sumbawa (Prihandini et al., 2023). Additionally, the riverine buffalo breed Silangit has also been recognized as an AnGR in Indonesia. Among these, four buffalo breeds (Gayo, Simeulue, Kuntu, Pampangan) are bred on Sumatra Island. West Sumatra Province, on Sumatra Island, has a swamp buffalo population of 30,913 heads as of 2023. Sijunjung Regency is one of the regions in West Sumatra where buffalo breeding occurs, although it is experiencing a decline in population (Rias et al., 2020; Syaiful et al., 2020).
Generally, female swamp buffaloes aged 6 years in Sijunjung Regency have an average body length of 109.50 ± 3.25 cm, a chest girth of 180.04 ± 4.05 cm, a withers height of 116.70 ± 3.28 cm, a hip height of 115.00 ± 1.91 cm, a rump width of 44.10 ± 0.74 cm, and a body weight of 327.58 ± 17.61 kg (Jesti, 2019). As a local livestock species, studying the origin lineage of swamp buffalo in Sijunjung Regency for genetic conservation and future breeding programs.
In mammals, the origin of maternal lineage can be predicted through the diversity of the cytochrome b (Cyt-b) region (Tobe et al., 2010). Cyt-b is a gene that plays a role in electron transport within the respiratory chain. It serves as a target for evolutionary analysis and species identification, which is particularly beneficial for comparing species within the same genus or family (Bruford et al., 2003). Besides, the Cyt-b gene has been used mostly for studying the evolution and genetic relationships of mammals, including buffalo (Yue et al., 2013; Sukri et al., 2014; Rusdin et al., 2020; Sun et al., 2020; Saputra et al., 2021).
In this study, the partial sequences of Cyt-b were used to identify the substitution pattern and haplotype diversity of Indonesian swamp buffalo, which are important for the genetic inventory, buffalo preservation, and improvement of the genetic potency of the buffalo population in Indonesia.
Materials and Methods
Animals and the research site
A total of 23 female swamp buffaloes were used for the study. These animals are kept by the farmers in the Villager Beeding Center at Jorong Koto Lamo Village, Muaro Takuang Urban Village, Kamang Baru District, Sijunjung Regency, West Sumatera Province of Indonesia (Fig. 1). This area is located at latitude 0˚ 18′ 43″–1˚ 41′ 46″ S and longitude 100˚ 37′ 40″–101˚ 30′ 52″ E with 500–1,000 m asl. In general, this place has 21˚C–37˚C of air temperature, 60%–100% of relative humidity, and 13.61 mm/d of rainfall.
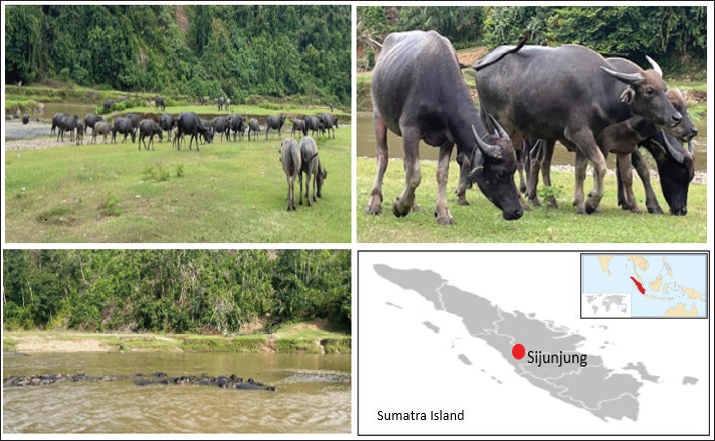

Fig. 1. A swamp buffalo herd at Sijunjung Regency, West Sumatra Province, Indonesia.
Management of the animals
The swamp buffaloes in this study were managed by the farmers under an extensive management system. The buffaloes grazed around the river from the morning (08.00 am) in the evening (17.00 pm). In addition, Brachiaria decumbens is the most common field grass around the river. However, no concentrate was given to the buffalo herds. Natural mating was managed in the buffalo herds. Regular health examination and medical treatment were performed every month.
DNA extraction and amplification
A total of 23 blood samples of swamp buffalo were collected from jugular veins using venoject and vacutainer tubes containing ethylenediaminetetraacetic acid. DNA extraction was performed using the Genomic DNA Extraction Kit (Geneaid, Taiwan) according to the manufacturer’s instructions. In this study, approximately 21.26 µl/ng of DNA concentration (λ260/280) and 1.31 of DNA purity were obtained from the DNA extraction analysis. The Polymerase Chain Reaction (PCR) reaction was carried out at a total volume of 30 µl containing 9 µl of DNA template, 0.6 µl of each primer (10 pmol), 15 µl of PCR mastermix (Bioline, USA), and 4.8 µl of nuclease-free water. A primer pair of Forward: 5’-CAT TCA TTG ACC TCC CTG CT-3’and Reverse: 5’-GCC GGA ACA TCA TAC TTC GT-3’ (Rusdin et al., 2020) was used to amplify the bubaline Cyt-b region (GenBank: AY702618) along 906 bp from 14,579th–15,484th nucleotide. The PCR program was run with one cycle of pre-denaturation at 94˚C for 5 minutes, and followed by 37 cycles of denaturation at 94˚C at 30 seconds, annealing at 59.4˚C at 30 seconds, extension at 72˚C at 30 seconds, and final extension at 72˚C at 5 minutes. The DNA was visualized by electrophoresis at 100 V for 30 minutes using 1% agarose, which was stained with FluoroSafe DNA Staining (1st Base, Singapore) above a UV Transilluminator machine (UVITEC, UK). According to the primer pair used, 906 bp of amplification was successfully amplified using 1% agarose gel (Fig. 2).
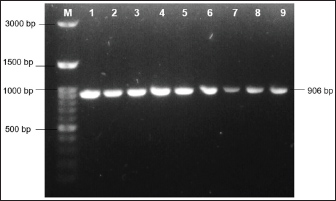
Fig. 2. Amplicons of bubaline Cyt-b along 906 bp at 1% agarose gel. Line 1–9=represented DNA samples; M=DNA ladder 100 bp.
Sequencing and analysis of data
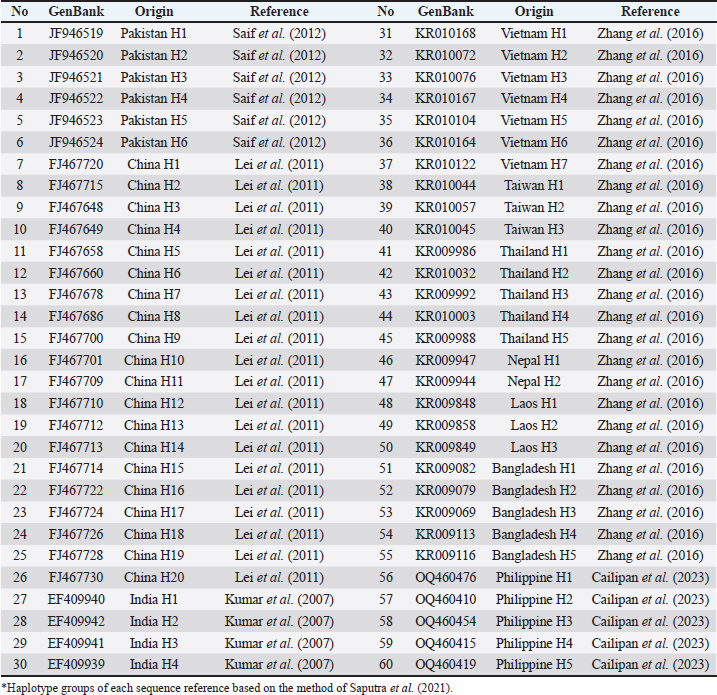
A forward sequencing analysis was executed for each amplicon by Apical Scientific (Malaysia). Sequence alignment analysis was performed using the BioEdit package (Hall, 2011). The DnaSP package (Librado and Rozas, 2009) was used to obtain the haplotype diversity (Hd) and nucleotide diversity (pi) values. An Neighbor-Joining tree was constructed with the Tajima-Nei model (10,000 × bootstrap replications) using the MEGA package (Hall, 2013). Therefore, 60 sequence references of bubaline Cyt-b were included in the phylogenetic tree (Table 1). Subsequently, the STRUCTURE package (Earl and vonHoldt, 2012) was used to determine the genetic admixture of buffaloes. The HKY+G5+F model (Hasegawa et al., 1985) was chosen to estimate the genetic divergence time of Asian buffalo because it describes the substitution pattern the best based on the Find Best DNA/Protein Models tool (Nei and Kumar, 2000) in MEGA 11 (Tamura et al., 2021). Subsequently, the BEAST package (Bouckaert et al., 2019; Bouckaert, 2022) was used with the Strict Clock model (Douglas et al., 2021). Furthermore, a Calibrated Yule model (Heled and Drummond, 2012) was constructed to show clear diverging points of every included species. The posterior tree file generated by BEAST was summarized using TreeAnnotator (Heled and Bouckaert, 2013) to produce a maximum clade credibility tree, with mean node heights and posterior probabilities assigned to each clade. Subsequently, the summarized tree performance and convergence were evaluated using Tracer v1.7.2, confirming that the analysis achieved reliable sampling of the posterior with Effective Sample Size values well above 200 for all key parameters, including clock rate and tree topology. For the BEAST analysis result, we added stratigraphy information with R software version 4.4.2.
Table 1. Sequence reference of bubaline Cyt-b region*.
Ethical approval
This research was approved by the Animal Ethics Committee of the National Research and Innovation Agency (approval issue no: 150/KE.02/SK/07/2023).
Results
Haplotype diversity
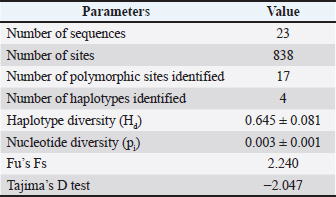
Seventeen polymorphic sites were identified within the partial Cyt-b region of the buffaloes examined, which exhibited four distinct haplotypes of sequence variation (Table 2). A total of 16 polymorphic sites were observed in the H4 animal, as indicated in Table 2. Mostly, mutation sites occurred in the H4 animal (15 sites). Therefore, the haplotype diversity (Hd) and nucleotide diversity (pi) values for the Cyt-b region in the studied animals were recorded as 0.645 ± 0.081 and 0.003 ± 0.001, respectively (Table 3). Furthermore, a positive value for Fu’s Fs statistic and a negative value for Tajima’s D test were found in the Cyt-b region of the buffaloes under investigation.
Table 2. Mutation site detection in each haplotype of swamp buffaloes (Bubalus bubalis) at Sijunjung Regency, West Sumatra, Indonesia.
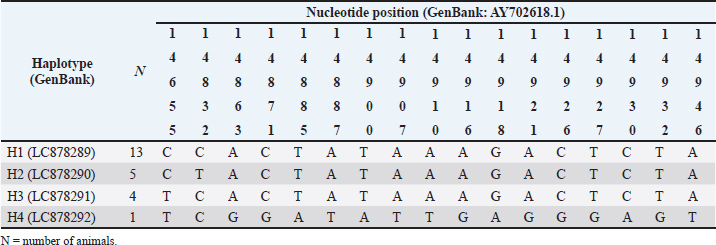
Table 3. Genetic diversity in the partial Cyt-b region of swamp buffalo (Bubalus bubalis) at Sijunjung Regency, Indonesia.
Phylogenetic tree and genetic admixture
The phylogenetic tree analysis indicated that the Sijunjung buffaloes were categorized within the Haplogroup 1 cluster (Fig. 3). Overall, the Sijunjung buffaloes exhibited a genetic composition similar to that of buffaloes from China, the Southeast region, Taiwan, and the Philippines, as demonstrated by STRUCTURE analysis (Fig. 4). Nevertheless, the Asian wild buffaloes were classified within the Haplogroup 1 cluster, showing similarity to the Sijunjung buffaloes. Based on the sequence of the Cyt-b region, the Sijunjung buffaloes were related to Haplogroup 1 buffaloes from China (Fig. 5). However, the genetic flow of Haplogroup 3 buffalo was not detected among Southeast Asian buffaloes, including those from Sijunjung. Additionally, the frequency of Haplogroup 1 buffaloes in Taiwan and the Philippines was higher than that in Thailand and Vietnam.
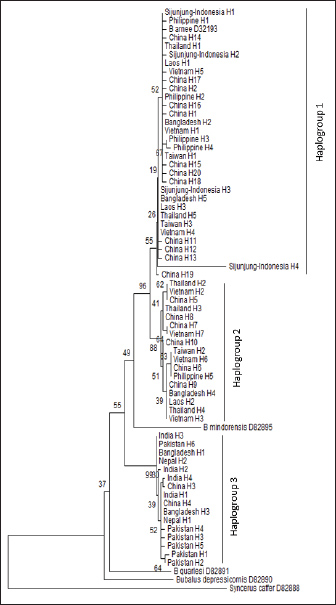

Fig. 3. Neighbor-joining tree (10,000 × bootstrap replications) among Cyt-b haplotypes of swamp buffaloes (Bubalus bubalis) from Sijunjung Regency and Asia.
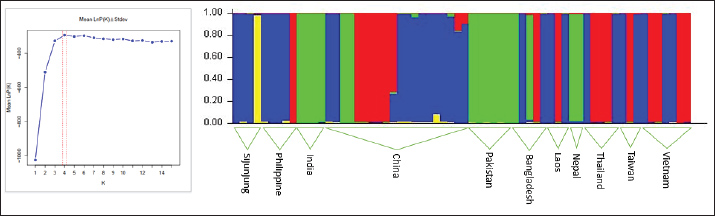
Fig. 4. The genetic admixture of the Cyt-b region in Asian buffaloes (Bubalus bubalis) is characterized in haplogroups 1 (blue and yellow), 2 (red) and 3 (green).
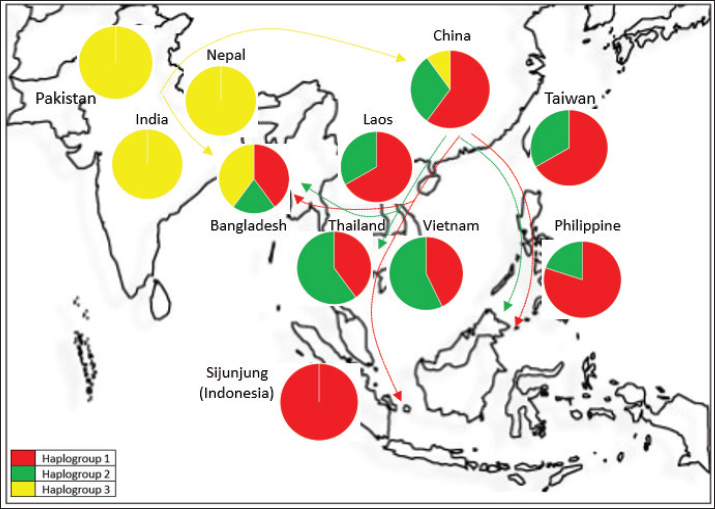
Fig. 5. Haplogroup map and genetic flow analysis of the Cyt-b haplogroup in Asian buffalo.
Genetic divergence
The genetic divergence analysis of Asian swamp buffaloes (Fig. 6) revealed that Haplotype 3 (riverine lineage) was an old cluster that existed during the early Calabrian period (1.8–0.8 mya). Haplotype 2 and Haplotype 1 were established during the last Calabrian period. Therefore, B. bubalis has been separated from Bubalus quarlesi since the last Gelasian period (2.6–1.80 mya). Interestingly, B. bubalis and Bubalus depressicornis have been separated since the middle of Tortonian (Neogene) period (11.6–7.2 mya). It can be predicted that the buffaloes migrated to the Sijunjung from 0.2 to 0.8 mya or during the middle of the Pleistocene period.
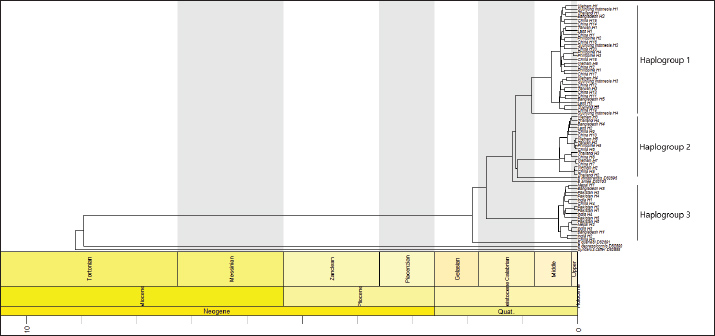
Fig. 6. Genetic divergence time of the Cyt-b haplogroup in Asian buffalo.
Discussion
The bubaline Cyt-b region in this study has several polymorphic sites. Rusdin et al. (2020) obtained nine polymorphic sites and 10 haplotypes for the Cyt-b region of Indonesian swamp buffaloes with a similar primer pair. However, Sukri et al. (2014) obtained 61 polymorphic sites and 16 haplotypes for Cyt-b region in Indonesian swamp buffaloes using a universal Cyt-b primer pair (L14841 and H15149). Sun et al. (2020) obtained 22–46 polymorphic sites in the Cyt-b region of Chinese swamp buffaloes with 9–6 haplotypes of sequence variation. Thus, the polymorphic sites in those buffaloes were found with a primer pair for 1,161 bp of amplicon length that is 15,617th–418th (circular) based on sequence reference (GenBank: AY702618.1). Subsequently, Zhang et al. (2016) identified 46 haplotypes and 59 polymorphic sites in the full Cyt-b region of the Chinese buffalo population.
Several studies have obtained similar Hd and pi values in the bubaline Cyt-b region with this study. Sun et al. (2020) reported similar findings in swamp buffaloes in Southwest China and North IndoChina regions, with Hd and pi values of 0.697 ± 0.016 and 0.003 ± 0.006, respectively. However, Yue et al. (2013) obtained a Hd value of 0.663 ± 0.101 for buffalo in Henan-China and a pi value of 0.003 ± 0.002 for buffalo in Southwestern China, which is similar to the present study. Furthermore, Zhang et al. (2016) reported similar results in the Cyt-b region of the Chinese buffalo population with the Hd and pi values of 0.637 and 0.003, respectively. The Hd value for the Cyt-b region in Indonesian buffalo was 0.659 (Rusdin et al., 2020), which is close to this study. According to Falconer and Mackay (1996), migration, selection, and inbreeding/crossbreeding can cause the genetic diversity of livestock. Nei and Kumar (2000) stated that the Hd > 0.50 is categorized as high and Hd < 0.50 as low. The pi value had three categories: low (0.01–0.04), moderate (0.05–0.07), and high (0.08–0.10). The negative value of Tajima’s D test in this study indicates an excess of low-frequency polymorphism relative to expectation (Ashfaq et al., 2014). Joshi et al. (2013) obtained a positive Fu’s Fs value in the D-loop region of Indian buffaloes, which is similar to the results of the present study. The positive Fu’s Fs statistic indicated that the investigated buffalo population was under a population bottleneck. Hence, the negative Tajima’s D value and positive Fu’s Fs value can indicate that the actual polymorphism frequency is lower than expected (Aminisarteshnizi et al., 2024).
Yue et al. (2013) previously identified three lineages (haplogroups) in Chinese buffaloes: Lineage A (swamp buffalo from the southeastern region), Lineage B1 (swamp buffalo from the central region), and Lineage B2 (Southwestern region/riverine lineage). Thus, the Lineage A Chinese buffaloes had a close genetic relationship with the Philippine and Thailand populations. Lineage A swamp buffaloes are distributed in Vietnam, Laos, Thailand, Taiwan, the Philippines, China, and Indonesia. Lineage B swamp buffaloes are not distributed in Indonesia (Cailipan et al., 2023). This argument supports this current finding that no Haplogroup 2 (Lineage B1) was observed in the Sijunjung population of Indonesia. Genetically, the buffaloes in South Asia are typical of riverine buffalo, and those in China and Southeast Asia are typical of swamp buffalo (Saif et al., 2012). In this study, a specific Cyt-b sequence, namely H4, was found in one sample of Sijunjung buffalo. Four swamp buffalo breeds (Gayo, Simeulue, Kuntu, and Pampangan) were kept on Sumatra Island and could have migrated to Sijunjung by human trade activity. Hence, the H4 buffalo may represent different matrilineal lineages of Sumatran buffalo.
During the Pleistocene era, buffaloes migrated to Indonesia through the Philippines and the Taiwan archipelagoes, known as the insular route (Tougard, 2001). Despite this, Bird et al. (2005) and De De Vos et al. (2007) argued that a migration route to Indonesia through ancient corridors via IndoChina and the Malaya Peninsula is known as the land-bridge route. Priyono et al. (2024) argued that the migration of buffalo through the insular route contains the various bubaline species in the Philippines (Bubalus mindorensis and Bubalus cebuensis) and the Wallacea region of Sulawesi (B. depressicornis and B. quarlesi). Borneo Island can be a part of the land-bridge and insular routes for buffalo migration. However, based on the mitochondrial D-loop region, the genetic admixture of the Bornean swamp buffaloes is related to the Fuling Chinese buffalo (Suhardi et al., 2023). The Cyt-b region is confirmed to cluster the Fuling buffalo in Lineage B2 (Yue et al., 2013). The haplogroup map in this study reveals a high frequency of buffalo with Haplogroup 1 in the insular route. Therefore, we argued that the haplotype 1 buffaloes can migrate across Borneo to Sumatra.
In this study, swamp buffalo migrated to Sijunjung multiple times through the Indo-China region between 0.2 and 0.8 mya. This period coincides with the era of significant glacial-interglacial fluctuations during the Quaternary. In the early Pleistocene, around 2.6 mya, alterations in the Earth's orbital patterns, influenced by Milankovitch cycles, led to temperature variations on Earth every 41,000 years (41 My) to 800 My, subsequently shifting to a cycle of every 100 My. These temperature changes resulted in the formation and melting of ice sheets in the Northern Hemisphere approximately every 1,000 years (Bintanja and Van De Wal, 2008; Sosdian and Rosenthal, 2009; Woodruff, 2010). Such fluctuations have caused substantial changes in global sea levels, with Sundaland and Sahul experiencing the most pronounced changes (Hall, 2012). During the last glacial maximum around 0.015 Mya, a vast savanna corridor was estimated to have existed from Indo-China to Java (Heaney, 1991; Bird et al., 2005). During each glacial period, this savanna corridor facilitated the movement of buffalo across the Sunda Shelf from China to Sumatra Island via land bridges and insular routes during interglacial periods, until the Sunda Shelf began to submerge and some areas became fragmented. These periodic oscillations have resulted in extensive genetic maternal introgression in the contemporary population of swamp buffalo. These findings highlight the significance of Sijunjung buffaloes as a potential reservoir of unique genetic resources with, important implications for livestock conservation and genetic management.
Study limitations
Studies that investigate the origin of an organism using mitochondrial DNA (mtDNA) sequences only reflect the evolutionary history of the maternal lineage and do not capture the entire biparental history of an organism. Despite this, genetic drift in small populations can lead to loss of mtDNA diversity, making it difficult to differentiate between individuals. Presently, next-generation sequencing can be performed to obtain the full mtDNA region of an organism and to accurately discriminate among individuals. Furthermore, in-depth study involving nuclear gene information with whole-genome sequencing can provide an accurate phylogenetic tree among individuals. Despite this, no Bornean buffalo samples in this study cannot completely explain the haplotype distribution in the migration routes.
Conclusion
The early investigation revealed that the swamp buffaloes in the Sijunjung Regency of Indonesia share a close genetic connection with the Chinese buffaloes from the Southeastern region (Lineage A), with genetic exchange occurring through land-bridge and insular pathways over the past 0.2–0.8 million years ago. A comprehensive study that includes various Indonesian swamp buffalo breeds is essential to accurately determine the origins of the swamp buffalo in Sijunjung.
Acknowledgments
The authors would like to be grateful to everybody who extended a helping hand to complete this research.
Conflict of interest
There are no conflicts of interest in this work.
Funding
This study funded by the National Research and Innovation Agency (BRIN) through the Research Project No: B-6800/III.5.5/HK.01.00/4/2023 and RKAT PTNBH Universitas Sebelas Maret fiscal year 2025 through Penguatan Kapasitas Grup Riset (PKGR-UNS) A research scheme with agreement decree No. 371/UN27.22/PT.01.03/2025
Authors’ contributions
All authors made equal contributions to the research.
Data availability
Data are available in the manuscript.
References
Aminisarteshnizi, M., Moyo, N.A.G. and Raphalo, M.E. 2024. Genetic and haplotype diversity of redbreast tilapia (Coptodon rendalli) based on cytochrome oxidase subunit I and D-loop. J. King. Saud. Univ. 36, 103585.
Ashfaq, M., Hebert, P.D.N., Mirza, J.H., Khan, A.M., Zafar, Y. and Mirza, M.S. 2014. Analyzing mosquito (Diptera: culicidae) diversity in Pakistan by DNA barcoding. PLos One. 9, e97268.
Bintanja, R. and Van De Wal, R.S.W. 2008. North American ice-sheet dynamics and the onset of 100,000-year glacial cycles. Nature 454(7206), 869–872.
Bird, M.I., Taylor, D. and Hunt, C. 2005. Palaeoenvironments of insular Southeast Asia during the Last Glacial Period: a savanna corridor in Sundaland ?. Quat. Sci. Rev. 24, 2228–2242.
Bouckaert, R., Vaughan, T.G., Barido-Sottani, J., Duchêne, S., Fourment, M., Gavryushkina, A., Heled, J., Jones, G., Kühnert, D., De Maio, N., Matschiner, M., Mendes, F.K., Müller, N.F., Ogilvie, H.A., Du Plessis, L., Popinga, A., Rambaut, A., Rasmussen, D., Siveroni, I., Suchard, M.A., Wu, C.H., Xie, D., Zhang, C., Stadler, T. and Drummond, A.J. 2019. BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLos Comput. Biol. 15(4), e1006650.
Bouckaert, R.R. 2022. An efficient coalescent epoch model for Bayesian phylogenetic inference. Systematic Biol. 71(6), 1549–1560.
Bruford, M.W., Bradley, D.G. and Luikart, G. 2003. DNA markers reveal the complexity of livestock domestication. Genetics 4, 900–910.
Cailipan, T.P.C., Paraguas, A.M., Cuanang, A.J.E., Soliven, N.F.J.A., Rono, J.G.A., Fontanilla, F.L., Servo, E.R., Cao, E.P., Fontanilla, I.K.C. and Villamor, L.P. 2023. Molecular data and karyotype revealed two distinct species of domesticated water buffaloes in the Philippines. PJS 152, 1861–1872.
De Vos, J., van den Hoek-Ostende, L.W. and van den Bergh, G.D. 2007. Patterns in insular evolution of mammals: a key to island palaeogeography. Biogeography, time, and place: distributions, barriers, and islands. Springer, 315-345.
Douglas, J., Zhang, R. and Bouckaert, R. 2021. Adaptive dating and fast proposals: revisiting the phylogenetic relaxed clock model. PLos Comput. Biol. 17.
Earl, D.A. and Vonholdt, B.M. 2012. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation. Genet. Resour. 4, 359–361.
Falconer, D.S. and Mackay, T.F.C. 1996. Introduction to quantitative genetics. Addison Wesley Longman.
Hall, B.G. 2013. Building phylogenetic trees from molecular Data with MEGA. Mol. Biol. Evol. 30, 1229 -1235. doi: 10.1093/molbev
Hall, R. 2012. Sundaland and Wallacea: geology, plate tectonics and palaeogeography. In Biotic evolution and environmental change in Southeast Asia. Gower, D., Johnson, K., Richardson, J., Rosen, B., Rüber, L. and Williams, S. (eds), Cambridge, UK: Cambridge University Press, pp: 32–78.
Hall, T. 2011. BioEdit: An important software for molecular biology. GERF Bullet. Biosci. 2, 60–61.
Hasegawa, M., Iida, Y., Yano,T., Takaiwa, F. and Iwabuchi, M. 1985. Phylogenetic relationships among eukaryotic kingdoms inferred from ribosomal RNA sequences. J. Mol. Evol. 22, 32–38. doi: 10.1007/BF02105802
Heaney, L.R. 1991. A synopsis of climatic and vegetational change in Southeast Asia. Climatic. Change. 19, 53–61.
Heled, J. and Bouckaert, R.R. 2013. Looking for trees in the forest: summary tree from posterior samples. BMC Evol. Biol. 13, 221.
Heled, J. and Drummond, A.J. 2012. Calibrated tree priors for relaxed phylogenetic and divergence time estimation. Syst. Bio. 61, 138–149.
Jesti, B.S. 2019. The body measurements of female swamp buffaloes at four regions of Sijunjung District, Sijunjung Regency, M. S. thesis, Andalas Univ., Padang, West Sumatera.
Joshi, J., Salar, R.K., Banerjee, P., S, U., Tantia, M.S. and Vijh, R.K. 2013. Genetic variation and phylogenetic relationships of Indian buffaloes of Uttar Pradesh. Asian-Australasian. J. Anim. Sci. 26, 1229–1236.
Kumar, R., Kamra, D. N., Agarwal, N. and Chaudhary, L.C. 2007. In vitro methanogenesis and fermentation of feeds containing oil seed cakes with rumen liquor of buffalo. Asian-Aust. J. Anim. Sci. 20, 1196–1200. doi: 10.5713/ajas.2007.1196
Kementan, R.I. 2024. Livestock and Animal Health Statistics 2024. Jakarta, Indonesia: Directorate General of Livestock and Animal Health, Indonesian Ministry of Agriculture.
Lei, C.Z., Zhang, C.M., Weining, S., Campana, M.G., Bower, M.A., Zhang, X.M., Liu, L., Lan, X.Y. and Chen, H. 2011. Genetic diversity of mitochondrial cytochrome b gene in Chinese native buffalo. Anim. Genet. 42, 432–436.
Librado, P. and Rozas, J. 2009. DnsSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452.
Nei, M. and Kumar, S. 2000. Molecular evolution and phylogenetic. Oxford: Oxford University Press.
Prihandini, P.W., Tribudi, Y.A., Hariyono, D.N.H., Sari, A.P.Z.N.L., Praharani, L., Handiwirawan, E., Tiesnamurti, B., Romjali, E., Matitaputty, P.R. and Wiyono, A. 2023. Biodiversity of Indonesian indigenous buffalo: first review of the status, challenges, and development opportunities. Vet. World. 16, 2217–2229.
Priyono, D.S., Solihin, D.D., Farajallah, A. and Purwantara, B. 2024. Biogeographic history of the endangered dwarf buffalo, subgenus Anoa (Bovidae: Bubalus quarlesi and Bubalus depressicornis): a perspective based on mitochondrial DNA phylogeny. Evol. Syst. 8, 311–321.
Rias, M.I., Putra, R.A. and Madarisa, F. 2020. Study on buffalo farming management system in Sijunjung Regency. Agrifo 5, 113–121.
Rusdin, M., Solihin, D.D., Gunawan, A., Talib, C. and Sumantri, C. 2020. Genetic variation of eight Indonesian swamp-buffalo populations based on Cytochrome b gene marker. Trop. Anim. Sci. J. 43, 1–10.
Saif, R., Wasim, M. and Babar, M.E. 2012. Molecular phylogeny of Pakistani riverine buffalo based on genetic variability of Mitochondrial cytochrome b gene. Mol. Biol. Rep. 39, 9707–9714.
Saputra, F., Anggraeni, A., Ishak, A.B.L., Hafid, A., Rusdin, M. and Sumantri, C. 2021. Haplotype diversity of swamp buffalo and river buffalo based on cytochrome B gene: a study of meta-analysis. Trop. Anim. Sci. J. 44, 399–407.
Sosdian, S. and Rosenthal, Y. 2009. Deep-Sea Temperature and Ice Volume Changes Across the Pliocene-Pleistocene Climate Transitions. Science 325, 306–310.
Suhardi, Wibowo, A., Putra, W.P.B. and Summpunn, P. 2023. The Phylogeny of Bornean swamp bufalo (Bubalus bubalis) analysis based on D-loop mitochondrial DNA sequence variation. Trop. Anim. Sci. J. 46, 139–145. doi: 10.5398/tasj.2023.46.2.139
Sukri, A., Amin, M., Winaya, A. and Gofur, A. 2014. Substitution and haplotype diversity analysis on the partial sequence of the mitochondrial DNA Cyt b of Indonesian swamp buffalo (Bubalus bubalis). Biol. Med. Natural Prod. Chem. 3, 59–63.
Sun, T., Wang, S., Hanif, Q., Chen, N., Chen, H. and Lei, C. 2020. Genetic diversity of mitochondrial cytochrome b gene in swamp buffalo. Anim.
Genet 51, 977–981.
Syaiful, F.L., Mundana, M. and Revar, F.H. 2020. Description and population structure of buffalo livestock of people’s in Sijunjung, West Sumatera. J. Embrio. 12, 14–22.
Tamura, K., Stecher, G. and Kumar, S. 2021. MEGA 11: molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 38(7), 3022–3027.
Tobe, S.S., Kitchener, A.C. and Linacre, A.M.T. 2010. Reconstructing mammalian phylogenies: a detailed comparison of the cytochrome b and cytochrome oxidase subunit i mitochondrial genes. PLos One 5, e14156.
Tougard, C. 2001. Biogeography and migration routes of large mammalfaunas in South ± East Asia during the Late Middle Pleistocene: focus on the fossil and extant faunas from Thailand. Palaeogeogr. Palaeoclimatol. Palaeoecol. 168, 337–358.
Woodruff, D.S. 2010. Biogeography and conservation in Southeast Asia: how 2.7 million years of repeated environmental fluctuations affect today’s patterns and the future of the remaining refugial-phase biodiversity. Biodivers. Conserv. 19, 919–941.
Yue, X.P., Li, R., Xie, W.M., Xu, P., Chang, T.C., Liu, L., Cheng, F., Zhang, R.F., Lan, X.Y., Chen, H. and Lei, C.Z. 2013. Phylogeography and domestication of Chinese swamp buffalo. PLos One 8, e56552.
Zhang, Y., Lu, Y., Yindee, M., Li, K.Y., Kuo, H.Y., Ju, Y.T., Ye, S., Faruque, M.O., Li, Q., Wang, Y., Cuong, V.C., Pham, L.D., Bouahom, B., Yang, B., Liang, X., Cai, Z., Vankan, D., Manatchaiworakul, W., Kowlim, N., Duangchantrasiri, S., Wajjwalku, W., Colenbrander, B., Zhang, Y., Beerli, P., Lenstra, J.A. and Barker, J.S.F. 2016. Strong and stable geographic differentiation of swamp buffalo maternal and paternal lineages indicates domestication in the China/Indochina border region. Mol. Ecol. 25, 1530.

















