| Research Article | ||
Open Vet. J.. 2026; 16(1): 679-685 Open Veterinary Journal, (2026), Vol. 16(1): 679-685 Research Article Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA geneKarar Ali Abdulkhuder1*, Mahmood Nabeel Awad2 and Khetam Qaid Mayea31Department of Medical Laboratory Technology, The Islamic University, Diwaniyah, Iraq 2College of Nursing, University of Altoosi, Al-Najaf, Iraq 3College of Veterinary Medicine, University of Al-Qadisiyah, Diwaniyah, Iraq *Corresponding Author: Karar Ali Abdulkhuder. Department of Medical Laboratory Technology, Submitted: 24/06/2025 Revised: 30/11/2025 Accepted: 21/12/2025 Published: 31/01/2026 © 2025 Open Veterinary Journal
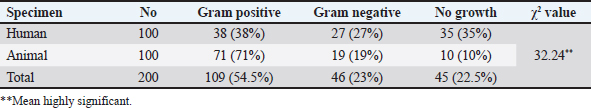
AbstractBackground: Staphylococcus aureus is an opportunistic pathogen responsible for a wide range of infections in humans and animals, including ocular infections in companion animals such as cats and dogs. Molecular identification based on the 16S rRNA gene provides a reliable approach for accurate bacterial detection and phylogenetic analysis. Aim: This study aimed to molecularly characterize S. aureus isolates obtained from humans, cats, and dogs using 16S rRNA gene sequencing in order to assess genetic diversity and evaluate the potential for zoonotic transmission. Methods: Ocular swab samples were collected from humans (n=100), cats (n=50), and dogs (n=50) between July 2023 and January 2024. Isolates were identified using conventional culture and biochemical methods, followed by polymerase chain reaction amplification of the 16S rRNA gene. Based on antibiotic resistance profiles, 25 representative isolates (10 from cats, 10 from dogs, and 5 from humans) were selected for sequencing and phylogenetic analysis. Results: Sequencing analysis confirmed the identity of all selected isolates as S. aureus. Genetic analysis demonstrated both similarities and variations among isolates derived from different hosts, indicating genetic diversity and suggesting possible cross-species transmission between humans and companion animals. Conclusion: The findings highlight the genetic diversity of S. aureus isolates from humans, cats, and dogs and support the potential role of companion animals in the zoonotic transmission of this pathogen. Molecular characterization using the 16S rRNA gene is a valuable tool for tracking bacterial evolution and understanding public health risks associated with pathogens shared between humans and animals. Continuous surveillance of antimicrobial resistance and molecular epidemiology is essential, particularly in environments with close human–animal contact. Keywords: Staphylococcus aureus, 16S rRNA, Zoonotic transmission, Genetic diversity, Molecular characterization. IntroductionStaphylococcus aureus is a versatile and opportunistic pathogen that colonizes the skin, mucosal surfaces, and nasal cavities of both humans and animals. Although it can exist as a commensal organism, it is also responsible for a wide range of infections, including skin and soft tissue infections, pneumonia, bacteremia, and endocarditis (Tong et al., 2015). The ability of S. aureus to colonize multiple host species raises concerns about its potential for cross-species transmission, particularly between humans and companion animals such as cats and dogs (Weese, 2010). Due to their close contact with humans, companion animals may serve as reservoirs for S. aureus, including antibiotic-resistant strains such as methicillin-resistant S. aureus (Boost et al., 2013). Several studies have suggested that pets can acquire S. aureus from their owners and vice versa. For example, a study found that S. aureus colonization was more prevalent in domestic (19.17%) than feral cats (8.3%), suggesting that close human contact may contribute to bacterial transmission (Kadlec et al., 2019). To better understand the genetic relationships and potential zoonotic transmission of S. aureus, molecular characterization techniques, such as 16S rRNA gene sequencing, are widely used. The 16S rRNA gene is highly conserved among bacterial species, making it a valuable tool for species identification and phylogenetic analysis (Janda and Abbott, 2007). By sequencing this gene, researchers can track bacterial evolution and assess interspecies transmission patterns (Becker et al., 2020). Recent genomic studies have demonstrated that S. aureus isolates from humans and animals share genetic similarities, supporting the idea of cross-species transmission (Harrison et al., 2020). This study aims to characterize S. aureus isolates obtained from humans, cats, and dogs using 16S rRNA gene sequencing to explore genetic diversity and interspecies transmission. Understanding these genetic relationships will provide insights into the epidemiology of S. aureus and its impact on public health. Materials and MethodsSample collection and isolation by using culture and biochemical testsIn the present study, (100%) ocular swab samples were collected from humans 63 (63%) male and 37 (37%) female, and (100%) from pet animals dog 50 (50%) dog and 50 (50%) cat pets between July 2023 and January 2024. Comparison of bacterial isolates from in humans and animals eye swabs. In the “human” group, gram-positive bacteria were found in 38 (38%) samples, gram-negative bacteria in 27 (27%), and no growth occurred in 35 (35%). In the “animal” group, gram-positive bacteria were found in 71 (71%) of samples, Gram-negative bacteria in 19 (19%), and no growth occurred in 10 (10%). The overall distribution across both groups shows the presence of gram-positive bacteria in 109 (54.5%) of samples (Table 1 and Fig. 1), gram-negative bacteria in 46 (23%), and no growth in 45 (22.5%).
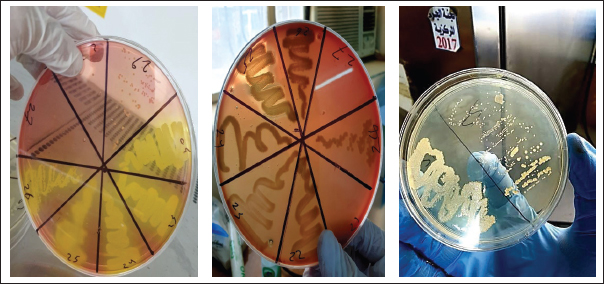
Fig. 1. S. aureus growth on mannitol salt agar, blood agar, and nutrient agar. Table 1. Percentage of bacteria isolated from eye swabs in humans and animals.
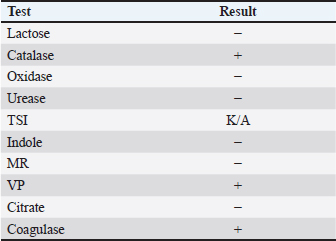
The present study involved collecting 200 ocular swab samples from humans and animals. When comparing these results with those of previous studies, the current study findings are consistent with the general trend observed in ocular infections, where Gram-positive bacteria are often predominant in animals. Biochemical identification of S. aureusBacterial samples that showed bacterial growth were initially diagnosed through biochemical tests. The isolate of S. aureus shown in Figure 1 was identified. Figure 1A shows S. aureus on mannitol salt agar medium, and Figure 1B shows the same bacteria on blood agar medium, where the type of hemolysis was clearly of the beta hemolytic type. Figure 1C shows S. aureus grown on nutrient agar showed a whitish yellow color (Table 2). Table 2. Identifaction of S.aureas by biochemical testing.
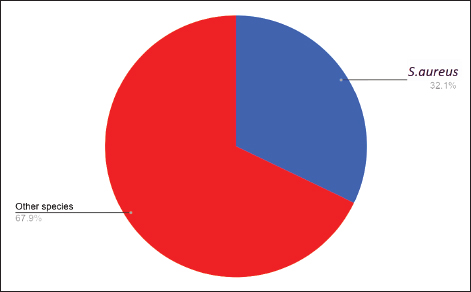
Staphylococcus aureus diagnosis via the VITEK 2 systemThe S. aureus was detected in 35 (32.1%) samples, while the remaining 74 (67.9%) samples contained other bacterial species. Gram-positive 109 (100%) samples were also detected. As shown in Figure 2. In the animal group, Gram-positive bacteria were found in 71 (71%) of samples, gram-negative bacteria in 19 (19%), and no growth occurred in 10 (10%).
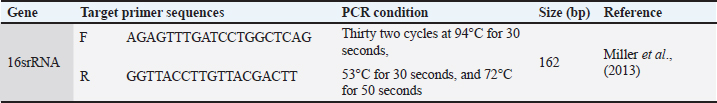
Fig. 2. Staphylococcus species isolated from ocular swabs from humans and small animals. In the human group, gram-positive bacteria were found in 38 (38%) of the samples, gram-negative bacteria in 27 (27%), and no growth occurred in 35 (35%). Detection antibiotic-resistant genesDetection of 16S rRNA was performed using polymerase chain reaction (PCR) through the following steps. DNA extractionThe first step was to extract DNA from the bacterial sample using boiling methods as previously reported (Mahuku, 2004). Primer designSpecific primers targeting the three genes were identified and validated using Primer 3 software (Table 3). These primers bind to the target DNA. Table 3. Primers used in the current study.
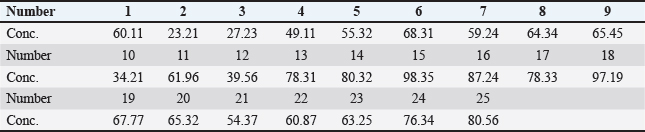
Ethical approvalThis study was conducted in accordance with the ethical standards of the responsible institutional and national research committees. Ethical approval for the collection of human and animal ocular samples was obtained from the Scientific and Ethical Committee of the College of Veterinary Medicine, University of Al-Qadisiyah, Diwaniyah, Iraq. Written informed consent was obtained from all human participants, and sample collection from animals was performed in compliance with animal welfare guidelines, with the permission of the owners and without causing unnecessary harm or distress. ResultsMolecular studyIn the molecular study, 25 (25%) isolates of S. aureus (10 from cats, 10 from dogs, and 5 from humans) were selected out of a total of 35 (35%) isolates. The most antibiotic-resistant isolates were selected for molecular analysis. DNA extractionThe DNA of all isolates was extracted and purified using a genomic DNA purification kit (Table 4). The results were detected by electrophoresis on a 1.5% agarose gel and exposed to UV light, where the DNA appeared as compact bands. Table 4. Genomic DNA concentrations of ng/ml.
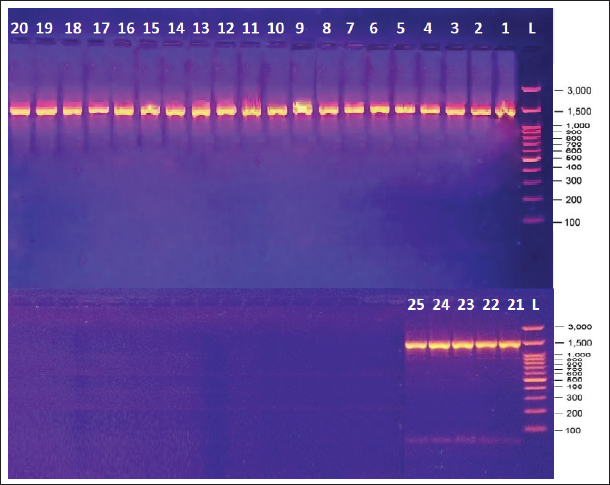
16s RNA gene detectionThe identification of the 16S rRNA gene in 25 (25%) isolates (10 cats, 10 dogs, and 5 humans) of S. aureus demonstrates the efficacy of molecular techniques such as PCR in bacterial detection and characterization. The amplification results, confirmed by electrophoresis, showed clear DNA bands corresponding to the expected size of 1,500 base pairs (Fig. 3). This size is consistent with the conserved nature of the 16S rRNA gene, which is commonly used as a molecular marker for bacterial identification (Janda and Abbott, 2007). The successful amplification of all isolates indicates the high specificity of the primers used. This binding specificity ensures accurate detection of the target gene, minimizing the risk of false positives or nonspecific amplifications. Visualization of single DNA bands under UV light, stained with ethidium bromide, further validates the purity and integrity of the amplified product. Such results are critical for downstream applications such as sequencing and phylogenetic analysis (Clarridge, 2004).
Fig. 3. DNA amplification of a partial sequence in local Staphylococcus sp. Detecting the 16srRNA gene using PCR. Lane 1: 1 adder, lane 1-25, positive results. lane 100: 3,000 bp marker (Ladder). Sequence of the 16s rRNAThe 16S rRNA gene is a well-established molecular marker because of its highly conserved nature across bacterial species, making it an indispensable tool for bacterial identification and classification (Janda and Abbott, 2007). The results of multiple sequence alignment (MSA) for the 25 Staphylococcus spp. strains, (Fig. 4) highlight the genetic similarities within the species, reflecting their close evolutionary relationships.
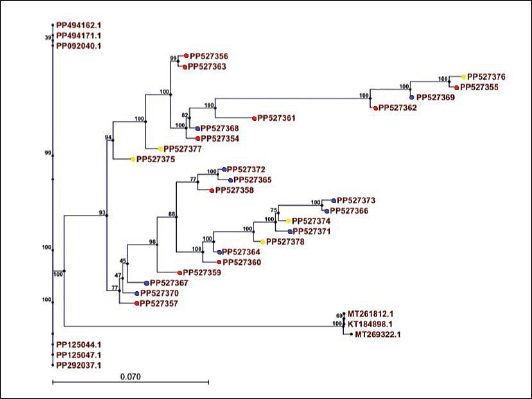
Fig. 4. Multiple sequence alignment analysis of the partial 16S ribosomal RNA gene in local S. auerus. This high degree of conservation is consistent with the 16S rRNA gene’s universal role in bacterial ribosomal function. Despite the overall similarity, the observed sequence variations, such as the single-nucleotide polymorphism (SNP) at position 54 between strains “PP4941711” and “PP494162158AAGC,” provide critical information for distinguishing individual strains or subspecies. Similarly, differences in nucleotide sequences toward the end of the gene between strains “PP292037” and “PP125047.1” indicate potential genetic diversity within the group. Such sequence-level variations are essential for fine-tuned taxonomic assignments and can aid in studying bacterial populations’ evolutionary and functional dynamics (Clarridge, 2004). These findings emphasize the dual utility of the 16S rRNA gene: its conserved regions enable broad bacterial identification, while its variable regions allow for strain-level differentiation. This capability is crucial for understanding microbial ecology, pathogenicity, and resistance mechanisms. Phylogenetic treeThe phylogenetic tree analysis based on the partial 16S rRNA gene sequences provides an in-depth look at the evolutionary relationships among different S. aureus strains from the current study and other geographic locations. The 16S rRNA gene is a widely recognized genetic marker for bacterial taxonomy because of its high conservation across bacterial species, yet it contains enough variation to distinguish between strains and species (Clarridge, 2004; Janda and Abbott, 2007). The phylogenetic tree illustrates that the S. aureus strains analyzed in this study are derived from various geographic regions, such as Russia, Iraq, Egypt, India, France, the USA, and Korea. This highlights the global distribution of S. aureus and reflects its adaptability to different environmental niches. The significant diversity among strains, as seen in the branching patterns of the phylogenetic tree, underscores the possibility of genetic variations arising from local environmental factors, host interactions, or genetic drift. The presence of strains with specific host origins (e.g., cats, dogs, and humans) further emphasizes the zoonotic potential and cross-species transmission capability S. aureus. Host species are represented with distinct color-coded nodes on the phylogenetic tree, such as red, blue, and yellow nodes for cats, dogs, and humans, respectively. This zoonotic characteristic has been widely documented in S. aureus studies and reflects its ability to jump across species boundaries (O’Neill et al., 2021; Watson et al., 2022). These observations align with recent insights into the evolutionary dynamics of S. aureus. The genetic diversity and phylogenetic clustering patterns in this study can inform public health strategies and contribute to the monitoring of emerging strains with zoonotic potential (Fig. 5) (Guo et al., 2023).
Fig. 5. Phylogenetic tree analysis based on the 16S ribosomal RNA gene partial sequence used for staphylococcus species typing detection of local S. aureus isolates (red, cats; blue, dogs; yellow, human). Key findings from phylogenetic tree analysis multiple of S. aureus strains share a common ancestor, reflecting evolutionary relatedness. Strain diversity correlates with geographical origins and host interactions. The zoonotic potential of S. aureus is supported by its ability to infect multiple species (e.g., cats, dogs, and humans). DiscussionThis molecular study demonstrates the effectiveness of PCR techniques in the detection and classification of S. aureus by targeting the 16S rRNA gene, which serves as a highly conserved molecular marker among bacterial species. The results indicate that the primers used have high specificity, ensuring accurate amplification of the target gene and minimizing false positives or nonspecific amplification. This specificity is critical for molecular diagnostic methods’ reliability (Clarridge, 2004; Janda and Abbott, 2007). Sequence analysis and alignment revealed both high conservation and subtle variations within the 16S rRNA gene among the studied S. aureus strains. These nucleotide variations, including SNPs, provide essential markers for differentiating strains at a finer taxonomic level, facilitating epidemiological tracking and understanding bacterial evolution (Clarridge, 2004). The phylogenetic analysis highlights the global distribution and genetic diversity of S. aureus, confirming its adaptability to various hosts and environments. The clustering patterns indicate that genetic variation correlates with geographic origin and host species, supporting the zoonotic potential bacterium. The ability of S. aureus to infect multiple species, including cats, dogs, and humans, underscores its significance in both veterinary and human medicine, especially within the One Health framework (O’Neill et al., 2021; Watson et al., 2022) the genetic diversity observed among local isolates may be influenced by environmental pressures, host-pathogen interactions, and genetic drift, which could impact antimicrobial resistance and virulence characteristics. This underlines the need for continuous surveillance and molecular characterization to inform effective control measures and public health interventions (Guo et al., 2023). ConclusionThis study successfully demonstrated the utility of molecular techniques, particularly PCR amplification of the 16S rRNA gene, for the accurate identification and characterization of S. aureus isolates from different hosts, including cats, dogs, and humans. The selection of antibiotic-resistant isolates for molecular analysis allowed the focused investigation of strains with potential clinical significance. The high quality and purity of the extracted genomic DNA, confirmed by agarose gel electrophoresis, facilitated reliable amplification of the 16S rRNA gene. Sequence analysis and MSA revealed a high degree of conservation among the isolates, with strain-level differentiation enabled by specific nucleotide variations. These findings underscore the dual role of the 16S rRNA gene in broad bacterial identification and detailed phylogenetic studies. Phylogenetic tree analysis illustrated the genetic diversity and evolutionary relationships among S. aureus strains from local and global origins, highlighting the zoonotic potential of S. aureus in multiple host species. The observed clustering based on geographic location and host species reinforces the need for continuous molecular surveillance to monitor emerging strains and track transmission pathways. Overall, this study contributes valuable molecular data to the understanding of S. aureus epidemiology and highlights the importance of integrating molecular tools to support effective diagnosis, treatment, and control strategies in veterinary and human infectious disease research. AcknowledgmentsThe authors would like to thank the staff of the Department of Medical Laboratory Technology, The Islamic University, Diwaniyah, Iraq, and the College of Veterinary Medicine, University of Al-Qadisiyah, for their technical assistance and support during sample collection and laboratory work. Conflict of interestThe authors declare that there is no conflict of interest regarding the publication of this article. FundingThis research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors. Authors’ contributionsKarar Ali Abdulkhuder: conceived and designed the study, performed the laboratory experiments, molecular analysis, data interpretation, and drafted the manuscript. Mahmood Nabeel Awad: contributed to sample collection, bacteriological identification, and critically revised the manuscript. Khetam Qaid Mayea: participated in data analysis, interpretation of results, and manuscript revision. All authors read and approved the final version of the manuscript. Data availabilityThe datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request. ReferencesBecker, K., Heilmann, C. and Peters, G. 2020. Coagulase-negative Staphylococci. Clin. Microbiol. Rev. 27(4), 870–926. Boost, M.V., O’Donoghue, M.M. and Siu, K.H. 2013. Characterization of methicillin-resistant Staphylococcus aureus (MRSA) isolates from dogs and their owners. EpidemiolInfection 141(10), 2186–2191. Clarridge, J.E. 2004. Impact of 16S rRNA gene sequence analysis for identifying of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev. 17(4), 840–862. Guo, F., Li, Y., Wang, Y., Zhang, X., Chen, S. and Liu, H. 2023. Evolutionary dynamics of zoonotic S. aureus strains in host and environment. Gut. Pathogens 15, 32. Harrison, E.M., Weinert, L.A., Holden, M.T.G., Welch, J.J., Wilson, K., Morgan, F.J.E., Harris, S.R., Loeffler, A., Boag, A.K., Peacock, S.J., Paterson, G.K., Waller, A.S., Parkhill, J. and Holmes, M.A. 2020. A shared population of epidemic methicillin-resistant Staphylococcus aureus 15 circulates in humans and companion animals. mBio 5(2), 98. Janda, J.M. and Abbott, S.L. 2007. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J. Clin. Microbiol. 45(9), 2761–2764. Kadlec, K., Feßler, A.T., Hauschild, T. and Schwarz, S. 2019. Novel insights into the epidemiology of Staphylococcus aureus colonization in domestic cats: higher prevalence in companion than in feral cats. Vet. Microbiol. 229, 89–94. Miller, C.S., Handley, K.M., Wrighton, K.C., Frischkorn, K.R., Thomas, B.C. and Banfield, J.F. 2013. Short-read assembly of full-length 16S amplicons reveals bacterial diversity in subsurface sediments. PLos One. 8(2), e56018. Mahuku, G.S. 2004. A simple extraction method suitable for PCR-based analysis of plant, fungal, and bacterial DNA. Plant Mol. Biol. Rep. 22(1), 71–81. O’Neill, J., Klemm, E.J., Shaw, R. and Piddock, L.J.V. 2021. Geographic distribution and diversity of Staphylococcus aureus strains. Front. Microbiol. 7(5), 000561. Tong, S.Y.C., Davis, J.S., Eichenberger, E., Holland, T.L. and Fowler, V.G. 2015. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 28(3), 603–661. Watson, A., Wright, J., Taylor, L.H. and Weese, J.S. 2022. Host specificity and cross-species transmission in S. aureus. BMC Microbiology Weese, J.S. 2010. Methicillin-resistant Staphylococcus aureus in animals. ILAR. J. 51(3), 233–244. | ||
| How to Cite this Article |
| Pubmed Style Abdulkhuder KA, Awad MN, Mayea KQ. Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Vet. J.. 2026; 16(1): 679-685. doi:10.5455/OVJ.2026.v16.i1.63 Web Style Abdulkhuder KA, Awad MN, Mayea KQ. Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. https://www.openveterinaryjournal.com/?mno=266451 [Access: February 04, 2026]. doi:10.5455/OVJ.2026.v16.i1.63 AMA (American Medical Association) Style Abdulkhuder KA, Awad MN, Mayea KQ. Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Vet. J.. 2026; 16(1): 679-685. doi:10.5455/OVJ.2026.v16.i1.63 Vancouver/ICMJE Style Abdulkhuder KA, Awad MN, Mayea KQ. Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Vet. J.. (2026), [cited February 04, 2026]; 16(1): 679-685. doi:10.5455/OVJ.2026.v16.i1.63 Harvard Style Abdulkhuder, K. A., Awad, . M. N. & Mayea, . K. Q. (2026) Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Vet. J., 16 (1), 679-685. doi:10.5455/OVJ.2026.v16.i1.63 Turabian Style Abdulkhuder, Karar Ali, Mahmood Nabeel Awad, and Khetam Qaid Mayea. 2026. Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Veterinary Journal, 16 (1), 679-685. doi:10.5455/OVJ.2026.v16.i1.63 Chicago Style Abdulkhuder, Karar Ali, Mahmood Nabeel Awad, and Khetam Qaid Mayea. "Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene." Open Veterinary Journal 16 (2026), 679-685. doi:10.5455/OVJ.2026.v16.i1.63 MLA (The Modern Language Association) Style Abdulkhuder, Karar Ali, Mahmood Nabeel Awad, and Khetam Qaid Mayea. "Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene." Open Veterinary Journal 16.1 (2026), 679-685. Print. doi:10.5455/OVJ.2026.v16.i1.63 APA (American Psychological Association) Style Abdulkhuder, K. A., Awad, . M. N. & Mayea, . K. Q. (2026) Molecular characterization of Staphylococcus aureus isolates from humans, cats, and dogs using the 16S rRNA gene. Open Veterinary Journal, 16 (1), 679-685. doi:10.5455/OVJ.2026.v16.i1.63 |