| Research Article | ||
Open Vet. J.. 2026; 16(1): 699-710 Open Veterinary Journal, (2026), Vol. 16(1): 699-710 Research Article Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidateChristina Safira Whinie Lestari1,2, Sabar Pambudi3, Gissi Novientri1, Fithriani Fithriani1, Novaria Sari Dewi Panjaitan2, Sunarno Sunarno2, Monica Dwi Hartanti2,4 and Suprihatin Suprihatin5*1Center for Research and Development of Biomedical and Basic Health Technology, National Institutes of Health Research and Development, Ministry of Health, Jakarta, Indonesia 2Center for Biomedical Research, Research Organization for Health, National Research and Innovation Agency (BRIN), Cibinong, Indonesia 3Research Center for Vaccine and Drug, Research Organization for Health, National Research and Innovation, Agency (BRIN), Cibinong, Indonesia 4Faculty of Medicine, Universitas Trisakti, Jakarta, Indonesia 5Faculty of Health Sciences, Universitas Nasional, Jakarta, Indonesia *Corresponding Author: Suprihatin Suprihatin. Faculty of Health Sciences, Universitas Nasional, Jakarta, Indonesia. Submitted: 27/06/2025 Revised: 11/11/2025 Accepted: 01/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
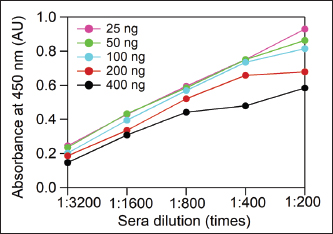
AbstractBackground: Dengue infection has increased significantly in subtropical and tropical regions in recent decades. As of 2020, 15 countries, including Indonesia, reported increasing cases, with mortality rates of approximately 0.5%–3.5%. Since 2014, the Dengue Vaccine Research Consortium under the Ministry of Health in the Republic of Indonesia has advanced a sharia-compliant dengue vaccine prototype in the form of the premembrane/envelope (prM/E) dengue virus (DENV)-1,-2,-3,-4 recombinant proteins. Aim: This study aims to develop and optimize an in-house indirect enzyme-linked immunosorbent assay (ELISA) for detecting IgG against prM/E DENV-3, serving as a preliminary antibody titer assessment for candidate dengue vaccines and potential diagnostic kits. Methods: Serum samples from mice immunized with 200 ng purified prM/E DENV-3 or 105 Plaque-forming units (PFU)/ml live DENV-3 or Phosphate-buffered saline (PBS) (negative control) were used. ELISA parameters, including antigen concentration (commercial DENV3), serum dilution, and conjugate antibody dilution, were systematically optimized. Optimal conditions were determined to be 100 ng antigen, 1:200 serum dilution, and 1:5,000 conjugate-Ab dilution. Results: Seropositive samples yielded absorbance values exceeding the established cutoff. The statistical analysis showed significant differences between groups (p < 0.001). Conclusion: The optimized in-house ELISA reliably detects IgG responses to recombinant prM/E DENV3, supporting its utility in early-stage evaluation of dengue vaccine candidates and in the development of diagnostic tools. This study is a successful stage supporting the development of prM/E DENV-1,-2,-3,-4 as a candidate dengue vaccine. Keywords: In-house ELISA, prM/E DENV-3, Dengue vaccine, Method optimization. IntroductionIn recent decades, dengue infection has increased in subtropical and tropical areas. Studies have shown that 390 million cases of dengue virus (DENV) infection occur annually, with 96 million cases manifesting severe clinical symptoms (Bhatt et al., 2013). The global incidence of dengue has dramatically increased in recent decades from 505,430 cases in 2000 to 14.6 million in 2024 (WHO, 2025). Another study showed that 3.9 billion people from 129 countries are at risk of contracting the DENV, and 70% of them are in the Asian region (Brady et al., 2012; Bhatt et al., 2013). Until 2020, 15 countries, including Indonesia, have reported increasing numbers of dengue cases. DENV is a member of the Flaviviridae family from the Flavivirus genus. Its morphology is spherical with a diameter of approximately 50 nm and is covered with a lipid membrane [envelope (E)] with a thickness of approximately 10 nm on the outside. The nucleocapsid inside the virus has a diameter of approximately 30 nm and contains a single-stranded positive-sense RNA genome of approximately 11 kilobases in length (Kuhn et al., 2002; Yuen et al., 2025). A recent study showed that the DENV E protein domain I interacts synergistically with the kringle-4 and kringle-5 domains of plasminogen, highlighting its important role in viral pathogenesis and host interactions (Yuen et al., 2025). Female Aedes aegypti and Aedes albopictus mosquitoes could transmit the DENV. The clinical manifestations of symptomatic dengue infection are differentiated based on their severity into dengue fever (DD), dengue hemorrhagic fever (DHF), and dengue shock syndrome (Jiang et al., 2013; Simmonds et al., 2017). This infection is a contribution by four serotypes of (DENV-1, -2, -3, and -4) that are known to be circulating in Indonesia. The death rate from this virus reaches 0.5%–3.5% (Lestari et al., 2017; Kementerian Kesehatan RI, 2017). In accordance with the 2020–2024 strategic plan of the Ministry of Health in the Republic of Indonesia, the prevention and control of vector-borne and zoonotic diseases in each district/city must fulfill the DHF <49 per 100,000 population by 90% (Kementerian Kesehatan, 2020). In previous years, the strategy relied mainly on reactive outbreak response and conventional vector control measures, which have shown limited long-term effectiveness and highlight the ne°°ed for more sustainable and integrated approaches. A new strategy is needed to achieve this goal. Vaccination is one of the most promising approaches to preventing dengue infection. Dengvaxia, the first dengue vaccine developed by Sanofi Pasteur and licensed in 2015, has low efficacy (56%) and produces severe clinical manifestations in seronegative individuals. Therefore, the Dengvaxia® vaccine is only aimed at populations of endemic areas with dengue prevalence of more than 80% (Chaturvedi et al, 2005). The newest dengue vaccine, which had been approved recently by the National Agency for Drug and Food Control (BPOM, Indonesia) and was being reviewed to be used in the European Union, is QDENGA® (TAK-003), whose usage is dedicated for 6 to 45 year-old individuals (Osorio et al., 2015; Saito and J, 2022). Although QDENGA® has been approved for use in several countries, data on long-term safety and effectiveness in real-world settings remain under evaluation through postmarketing surveillance programs. DENV consists of three structural proteins (capsid, membrane, and E proteins) located at the N-terminus and seven nonstructural proteins nonstructural 1 (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5) encoded at the C-terminus (Costa et al., 2011; Lestari et al., 2016; Fajardo-Sánchez et al., 2017; Zonetti et al., 2018). Premembrane (PrM), E, NS1, and NS3 proteins in DENV are known to have immunogenic properties and have been evaluated for their ability as vaccine candidates (Rothman, 2004; Konishi et al., 2006). Only protein E can induce a humoral immune response in the form of neutralizing antibodies. Protein E itself can be the main protective mechanism to avoid the formation of ADE in the severity of DENV pathogenesis caused by sub-neutralizing antibodies (Pengsaa et al., 2011; Zheng et al., 2011). Since 2014, the Dengue Vaccine Research Consortium under the Ministry of Health in the Republic of Indonesia has developed a prototype of a sharia-compliant dengue vaccine candidate in the form of the prM/E DENV-1,-2,-3,-4 recombinant protein subunit (Marbawati and Wijayanti, 2014). DENV-2 and DENV-3 have been proven to cause higher severity of dengue than that caused by DENV-4, where DENV-4 only caused milder symptoms. In 2019, a study showed that 75.9% of the dominant DENV-3 serotype was the cause of the DHF epidemic in Manado, Indonesia (Tatura et al., 2021). The enzyme-linked immunosorbent assay (ELISA) method can be used to determine the anti-dengue IgG titer and its ability to bind the protein subunits or the whole DENV virus as antigens (Lazo et al., 2014; Liu et al., 2014; Gil et al., 2016). Previous approaches to detect anti-dengue antibodies have relied on commercial ELISA kits using whole-virus antigens, which often lack serotype specificity and produce high background signals (Marfia et al, 2025). These limitations can compromise the accurate assessment of immune responses to recombinant subunit vaccine candidates, particularly in regions with cocirculating flaviviruses. To address this limitation, this study aimed to develop and optimize an in-house indirect ELISA system for evaluating vaccine effectiveness, with a focus on detecting IgG responses against the prM/E DENV-3 candidate vaccine. A recombinant DENV-3 E protein was employed as the coating antigen to ensure specificity in detecting antigen-induced responses in murine sera. The optimized assay may also serve as a model for extending the same approach to other serotype-based vaccine candidates using their respective RAs. Materials and MethodsSamplesResearch samples were obtained from the Dengue Vaccine Research Consortium under the Ministry of Health, Republic of Indonesia, in the form of sera collected from Balb/c mice. The sera were divided into three groups: group 1, mice immunized with 200 ng of purified prM/E DENV-3 protein; group 2, mice immunized with 1 × 105 PFU/ml live wild-type DENV-3 virus; and group 3, negative controls injected with PBS. Positive control sera were obtained from a separate batch of mice immunized with live wild-type DENV-3 virus, which had been previously confirmed to be seropositive and consistently produced high absorbance values in the ELISA. All sera samples were stored in aliquoted form at 20°C to avoid repeated freeze–thaw cycles, with a maximum storage duration of 4 months before use. Protein-based vaccine formulation and immunizationRecombinant protein prM/E DENV-3 was a genetically engineered product of the DENV-3 of the Jakarta 2010 strain, developed by the National Dengue Vaccine Consortium Research and available in the National Institute of Research and Development Laboratory, Indonesia. Recombinant protein prM/E DENV-3 was developed with an adjuvant before immunization. For the primary dose, 200 µg of recombinant protein was emulsified with complete Freund’s adjuvant (1:1 v/v) in a final volume of 100 µl per mouse. Booster doses were prepared with the same amount of protein emulsified with incomplete Freund’s adjuvant (1:1 v/v). Three doses were administered via the intraperitoneal route, with 3-week intervals between immunizations. Preparation and immunization of live virus vaccineThe DENV-3 of Jakarta 2010 strain was propagated in Vero CCL-81 cells maintained in Advanced Roswell Park Memorial Institute (RPMI) medium 1,640 medium (Gibco-Invitrogen) supplemented with 1% fetal bovine serum, 2 mM L-glutamine, and 1× antibiotic/antimycotic (Gibco-Invitrogen). Vero cells were subcultured using 1× TrypLE Select (Porcine-free, Gibco-Invitrogen) instead of trypsin. Cultures were incubated at 37°C with 5% CO2 and inspected daily for cytopathogenic effect OR cytopathic effect (CPE). Viral supernatants were harvested either upon the appearance of CPE or at day 7 post-infection, clarified by centrifugation at 3,000 × g for 10 minutes at room temperature, and concentrated to a final volume of 3–4 ml using Amicon Ultra 100 kDa filters. The concentrated virus was aliquoted and stored at −80°C until further use. Viral titers were determined using the plaque assay. For immunization, the viral stock was diluted in sterile phosphate-buffered saline (pH 7.4) to obtain 1 × 105 PFU per dose. Mice were immunized with 100 µl intraperitoneal injections of live virus. Three doses were administered at 3-week intervals. Optimization of the indirect ELISA protocolVarious antigen concentrations (25, 50, 100, and 200 ng/well) were tested based on standard coating ranges commonly used for recombinant proteins to determine the optimal ELISA conditions. Serial dilutions of serum samples (1:100–1:800) and secondary antibody (1:5,000–1:20,000) were evaluated to identify the conditions that offered the highest signal-to-background ratio. The rationale was to use the minimal amount of antigen and antibody dilutions necessary to maintain high assay sensitivity, specificity, and low background, while being cost-effective for downstream applications. Each well (96-well plate Maxisorp, Nunc) was coated with 100 µl antigen of E DENV-3 recombinant protein (Mybiosource MBS596112) at concentrations of 25, 50, 100, and 200 ng/well per 100 µl coating buffer and incubated overnight at 4°C. The coating buffer consisted of 0.015 M Na2CO3 and 0.035 M NaHCO3 dissolved in 1 l of sterile distilled water, adjusted to pH 9.5. After incubation, it was washed three times with 200 l of 0.01 M phosphate-buffered saline + 0.05% Tween 20 (PBS-T) 200 µl three times. Blocking buffer (bovine serum albumin, BSA 3%) was added 100 µl/well and incubated for 1 hour at 37°C. The washing step was performed again with PBS-T 0.05%, 200 µl, three times. Each well was serially diluted with 100 µl of sera from each group (1:200; 1:400; 1;800; 1:1,600; 1:3,200) in PBS 1× and incubated for 1 hour at 37°C. Wash again with 200 l of PBS-T 0.05%, 200 µl three times. IgG binding was detected using goat anti-mouse IgG-peroxidase conjugate (Cat# A4416, Sigma Aldrich). Secondary antibodies were diluted at a ratio of 1:5,000, 1:10,000, and 1:20,000 in 0.05% PBS-T. Plates were then incubated in the dark at 37°C for 1 hour, covered with aluminum foil. Wash again with the same buffer three times. Add a 3,3′,5,5′-tetramethylbenzidine (TMB) substrate (Thermo Scientific, Cat# 34021) and incubate at room temperature for 15 minutes. The reaction was stopped by adding 50 µl of 2 M H2SO4. The absorbance was measured using an ELISA reader at a wavelength of 450 nm. A positive result was defined based on a cut-off value calculated as the mean absorbance of the negative control group (PBS-injected mice) plus two standard deviations (Mean + 2SD). This approach minimizes false positives by accounting for background variability in negative samples. Statistical analysisData processing and analysis were performed using SPSS version 21.0. A normality test was performed on all data using the Kolmogorov-Smirnov test. An independent parametric t-test analyzed the normally distributed and homogeneous data to show the significance between groups. Ethical approvalThis research received ethical approval from the Health Research Ethics Committee, National Institutes of Health Research and Development (approval number: LB.02.01/2/KE.189/2021) dated April 16, 2021. ResultsAntigen concentration optimization and sample dilutionThe indirect ELISA was optimized to determine the most effective antigen concentration and serum dilution that provided high sensitivity and specificity while minimizing non-specific binding. Therefore, the plate was coated with various concentrations of DENV-3 E recombinant protein as an antigen (25, 50, 100, 200, and 400 ng/well). For blocking, different agents, including 5% non-fat dry milk and 3% BSA, were evaluated. Although non-fat milk is commonly used, it resulted in slightly higher background reactivity in murine sera. Therefore, 3% bovine serum albumin was selected as the optimal blocking agent for this assay. Serum samples from DENV-3-seropositive mice were tested at serial dilutions of 1:200–1:3,200. The dilution range was selected considering the limited serum volume available from each mouse, since the same samples were also required for Plaque reduction neutralization test (PRNT) assays. The conjugate dilution was fixed at 1:5,000 during this optimization stage. The goal of this study was to identify the optimal balance between reagent use and signal-to-background performance. The parameter to determine our optimal condition is the highest OD450 ratio from positive samples to negative samples. The negative control samples consistently showed negligible absorbance across all conditions (OD450=0.047), confirming low background reactivity. Among the tested serum dilutions, 1:200 produced the highest absorbance value for the positive samples while maintaining a low background, making it the optimal dilution. Antigen concentrations of 200 and 400 ng showed saturation at a dilution of 1:200. Therefore, 100 ng/well was selected as the optimal antigen concentration due to its strong and specific signal, while minimizing the reagent use. In this context, the term optimal refers to the condition that maximizes sensitivity for distinguishing seropositive from seronegative samples, thereby reducing the likelihood of false-negative results. We acknowledge that a 1:200 dilution lies within the saturation plateau of the curve, which may limit the ability of the assay to discriminate subtle differences in antibody titers across samples. For semi-quantitative or quantitative measurements of antibody responses, dilutions within the linear range of the dose-response curve (1:800–1:600) would provide better resolution. Nevertheless, for this optimization, which aimed to establish a robust in-house ELISA for qualitative detection of anti-DENV-3 antibodies, 1:200 was considered the most suitable condition (Fig. 1).
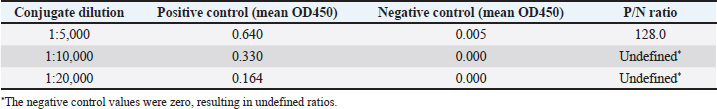
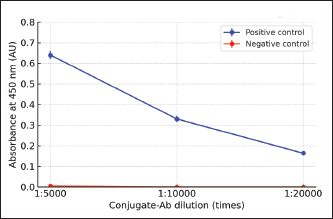
Fig. 1. Antigen concentration and sample dilution optimization. The sample used was seropositive DENV-3 post-vaccination in mice. Variations in antigen concentration are 25, 50, 100, 200, and 400 ng/well. Sera dilution was performed from 1:200 to 1:3,200. The negative control showed minimal absorbance (OD450=0.047) across all conditions and was therefore not plotted due to overlap with the baseline. Note: At a dilution of 1:200, the assay reached maximal absorbance values, which were defined here as the ‟optimal” point for seropositivity detection. For quantitative discrimination of antibody titers, dilutions within the linear range (1:800–1:600) would be preferable. Optimization of the conjugate antibody dilutionA serum dilution of 1:200 and an antigen concentration of 100 ng were used to determine the optimum conjugate dilution. Based on the comparison of positive-to-negative (P/N) ratios, the conjugate dilution of 1:5,000 was selected as the optimum point. At this dilution, the mean OD450 values of the P/N controls were 0.64 and 0.005, respectively, resulting in a P/N ratio of 128. At higher dilutions (1:10,000 and 1:20,000), the negative control values were zero, which yielded undefined P/N ratios, whereas the positive control absorbance values decreased to 0.33 and 0.164, respectively. These results indicate that the 1:5,000 dilutions not only provided the highest measurable P/N ratio but also maintained a sufficiently strong signal, justifying its use as the optimal conjugate antibody dilution. As summarized in Table 1, the 1:5,000 dilution yielded the highest P/N ratio, indicating the best balance between signal intensity and background reduction. This dilution was selected for subsequent ELISA assays. Table 1 shows the calculated P/N ratios for all dilutions. Importantly, all measurements were performed in triplicate, and error bars representing standard error of the mean were included in Figure 2 to illustrate the assay’s reproducibility and reliability.
Fig. 2. Optimization of the conjugate antibody dilution. The samples used were seropositive DENV-3 post-vaccination in mice (positive control) and negative controls (PBS treatment). The conjugate dilutions ranged from 1:5,000 to 1:20,000. Data represent mean OD450 values (n=3 technical replicates) with standard deviations indicated by error bars. The 1:5,000 dilution yielded the highest measurable P/N ratio (128), supporting its selection under the optimum condition. Table 1. Mean OD values of the P/N controls at different conjugate antibody dilutions and corresponding P/N ratios.
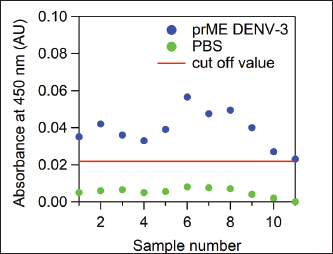
The 1:5,000 dilution yielded the strongest specific signal with minimal background, as reflected in the highest OD450 ratio between the P/N samples. The parameter to determine our optimal conditions is the highest OD450 ratio from positive samples to negative samples. Based on this, a conjugate dilution of 1:5,000 was chosen as the optimum concentration. Detection of IgG antibodies against prM/E DENV-3 and live wild-type virus DENV-3Detection of IgG anti-dengue prM/E DENV-3 was performed based on the ELISA optimization results using a sample dilution of 1:200, a coating antigen concentration of 100 ng, and a conjugate antibody dilution of 1:5,000. The cutoff value, calculated as 2 × (mean + 2 SD) of the prevaccinated sera, was 0.022. All serum samples in the group of mice that received prM/E DENV-3 vaccination showed absorbance values above this cut-off, with a mean absorbance of 0.039. In contrast, all samples from the negative control group (PBS-injected mice) had absorbance values below the cut-off value, with a mean absorbance of 0.0057 (Fig. 3). These results indicate that the optimized in-house ELISA system is technically functional under the tested conditions. However, the relatively low OD450 values obtained in this study should be interpreted with caution, as OD450 values below 0.04 are generally considered negative when using a TMB substrate. Therefore, further validation using well-accepted methods, such as commercial ELISA kits or virus neutralization assays, to confirm the antibody responses is required.
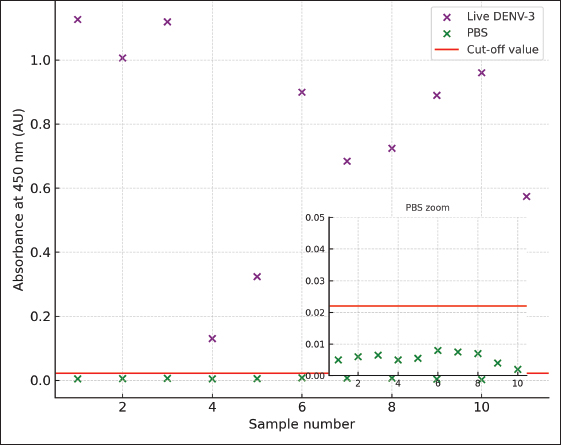
Fig. 3. The results of IgG anti-dengue detection using the optimized in-house ELISA method on the seropositive mice injected with prM/E DENV-3 recombinant protein (blue dots) and the seronegative mice injected with PBS (green dots). The cutoff value is two times above the mean + 2SD of the absorbance value of pre-vaccinated sera. IgG anti-DENV-3 wt was detected using the optimized in-house ELISA system. The results showed a mean absorbance value of 0.777 (SD=0.3056), all of which remained above the cut-off value, indicating that the optimized ELISA system can reliably discriminate between P/N samples. In contrast, the sera from PBS-injected control mice remained below the cut-off threshold. To enhance the visualization of the differences in the low-absorbance range, particularly for the PBS control group, an inset plot was used to magnify the low-range OD450 values relative to the cutoff (Fig. 4). This allows for a clearer and more accurate interpretation of the background reactivity in the negative controls.
Fig. 4. IgG anti-dengue detection using the optimized in-house ELISA. Mice inoculated with live wild-type DENV-3 (purple dots) showed consistently higher absorbance values than PBS-injected control mice (green dots). An inset plot is provided for the PBS control group to enhance visualization of the low-range absorbance. The cut-off value (red line) was determined as twice the mean + 2SD of pre-vaccinated sera absorbance values. Data processing and analysis were performed using SPSS version 21.0 and visualized using GraphPad 9.1.0. All data were distributed normally and passed the Kolmogorov-Smirnov test with p >0.1000 (Fig. 5a). All data were analyzed using the independent parametric t-test to assess statistical significance between groups. Significant differences (p <0.0001) were detected between seropositive prM/E DENV-3 samples (n=12) and seronegative PBS controls, as well as between seropositive live wild-type DENV-3 samples (n=11) and seronegative PBS controls. Notably, the live DENV-3 group displayed higher inter-animal variability than the prM/E DENV-3 group, showing heterogeneity in antibody responses following live virus exposure. These results indicate that the in-house ELISA system can reliably distinguish between seropositive and seronegative samples (Fig. 5b). Although a statistically significant difference was observed between the PrM/E DENV-3 and negative control groups (p <0.0001), OD450 values for both groups remained within the assay’s background range (<0.0001). Therefore, this result is not considered biologically significant, and the PrM/E DENV-3 immunized group is interpreted as seronegative in this context.
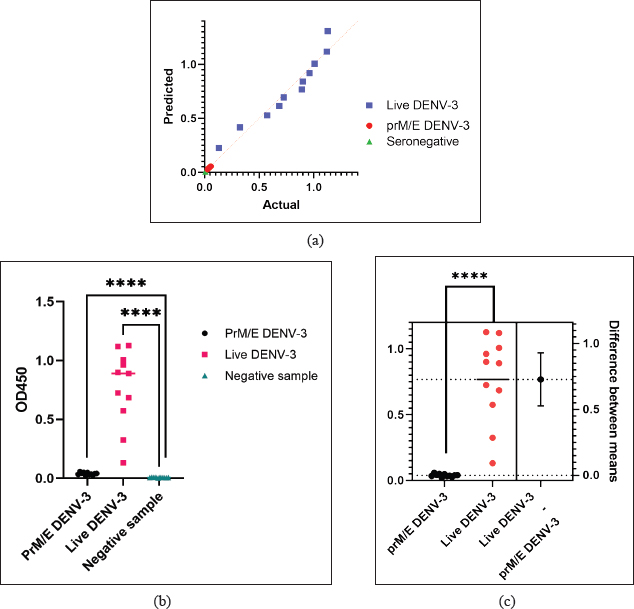
Fig. 5. (a): Q-Q plot demonstrating the normal distribution of all sample groups, confirmed by the Kolmogorov-Smirnov (K-S) test. (b): Comparison of IgG anti-dengue titers among seronegative samples (n=10), seropositive prM/E DENV-3 immunized mice (n=11), and seropositive live DENV-3 wt immunized mice (n=11). Data are presented as mean ± standard deviation. (c): Estimation plot showing the difference in mean IgG anti-dengue titers between mice immunized with prM/E DENV-3 (n=11) and live DENV-3 wt (n=11). The difference between group means with standard deviation is shown to emphasize the effect size and statistical confidence in comparing the antibody responses. This plot format emphasizes the effect size and statistical confidence by displaying the mean difference (SD). The data were analyzed using an independent t-test (assuming normal distribution). Asterisks (****) indicate p <0.0001. Significant differences in IgG anti-dengue titers were observed between the prM/E DENV-3 immunized group and the live wild-type DENV-3 virus immunized group (p <0.0001). The results demonstrate that the optimized in-house ELISA system can reliably distinguish between antibody responses elicited by different antigenic exposures. Figure 5c illustrates the comparative effect sizes and their corresponding confidence intervals, which presents the difference between means format to highlight the magnitude and statistical confidence of group differences and directly reflect the extent of serological response and assay specificity. DiscussionDD, caused by DENV, remains a major acute infectious disease with cases continuously reported worldwide. Despite the national prevention and control programs established by the Indonesian Ministry of Health, the incidence of dengue remains high (Kementerian Kesehatan RI, 2017). Dengvaxia®, the first licensed dengue vaccine, has demonstrated contraindications in children aged 9 years, emphasizing the urgent need for safer and more broadly applicable vaccine alternatives (Guy et al., 2017). Several vaccine candidates have been reported in recent reviews (Dighe et al., 2019), but most remain at the early stages of development and require further evaluation. More recently, QDENGA® (TAK-003) was approved for use in Indonesia, although its nationwide impact requires rigorous monitoring and time to generate comprehensive surveillance data. In addition, a prototype of a sharia-compliant dengue vaccine, based on recombinant prM/E DENV-1–4 proteins, has been developed but remains insufficiently studied (Marbawati and Wijayanti, 2014). These gaps underscore the need for reliable preclinical evaluation platforms. Although several commercial ELISA kits are available for dengue IgG detection, they are primarily intended for clinical use in humans and generally employ whole-virus antigens or pan-serotype E proteins. Such designs limit their utility for experimental vaccine studies using animal sera and hinder the evaluation of antibody responses against specific RAs (Medina et al., 2024). In this study, we used sera from mice immunized with a recombinant prM/E DENV-3 vaccine candidate. Commercial kits are not validated for murine samples and do not target prM/E-specific IgG; therefore, an in-house indirect ELISA was required. Establishing assay conditions tailored to RAs and murine sera ensured antigen specificity and compatibility with the experimental system. Similar approaches have been reported for dengue and other viral infections, highlighting the value of optimized recombinant antigen-based ELISA for sensitive and specific antibody detection (Feng et al., 2020; Marfia et al., 2025). The recombinant DENV E protein was used as the coating antigen in the ELISA. The prM protein, consisting of approximately 166 amino acids, mainly contributes to viral assembly and maturation and is not a primary antigenic determinant for antibody recognition. The E protein (~354 amino acids) is the major structural component exposed on the viral surface and is well established as the principal target of dengue-specific binding and neutralizing antibodies. Recent studies have consistently demonstrated that antibodies generated during natural dengue infection or vaccination predominantly target the E protein, supporting its diagnostic value. Durham et al. (2019) showed that broadly reactive antibodies from dengue patients recognize conserved regions of the E protein (Durham et al., 2019). Similarly, Uno and Ross (2021) reported that E-based antigens elicit strong cross-serotype IgG responses, while Yen et al. (2023) identified novel epitopes in the E protein suitable for diagnostic applications (Uno and Ross, 2021; Yen et al., 2023). These findings reinforce the rationale for using the E protein as the antigen in ELISA-based detection of IgG against DENV-3. The recombinant antigen used represents only the DENV E protein rather than the combined prM/E construct. However, as the E protein constitutes the major antigenic determinant and the primary target of DENV-specific antibodies, its use remains highly relevant for the development of ELISA-based diagnostic assays. In this study, recombinant prM/E protein and live DENV-3 virus immunizations were designed to evaluate antibody responses in a preliminary vaccine development setting. The dosing strategy aimed to ensure detectable IgG and neutralizing antibody responses, with recombinant protein requiring higher antigen input in line with previous preclinical dengue vaccine reports. A limitation of this study is that doses were not normalized to equivalent prM/E antigen content between the recombinant protein and live virus groups. In this preliminary evaluation, the recombinant protein dose was chosen to ensure measurable neutralizing antibody responses, but this difference constrains the direct quantitative comparison of IgG titers. Future studies will incorporate antigen quantification and normalization to allow rigorous head-to-head comparisons. In some experimental cases, the optimization of a certain detection to increase the specificity and efficiency is required. In recent studies, an in-house Dot-ELISA rapid test for the detection of Toxoplasma gondii infection (Teimouri et al., 2019), a recombinant nucleoprotein-based indirect ELISA for Crimean-Congo hemorrhagic fever virus-specific antibodies (Shrivastava et al., 2019), and even helps to solve the problem faced clinically for detecting the antibody form patients receiving Tumor necrosis factor alpha (TNF-α) inhibitors (Ogrič et al., 2019). Besides, there were also changes and validations performed in some previous studies to improve the detection while still using the available commercial ELISA kit (Gono et al., 2019). Until now, there is no available gold standard for detecting the anti-dengue antibody due to the recombinant protein antigen. Previous studies showed that the Immunochromatographic test assay, a relatively cost-effective assay, showed low sensitivity for anti-dengue IgM and IgG, whereas the ELISA system has limitations for rapid detection (Moorthy et al., 2009; Murugananthan et al., 2018). Regarding this observation, a note to the clinical laboratories had been addressed in a previous scientific communication letter (Lee et al., 2019). The optimization of serum dilution was carefully considered in this study due to the limited volume of mouse serum available, as the same samples were also utilized for PRNT, which is the gold standard for functional neutralizing antibody measurement. Serial dilutions of 1:3,200, 1:1,600, 1:800, 1:400, and 1:200 were tested. The 1:200 dilution provided the most favorable balance between sensitivity and specificity, whereas higher dilutions, particularly in low-titer sera, substantially reduced the signal. Although we acknowledge that very low antibody titers may still be challenging to detect, the use of recombinant antigens and optimized coating concentrations enhanced assay sensitivity at this dilution. Furthermore, we performed antigen concentration optimization at 25, 50, 100, and 400 ng, and 100 ng was ultimately selected because it provided the most consistent performance with reduced background in negative sera, although lower concentrations yielded relatively higher absorbance values. We acknowledge that a broader comparative analysis across multiple antigen concentrations could provide further insights; however, the present work primarily focused on establishing a standardized and reliable in-house ELISA protocol. Future studies may expand on this by systematically evaluating the efficiency of antigen usage to refine assay performance. A previous report also highlighted that selecting the appropriate antigen concentration is critical for maximizing the signal-to-noise ratio while avoiding excessive background or plate saturation, supporting our rationale for selecting 100 ng as the optimal condition (Alandijany et al., 2020). We also considered the use of a blocker and a substrate as an important stage. There are several types of protein blockers and substrates available for ELISA, and each blocker and substrate has its own advantages and disadvantages. Different blocking agents were compared. Although 5% non-fat dry milk in PBS/TBS with Tween-20 is a standard choice, it generated a higher background in negative murine sera in our system (data not shown), possibly due to casein-derived protein interactions. However, Bovine serum albumin (BSA) was chosen to be used as the protein blocker in this study because it has been proven to be a well-working blocker that successfully blocks non-specific protein-surface bindings (Ahirwar et al., 2015). In addition, BSA is a relatively cost-effective and hassle-free storage reagent. Compared with other types of protein blockers, such as non-ionic detergent blockers, which still facilitate non-specific binding, BSA showed much higher sensitivity (Hakami et al., 2015). For the substrate, TMB was chosen as the substrate for this ELISA system optimization. TMB has been demonstrated to have far higher sensitivity, even for the detection of tiny concentrations in picograms, and faster reaction rate than other types of ELISA system substrates, such as ABTS (2,2’-azino-di[3-ethylbenzthizoline]), ONGP (ortho-nitrophenyl-β-galactoside), and OPD (O-phenylenediamine dihydrochloride) in previous studies (Sanders et al., 2016; Minic and Zivkovic, 2021). In addition, TMB is safe for routine laboratory use, whereas OPD is cancerogenic (Minic and Zivkovic, 2021). We also observed strong significant differences between seropositive prM/E DENV-3 and seronegative PBS controls, as well as between seropositive live wild-type DENV-3 and controls, confirming that our optimized in-house ELISA can robustly differentiate seropositive from seronegative sera. However, the live DENV-3 group exhibited substantially greater inter-animal variability in antibody titers than the prM/E group. This elevated heterogeneity likely reflects biological diversity in immune responses triggered by live virus, possibly driven by differences in viral replication kinetics, host innate immune activation, and individual genetic or microbiota influences. In contrast, immunization with recombinant prM/E antigen leads to more homogeneous antibody responses, consistent with a more uniform antigen exposure without active replication. Importantly, such interindividual variability is consistent with observations in murine dengue models (e.g., non-lethal DENV-3 infection in AG129 mice) (Sarathy et al., 2018), as well as recognized influences of genetic, environmental, and microbiome factors on immune variability in mice (Enriquez et al., 2020). These findings underscore that the biological variability among animals must be considered when interpreting serological endpoints and evaluating immunogenicity in vaccine or challenge studies. Although additional controls, such as irrelevant antigens or higher serum dilutions, were not included in this initial optimization study, the use of nonimmunized sera and PBS blanks demonstrated low background binding, supporting assay specificity. Future studies will expand on this by including broader specificity assessments, including cross-reactivity analyses and irrelevant antigens. In this study, we established and optimized an in-house indirect ELISA for the detection of IgG anti-dengue prM/E DENV-3 antibodies in mice. The results demonstrated that under controlled experimental conditions, the optimized assay could reliably distinguish between seropositive and seronegative samples. However, this work represents a methodological optimization rather than the development of a validated diagnostic assay. Comprehensive validation, including sensitivity, specificity (particularly cross-reactivity with other flaviviruses), and assay reproducibility assessments, is required before this approach can be applied as a diagnostic tool. Therefore, our findings should be considered as proof of concept optimization that may provide a basis for future diagnostic validation studies. A limitation of this study is that the analytical sensitivity of the assay was not evaluated by spiking seronegative murine sera with a well-characterized anti-DENV-3 E protein antibody. Such an approach would allow the determination of the detection limit and provide a more comprehensive assessment of assay performance, which we plan to address in our future work. ConclusionBased on our results, the in-house indirect ELISA system was successfully optimized in terms of sample dilution, antigen concentration, and conjugate antibody dilution. This optimization addressed the limitations of a commercial ELISA kit. The optimized in-house system provides a protocol for detecting preM/E DENV-3 IgG antibodies using mouse sera. Although this study was limited to the DENV-3 E protein, the findings may serve as a technical reference for future adaptations to other serotypes. However, we refrain from making generalized claims regarding tetravalent applicability until further data are generated. We encourage future research to extend this approach to other DENV serotypes, to explore additional diagnostic biomarkers, and to refine standardized methods for reliable anti-dengue IgM and IgG detection using mouse sera. AcknowledgmentsAll authors would like to extend our gratitude to Dr. Vivi Setiawaty, M. Biomed, as Head of the Center for Research and Development of Biomedical and Basic Health Technology, National Institute of Health Research and Development, Indonesia Ministry of Health, who provided us with funding and allowance for performing the experiments in the Immunology Laboratory in the Prof. Dr. Sri Oemijati Infectious Disease Research Laboratory. In addition, many thanks should be conveyed to Dr. Neni Nuraeny and her team from the R&D division of PT. Biofarma which prepared the prM/E DENV-3 recombinant protein antigen for vaccination in mice. We would like to thank Dr. Tjahjani Mirawati Soediro, Beti Ernawati Dewi, Ph. D., Dr. Fithriyah, and the team who carried out the mouse vaccination procedure and facilitated the laboratory of experimental animals at the Department of Microbiology of the Faculty of Medicine, University of Indonesia. All authors would also like to thank the colleagues in the Immunology and Virology Laboratory of the Center for Research and Development of Biomedical and Basic Health Technology, NIHRD. Conflict of interestAll authors declare no conflicts of interest. FundingThis study was funded by the Center for Research and Development of Biomedical and Basic Health Technology, NIHRD, Indonesia Ministry of Health, with Decision Letter No. HK.02.03/1/354/2021. Authors’ contributionCSWL, GN, and Suprihatin conceived and designed the study. GN and Fithriani performed most of the experiments. CSWL, Sunarno, NSDP, and GN analyzed the data. CSWL, GN, and NSDP wrote the first draft of the manuscript. Sunarno, MDH, and SP provided the required corrections and comments. CSWL, NSDP, and Suprihatin revised the manuscript. All authors have read and approved the final version of the manuscript. Data availabilityThe information supporting the discoveries in this regard is accessible in the revised manuscript. ReferencesAhirwar, R., Bariar, S., Balakrishnan, A., and Nahar, P. 2015. BSA blocking in enzyme-linked immunosorbent assays is a non-mndatory step: A perspective study on mechanism of BSA blocking in common ELISA protocols. RSC Adv. 5, 100077–100083. doi: 10.1039/c5ra20750a Alandijany, T.A., El-Kafrawy, S.A., Tolah, A.M., Sohrab, S.S., Faizo, A.A., Hassan, A.M., Alsubhi, T.L., Othman, N.A. and Azhar, E.I. 2020. Development and optimization of in-house ELISA for detection of human IgG antibody to SARS-CoV-2 full-length spike protein. Pathogens 9, 1–11; doi:10.3390/pathogens9100803 Bhatt, S., Gething, P.W., Brady, O.J., Messina, J.P., Farlow, A.W., Moyes, C.L., Drake, J.M., Brownstein, J.S., Hoen, A.G., Sankoh, O., Myers, M.F., George, D.B., Jaenisch, T., Wint, G.R., Simmons, C.P., Scott, T.W., Farrar, J.J. and Hay, S.I. 2013. The global distribution and burden of dengue. Nature 496, 504–507; doi:10.1038/nature12060 Brady, O.J., Gething, P.W., Bhatt, S., Messina, J.P., Brownstein, J.S., Hoen, A.G., Moyes, C.L., Farlow, A.W., Scott, T.W. and Hay, S.I. 2012. Refining the Global Spatial Limits of Dengue Virus Transmission by Evidence-Based Consensus. PLoS Negl. Trop. Dis. 6(8), e1760. doi: 10.1371/journal.pntd.0001760 Chaturvedi, U.C., Shrivastava, R. and Nagar, R. 2005. Dengue vaccines : problems & prospects. Indian J. Med. Res. 121(5), 639–652. Costa, S.M., Yorio, A.P., Gonçalves, A.J.S., Vidale, M.M., Costa, E.C.B., Mohana-Borges, R., Motta, M.A., Freire, M.S. and Alves, A.M.B. 2011. Induction of a protective response in mice by the dengue virus NS3 protein using DNA vaccines. PLos One 6, 18–19; doi:10.1371/journal.pone.0025685 Dighe, S.N., Ekwudu, O., Dua, K., Chellappan, D.K., Katavic, P.L. and Collet, T.A. 2019. Recent update on anti-dengue drug discovery. Eur. J. Med. Chem. 176, 431–455; doi:10.1016/j.ejmech.2019.05.010 Durham, N.D., Agrawal, A., Waltari, E., Croote, D., Zanini, F., Fouch, M., Davidson, E., Smith, O., Carabajal, E., Pak, J.E., Doranz, B.J., Robinson, M., Sanz, A.M., Albornoz, L.L., Rosso, F., Einav, S., Quake, S.R., Mccutcheon, K.M. and Goo, L. 2019. Broadly neutralizing human antibodies against dengue virus identified by single B cell transcriptomics. Elife 8, 1–29; doi:10.7554/eLife.52384 Enriquez, J., Mims, B.M.D., Trasti, S., Furr, K.L. and Grisham, M.B. 2020. Genomic, microbial and environmental standardization in animal experimentation limiting immunological discovery. BMC. Immunol. 21, 1–19; doi:10.1186/s12865-020-00380-x Fajardo-Sánchez, E., Galiano, V. and Villalaín, J. 2017. Spontaneous membrane insertion of a dengue virus NS2A peptide. Arch. Biochem. Biophys. 627, 56–66; doi:10.1016/j.abb.2017.06.016 Feng, K., Zheng, X., Wang, R., Gao, N., Fan, D., Sheng, Z., Zhou, H., Chen, H. and An, J. 2020. Long-Term Protection Elicited by a DNA Vaccine Candidate Expressing the prM-E Antigen of Dengue Virus Serotype 3 in Mice. Front. Cell. Infect. Microbiol. 10, 1–12; doi:10.3389/fcimb.2020.00087 Gil, L., Cobas, K., Lazo, L., Marcos, E., Hernández, L., Suzarte, E., Izquierdo, A., Valdés, I., Blanco, A., Puentes, P., Romero, Y., Pérez, Y., Guzmán, M.G., Guillén, G. and Hermida, L. 2016. A Tetravalent Formulation Based on Recombinant Nucleocapsid-like Particles from Dengue Viruses Induces a Functional Immune Response in Mice and Monkeys. J. Immunol. 197, 3597–3606; doi: 10.4049/jimmunol.1600927 Gono, T., Okazaki, Y., Murakami, A. and Kuwana, M. 2019. Improved quantification of a commercial enzyme-linked immunosorbent assay kit for measuring anti-MDA5 antibody. Mod. Rheumatol. 29, 140–145; doi:10.1080/14397595.2018.1452179 Guy, B., Noriega, F., Ochiai, R.L., L’Azou, M., Delore, V., Skipetrova, A., Verdier, F., Coudeville, L., Savarino, S. and Jackson, N. 2017. A recombinant live attenuated tetravalent vaccine for the prevention of dengue. Expert. Rev. Vaccines. 16, 671–683; doi:10.1080/14760584.2017.1335201 Hakami, A.R., Ball, J.K. and Tarr, A.W. 2015. Non-ionic detergents facilitate non-specific binding of M13 bacteriophage to polystyrene surfaces. J. Virol. Methods 221, M1–M8; doi:10.1016/j.jviromet.2015.04.023 Jiang, L., Wu, X., Wu, Y., Bai, Z., Jing, Q., Luo, L., Dong, Z., Yang, Z., Xu, Y., Cao, Y., Di, B., Wang, Y. and Wang, M. 2013. Molecular epidemiological and virological study of dengue virus infections in Guangzhou, China, during 2001-2010. Virol. J. 10, 1–9. PLoS Negl. Trop. Dis. 6(8), e1760. Kementerian Kesehatan RI (n.d.). Situasi Penyakit Demam Berdarah Di Indonesia 2017. 71–8. Konishi, E., Kosugi, S. and Imoto, J.I. 2006. Dengue tetravalent DNA vaccine inducing neutralizing antibody and anamnestic responses to four serotypes in mice. Vaccine 24, 2200–2207; doi:10.1016/j.vaccine.2005.11.002 Kuhn, R.J., Zhang, W., Rossmann, M.G., Pletnev, S.V., Corver, J., Lenches, E., Jones, C.T., Mukhopadhyay, S., Chipman, P.R., Strauss, E.G., Baker, T.S. and Strauss, J.H. 2002. Structure of Dengue Virus: implications for Flavivirus Organization, Maturation, and Fusion. Cell 108, 717–725; doi:10.1142/9789814513357_0037 Lazo, L., Izquierdo, A., Suzarte, E., Gil, L., Valdés, I., Marcos, E.., Álvarez, M., Romero, Y., Guzmán, M.G., Guillén, G. and Cruz, L.H. 2014. Evaluation in mice of the immunogenicity and protective efficacy of a tetravalent subunit vaccine candidate against dengue virus. Microbiol. Immunol. 58(4), 219–226., doi: 10.1111/1348-0421.12140 Lee, H., Ryu, J.H., Park, H.S., Park, K.H., Bae, H., Yun, S., Choi, A.R., Cho, S.Y., Park, C., Lee, D.G., Lim, J., Lee, J., Lee, S., Shin, S., Park, H. and Oh, E.J. 2019. Comparison of Six Commercial Diagnostic Tests for the Detection of Dengue Virus Non-Structural-1 Antigen and IgM/IgG Antibodies. Ann. Lab. Med. 39(6), 566–571. Lestari, C., Narita, V., Soediro, T. and Sjarurachman, A. 2016. Ekspresi dan Purifikasi Protein Rekombinan Non-Struktural NS1 Virus Dengue Serotipe 1 Strain Indonesia Pada Pichia Pastoris. J. Biotek Medisiana Indones. 5(2), 121–132. Lestari, C.S.W., Yohan, B., Yunita, A., Meutiawati, F., Hayati, R.F., Trimarsanto, H. and Sasmono, R.T. 2017. Phylogenetic and evolutionary analyses of dengue viruses isolated in Jakarta, Indonesia. Virus. Genes. 53, 778–788. Liu, Y., Zhou, J., Yu, Z., Fang, D., Fu, C., Zhu, X., He, Z., Yan, H. and Jiang, L. 2014. Tetravalent recombinant dengue virus-like particles as potential vaccine candidates: immunological properties. BMC Microbiol. 14, 1–13; doi:10.1186/s12866-014-0233-3 Marbawati, D., and Wijayanti, T. 2014. Vaksin Dengue, Tantangan, Perkembangan, dan Strategi. Balaba 10, 39–46. Marfía, J.I., Smith, I., Targovnik, A.M., Di Lello, F.A., Wolman, F.J., Flichman, D.M., Miranda, M.V. and Valdez, S.N. 2025. Development and validation of dengue virus envelope protein domain III IgG antibody enzyme-linked immunosorbent assay. Indian J. Med. Microbiol. 43, 29–36; doi:10.1016/j.ijmmb.2024.09.004 Medina, F.A., Vila, F., Adams, L.E., Cardona, J., Carrion, J., Lamirande, E., Acosta, L.N., De León-rodríguez, C.M., Beltran, M., Grau, D., Rivera-Amill, V., Balmaseda, A., Harris, E., Madewell, Z.J., Waterman, S.H., Paz-Bailey, G., Whitehead, S. and Muñoz-Jordán, J.L. 2024. Comparison of the sensitivity and specificity of commercial anti-dengue virus IgG tests to identify persons eligible for dengue vaccination. J. Clin. Microbiol. 62, 1–16; doi:10.1128/jcm.00593-24 Minic, R., and Zivkovic, I. 2021. Optimization, Validation and Standardization of ELISA. Norovirus. doi: 10.5772/intechopen.94338 Moorthy, M., Chandy, S., Selvaraj, K. and Abraham, A.M. 2009. Evaluation of a rapid immunochromatographic device for the detection of IgM & IgG antibodies to Dengue viruses (DENV) in a tertiary care hospital in South India. Indian. J. Med. Microbiol. 27, 254–256; doi:10.4103/0255-0857.53210 Murugananthan, K., Coonghe, P.A.D., Ketheesan, N. and Noordeen, F. 2018. Comparison of a rapid immuno-chromatography assay with a standard ELISA for the detection of IgM and IgG antibodies against dengue viruses. VirusDisease 29, 199–202; doi:10.1007/s13337-018-0440-x Ogrič, M., Žigon, P., Lakota, K., Praprotnik, S., Drobne, D., Štabuc, B., Sodin-Semrl, S. and Čučnik, S. 2019. Clinically important neutralizing anti-drug antibodies detected with an in-house competitive ELISA. Clin. Rheumatol. 38, 361–370; doi:10.1007/s10067-018-4213-0 Osorio, J.E., Partidos, C.D., Wallace, D. and Stinchcomb, D.T. 2015. Development of a recombinant, chimeric tetravalent dengue vaccine candidate. Vaccine 33, 7112–7120; doi:10.1016/j.vaccine.2015.11.022 Pengsaa, K., Limkittikul, K., Yoksan, S., Wisetsing, P. and Sabchareon, A. 2011. Dengue antibody in Thai children from maternally transferred antibody to acquired infection. Pediatr. Infect. Dis. J. 30, 897–900; doi:10.1097/inf.0b013e31821f07f6 Rothman, A.L. 2004. Dengue: defining protective versus pathologic immunity. J. Clin. Invest. 113, 946–951; doi:10.1172/JCI21512 Saito, J. 2022. Takeda. Available via https://www.takeda.com/newsroom/newsreleases/2022/takedas-qdenga-dengue-tetravalent-vaccine-live-attenuated-approved-in-indonesia-for-use-regardless-of-prior-dengue-exposure/ (Accessed September6, 2022). Sanders, M., McPartlin, D., Moran, K., Guo, Y., Eeckhout, M., O’Kennedy, R.,De Saeger, S. and Maragos, C 2016. Comparison of Enzyme-linked Immunosorbent assay, surface plasmon resonance and biolayer interferometry for screening of deoxynivalenol in Wheat and Wheat Dust. Toxins. (Basel). 8, 1–14; doi:10.3390/toxins8040103 Sarathy, V.V., White, M., Li, L., Kaiser, J.A., Campbell, G.A., Milligan, G.N., Bourne, N. and Barrett, A.D.T. 2018. Characterization of a murine model of non-lethal, symptomatic dengue virus infection. Sci. Rep. 8, 1–16; doi:10.1038/s41598-018-22618-w Shrivastava, N., Shrivastava, A., Ninawe, S.M., Sharma, S., Kumar, J.S., Alam, S.I., 2019. Development of multispecies recombinant nucleoprotein-based indirect ELISA for high-throughput screening of Crimean-Congo hemorrhagic fever virus-specific antibodies. Front. Microbiol. 10, 1–14; doi:10.3389/fmicb.2019.01822 Simmonds, P., Becher, P., Bukh, J., Gould, E.A., Meyers, G., Monath, T., Muerhoff, S., Pletnev, A., Rico-Hesse, R., Smith, D.B. and Stapleton, J.T. 2017. ICTV virus taxonomy profile: flaviviridae. J. Gen. Virol. 98(2–3), 2; doi:10.1099/jgv.0.000672 Tatura, S.N.N., Denis, D., Santoso, M.S., Hayati, R.F., Kepel, B.J., Yohan, B. and Sasmono, R.T. 2021. Outbreak of severe dengue associated with DENV-3 in the city of Manado, North Sulawesi, Indonesia. Int. J. Infect. Dis. 106, 185–196; doi:10.1016/j.ijid.2021.03.065 Teimouri, A., Modarressi, M.H., Shojaee, S., Mohebali, M., Rezaian, M. and Keshavarz, H. 2019. Development, optimization, and validation of an in-house dot-ELISA rapid test based on SAG1 and GRA7 proteins for serological detection of Toxoplasma gondii infections. Infect. Drug Resist. 12, 2657–2669; doi:10.2147/IDR.S219281 Uno, N. and Ross, T.M. 2021. Universal Dengue Vaccine Elicits Neutralizing Antibodies against Strains from All Four Dengue Virus Serotypes. J. Virol. 95, 1–16; doi:10.1128/jvi.00658-20 WHO. 2025. Dengue. Available via https://www.who.int/en/news-room/fact-sheets/detail/dengue-and-severe-dengue?utm_source=chatgpt.com. (Accessed August28, 2025). Yen, L.C., Chen, H.W., Ho, C.L., Lin, C.C., Lin, Y.L., Yang, Q.W., Chiu, K.C., Lien, S.P., Lin, R.J. and Liao, C.L. 2023. Neutralizing antibodies targeting a novel epitope on envelope protein exhibited broad protection against flavivirus without risk of disease enhancement. J. Biomed. Sci. 30, 1–12; doi:10.1186/s12929-023-00938-y Yuen, Y.J., Sabitha, T., Li, L.J., Walvekar, V.A., Ramesh, K., Kini, R.M., Sivaraman, J. and Mok, Y.K. 2025. Hijacking of plasminogen by dengue virus: the kringle-4 and -5 domains of plasminogen binds synergistically to the domain I of envelope protein. Protein Sci. A Publ. Protein Soc. 34, 1–15; doi:10.1002/pro.70035 Zheng, Q., Fan, D., Gao, N., Chen, H., Wang, J., Ming, Y., Li, J. and An, J. 2011. Evaluation of a DNA vaccine candidate expressing prM-E-NS1 antigens of dengue virus serotype 1 with or without granulocyte-macrophage colony-stimulating factor (GM-CSF) in immunogenicity and protection. Vaccine 29, 763–771; doi:10.1016/j.vaccine.2010.11.014 Zonetti, L.F.C., Coutinho, M.C. and De Araujo, A.S. 2018. Molecular Aspects of the Dengue Virus Infection Process: a Review. Protein. Pept. Lett. 25, 712–719; doi:10.2174/0929866525666180709115506 | ||
| How to Cite this Article |
| Pubmed Style Lestari CSW, Pambudi S, Novientri G, Fithriani F, Panjaitan NSD, Sunarno S, Hartanti MD, Suprihatin S. Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 Web Style Lestari CSW, Pambudi S, Novientri G, Fithriani F, Panjaitan NSD, Sunarno S, Hartanti MD, Suprihatin S. Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. https://www.openveterinaryjournal.com/?mno=267210 [Access: February 04, 2026]. doi:10.5455/OVJ.2026.v16.i1.65 AMA (American Medical Association) Style Lestari CSW, Pambudi S, Novientri G, Fithriani F, Panjaitan NSD, Sunarno S, Hartanti MD, Suprihatin S. Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 Vancouver/ICMJE Style Lestari CSW, Pambudi S, Novientri G, Fithriani F, Panjaitan NSD, Sunarno S, Hartanti MD, Suprihatin S. Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 Harvard Style Lestari, C. S. W., Pambudi, . S., Novientri, . G., Fithriani, . F., Panjaitan, . N. S. D., Sunarno, . S., Hartanti, . M. D. & Suprihatin, . S. (2026) Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 Turabian Style Lestari, Christina Safira Whinie, Sabar Pambudi, Gissi Novientri, Fithriani Fithriani, Novaria Sari Dewi Panjaitan, Sunarno Sunarno, Monica Dwi Hartanti, and Suprihatin Suprihatin. 2026. Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 Chicago Style Lestari, Christina Safira Whinie, Sabar Pambudi, Gissi Novientri, Fithriani Fithriani, Novaria Sari Dewi Panjaitan, Sunarno Sunarno, Monica Dwi Hartanti, and Suprihatin Suprihatin. "Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate." doi:10.5455/OVJ.2026.v16.i1.65 MLA (The Modern Language Association) Style Lestari, Christina Safira Whinie, Sabar Pambudi, Gissi Novientri, Fithriani Fithriani, Novaria Sari Dewi Panjaitan, Sunarno Sunarno, Monica Dwi Hartanti, and Suprihatin Suprihatin. "Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate." doi:10.5455/OVJ.2026.v16.i1.65 APA (American Psychological Association) Style Lestari, C. S. W., Pambudi, . S., Novientri, . G., Fithriani, . F., Panjaitan, . N. S. D., Sunarno, . S., Hartanti, . M. D. & Suprihatin, . S. (2026) Optimization of in-house indirect ELISA for the detection of mice IgG anti-dengue prM/E DENV-3 as a vaccine candidate. doi:10.5455/OVJ.2026.v16.i1.65 |