| Research Article | ||
Open Vet. J.. 2026; 16(2): 1289-1296 Open Veterinary Journal, (2026), Vol. 16(2): 1289-1296 Research Article Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, IraqNuha Jabbar Alrikaby*, Hasan Jasim Hami, and Amal F. AL-Gorani*Biology Department, College of Science, University of Thi Qar, Nasiriyah, Iraq *Corresponding Author: Nuha Jabbar Alrikaby. Biology Department, College of Science, University of Thi Qar, Nasiriyah, Iraq. Email: Nuha_bio [at] sci.utq.edu.iq Submitted: 16/05/2025 Revised: 10/10/2025 Accepted: 20/10/2025 Published: 28/02/2026 © 2026 Open Veterinary Journal
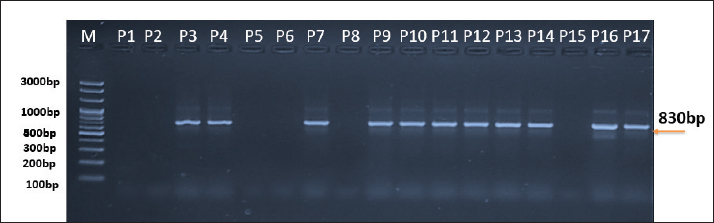
AbstractBackground: Cryptosporidium is a parasite that affects birds and humans, it play role in potential zoonotic interspecies transmission. adequate diagnosis is important to study the epidemiological diversity of this genus. Aim: This study aimed to diagnose the species of Cryptosporidium in domestic pigeons (Columba livia domestica) in Nasiriyah city in Iraq, by using nested polymerase chain reaction (Nested PCR). Methods: Seventeen fecal samples from domestic pigeons were collected. Nested polymerase chain reaction targeting the 18S rRNA gene was done to detect and molecularly characterize the parasite, and we study the genetic similarities with known Cryptosporidium species in humans and birds in gene bank. Results: The rate of infection Pigeons was high 64.7% , the Cryptosporidium was detected in 11 out of 17 samples. Three species were detected : C. meleagridis, C. baileyi, and C. avium. The present study confirms for the first time the presence of Cryptosporidium avium in domestic pigeons in Iraq using molecular analysis. The samples showed a high genetic similarity of 98.50%–99.0% with known human species, indicating a close genetic relationship. This is the first molecular identification of C. avium parasites in pigeons in Nasiriyah, with one sample showing a 99.45% match with the species isolated from birds. Minor genetic mutations were also observed in the 18S rRNA gene, indicating slight genetic diversity that may be related to the adaptation of the parasite. Conclusion: This study highlights the presence and diversity of Cryptosporidium parasites in domestic pigeons in Iraq, emphasizing the potential for parasite transmission between birds and humans. These findings provide a foundation for future research into transmission pathways and infection. Keywords: 18SrRNA, Bird-to-human transmission, Columba livia domestica, C. avium, C. baileyi, C. meleagridis, Nested PCR. IntroductionCryptosporidium is an intestinal protozoan parasite belonging to the Eimeriidae family. It is recognized as a major causative agent of acute watery diarrhea in both humans and animals, posing serious health risks, particularly to immunocompromised individuals (Meinhardt et al., 1996). This parasite is transmitted through contaminated water and food as, well as by contact with infected animals. The potential reservoir for infection with this parasite in humans and other animals is birds, including infected domestic pigeons (Wang et al., 2021). The detecting of Cryptosporidium in avian hosts is very importance, as pigeon , especially in urban area , can act as reservoirs for the parasite and contribute to its transmission. Cryptosporidium is still a non – studied pathogen in the population of birds in Iraq, especially in the Nasiriyah city. To date, no comprehensive studies about its prevalence have performed in this region. So, this study is the first investigation to detect Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah Nested PCR was utilized for the molecular detection of Cryptosporidium in stool samples collected from domestic pigeons. Nested PCR it is most suitable for sensitive and specific molecular detection, and this technique has been confirmed to detect a submicroscopic stool parasite with low density in stool. This method includes two amplification rounds: the first round uses general primers to detect the Cryptosporidium Deoxyribonucleic Acid (DNA) , while the second round uses specific primers to identify particular species with high accuracy (McConkey, 2024). Some studies have confirmed the prevalence of Cryptosporidium in Iraqi pigeons For example, a recent study conducted in 2024 in Baghdad showed a high overall infection rate of 70.83% using conventional PCR. The study also indicated seasonal variation in infection rates, with the highest rate being in February (95%) and the lowest in March (85%), with no statistically significant differences between males and females (Hashim and Al Zubaidi, 2024). New study disciplined the transmission methods of these parasites, its diagnosis , and its infection strategies, confirmed its global scope, especially in developing countries, to reduce the rates of infection through improving the general health and controlling vectors. The intestinal parasites still form a huge challenge in many world regions. (Al-Ataby and Radhi, 2025). According to Liao et al. (2021) Cryptosporidium was detected in birds from pet markets in Wuhan, China, That Shows the ability of its transmission to humans. Intestinal parasites in domestic pigeons may contribute to an imbalance in the trace elements inside the infected host (Maktoof et al., 2023). This study aimed to detect Cryptosporidium parasite in domestic pigeons in Nasiriyah city and to determine the species of this parasite by using nested polymerase chain reaction (PCR). This work contributes to the understanding the pigeon’s role in the prevalence of the parasite in urban environments and paves the way for more research about the common parasites between humans and animals in Iraq. Materials and MethodsSample collection and initial microscopicSeventeen fecal samples were collected from domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. These samples were collected from bird cages or their barn and put in sterile containers, and transferred to laboratory in proper refrigeration circumstances . Then treated immediately to keep its quality. The number of samples was limited relatively due to field conditions and the lack of periodic sample collection . which also requires molecular analysis, substances, and sophisticated laboratory techniques which limits the collection of a larger number of samples. Nevertheless , samples were carefully selected to ensure adequate representation of the available situations , that enhancing the sustainability, the samples collected between February and March 2025 to ensure coverage of the bird season and diversity as well. All samples were submitted for preliminary microscopic examination by using the formalin-ether fixation technique, followed by light microscopy, and modified Ziehl-Neelsen was also used for staining and detecting Cryptosporidium oocysts (Buján et al., 2024), some samples showed distinguish structures that were considered positive. The microscopic results were used to identify samples analyzes by molecular study using polymerase chain reaction ( Nested PCR). DNA extractionGenomic DNA was extracted from the fecal samples of 11 pigeons. Samples were stored at 4°C for 24 hours before molecular extraction. The Presto™ Stool DNA Extraction Kit (Geneaid, Taiwan) was used following the manufacturer’s instructions. The extracted DNA was quantified using a NanoDrop spectrophotometer (260/280 nm) to assess purity and concentration. Primary PCR amplificationPrimary PCR was performed using GoTaq® Green Master Mix (Promega, USA) with a pair of primers targeting the 18S rRNA gene of Cryptosporidium species. The expected product size resulting from this reaction was approximately 435 base pairs. (Rafiei et al., 2014): Primary PCR primers: Forward: 5'-TTCTAGAGCTAATACATGCG-3' Reverse: 5'-CCCATTTCCTTCGAAACAGGA-3' The 25 μl reaction volume contained the following: • 12.5-μl GoTaq® Green Master Mix (2X) • 1 μl of the primary PCR forward primer (10 μM) • 1 μl of the primary PCR reverse primer (10 μM) • 2.5 μl of the extracted DNA • 8 μl of nuclease-free water The thermal cycling conditions for the primary PCR were as follows: Initial denaturation at 95°C for 5 minutes, followed by 35 cycles of the following: • Denaturation at 95°C for 30 seconds • Annealing at 55°C for 30 seconds • Extension at 72°C for 1 minute • Final extension at 72°C for 7 minutes. Nested PCR amplificationNested PCR was performed using the GoTaq® Green Master Mix (Promega, USA) and the following primer pair targeting a region within the primary PCR product. The expected size of the product resulting from this reaction was approximately 305 base pairs. (Rafiei et al., 2014): Nested PCR primers: Forward:5'-GGAAGGGTTGTATTTATTAGATAAAG-3' Reverse: 5'-AAGGAGTAAGGAACAACCTCCA-3' The 25 μl reaction volume contained the following: • 12.5-μl GoTaq® Green Master Mix (2X) • 1 μl of nested PCR forward primer (10 μM) • 1 μl of nested PCR reverse primer (10 μM) • 2 μl of the primary PCR product • 8.5 μl of nuclease-free water. The thermal cycling conditions for the nested PCR were the same as those for the primary PCR, except for the annealing temperature, which was set at 58°C. DNA sequencing and identificationGel extraction method was used for accurate separation of PCR products of positive samples 11 and removal of impurities. The nested PCR primers were sequenced by Macrogen (Korea) in both directions. The obtained DNA sequences were compared with reference sequences in the National Center for Biotechnology Information database using the BLAST tool. Clustal W algorithm for the purpose of multiple sequence, and phylogenetic analysis was performed using neighbor joining method in MEGA software ( version 6.0). Ethical approvalPigeon fecal samples were collected from different local environment were collected from different local environment without harm to birds and no living studies were performed and birds were not captured hence, the ethical approval was not needed in accordance with the Research Ethics Committee of Thi Qar. ResultsSeventeen fecal samples collected from Columba livia domestica pigeons in Nasiriyah city , Southern Iraq to detect the Cryptosporidium using Nested PCR. This study detected the results of molecular diagnostic 11(64.7%) indicating the active infection. The reasons of high percent of infection may because of many factors such as closed rearing conditions, poor hygiene, food and water resources. This results highlighted the pigeon' s role as reservoir for the parasite. This make it a source of transmission of infection to another birds or to human. These species of Cryptosporidium were determined in the infected samples as C. baileyi, C. meleagridis and C. avium. The results of this study revealed high genetic similarity between the Cryptosporidium isolates in this study and human infected species, it varied between 98.75% , this demonstrated to a strong genetic link between the species recorded in this study and the known human infected species. It should be noted that Cryptosporidium avium was diagnostic for the first time in Iraq. This study showed 99.45% genetic similarity with reference isolates. The findings suggested concerns about the potential for zoonotic transmission especially to immunocompromised people. Besides that, slight genetic mutations were identified in the regions of the 18S rRNA gene of the local parasite compared with GenBank reference sequences, this mutations effect on the parasite's adaptation to new hosts and on its transmission routes Figure 2. The present investigation contribute to clearer insight of parasite’s genetic diversity in Iraq and prepare the ground for future researches about its transmission methods between birds and human. This helps refinement of prevention strategies against infection. This investigation provides significant step in the direction of identifying the Cryptosporidium species fauna in Iraq. Draw attention to their relatedness to human parasites. This reinforces the need for further studies to explore methods for preventing infection. Figure 1 shows an image of an agarose gel electrophoresis showing the results of nested PCR products targeting a conserved region of the 18S rRNA gene of the parasite Cryptosporidium extracted from domestic pigeon fecal samples. M (McClean): contains standard molecular weight markers ranging from 3000 to 100 bp for size determination. Samples (P1 - P17): The results for the samples are shown in lanes 1–17. Some samples were positive for Cryptosporidium at 830 bp, indicating the parasite in these samples. The use of nested PCR increased the sensitivity, which enabled the detection of the parasite in samples with low DNA concentrations (Fig. 1). DNA sequenceThe DNA sequencing method was performed for genetic species typing analysis in 18S ribosomal RNA gene in local Cryptosporidium species from domestic pigeon isolates that compared with NCBI-Genbank related Cryptosporidium species isolates. Local Cryptosporidium sp. IQN-No.1, No.4, No.8, and No.10 isolates were closely related to NCBI-BLAST Cryptosporidium meleagridis (KP730308.1), local Cryptosporidium sp. IQN-No.2, No.3, No.4, No.5, No.6, No.7, and No.11 isolates were closely related to NCBI-BLAST Cryptosporidium baileyi (KY448456.1), and the local Cryptosporidium sp. IQN-No.9 isolate was closely related to NCBI-BLAST Cryptosporidium avium (MW783461.1) at total genetic changes (0.01%), as shown in Figure 2. The homology sequence identity between local Cryptosporidium sp. domestic pigeon isolates was close to that of NCBI -BLAST Cryptosporidium species. The genetic homology sequence identity ranged from 98.50 to 99.45% (Table 1). The phylogenetic tree was built using the neighbor-joining method in MEGA X software with 1,000 bootstrap replicates to assess the branch reliability. Figure 2 shows the bootstrap values at the nodes. That positive and negative controls were not included in this phylogenetic analysis. Finally, local Cryptosporidium species from domestic pigeon isolates were submitted to the National Center for Biotechnology Information GenBank and identified by accession numbers (PV478411-PV478421). Figures 2 and 3.
Fig. 1. Agarose gel electrophoresis showing nested PCR amplification of the 18S rRNA gene region in Cryptosporidium species isolated from domestic pigeon feces. The M markerindicates DNA size standards, with the positive PCR products identified at 830 bp. DiscussionCryptosporidium is one of the most important protozoan parasites affecting birds and infects several hosts, including both wild and domesticated avian species. Pigeons (Columba livia domestica) are commonly found in urban and agricultural environments, making them natural candidates for multiple Cryptosporidium species, including C. avium. Several Cryptosporidium species were detected in fecal samples collected from domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. The results revealed a high level of genetic similarity between the identified species and previously known human-infecting Cryptosporidium strains, reinforcing the potential risk of bird-to-human transmission in urban settings. Encompassing the Cryptosporidium species that were documented in this study C. melegardis , C. bailey and C. avium .This consider the first documentation of C. avium in Iraq.
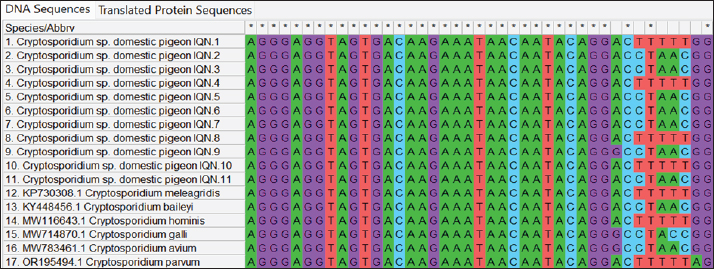
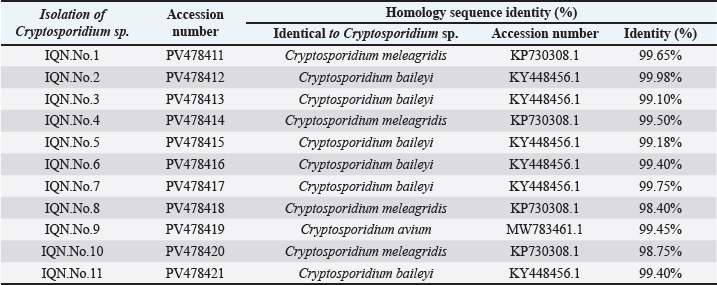
Fig. 2. Multiple sequence alignment analysis of the 18S ribosomal RNA gene in local Cryptosporidium species isolated from domestic pigeon NCBI-GenBank-related Cryptosporidium species isolates. Multiple alignment analysis was performed using the (ClustalW alignment tool. Online). The alignment analysis showed the nucleotide alignment similarity as () and substitution mutations in the 18S ribosomal RNA gene between isolates. Table 1. NCBI-BLAST homology sequence identity percentage between Cryptosporidium sp. domestic pigeon isolates and NCBI-BLAST closed genetic-related Cryptosporidium species isolates.
Cryptosporidium meleagridisCryptosporidium meleagridis is a significant parasite that principally infect birds in particular turkey, but it has high ability to transmission between animalis and human ,notably in immunocompromised patients. This species was recorded earlier in Babil province by Al- Tamimi and Al Zabaidi (2020), where it was recovered from domestic pigeon Columba livia domestica with high prevalence 61.9% by using nested polymerase chain reaction . This was regarded as the first scientific registration of C. meleagridis within Iraq particularly and prompts concern about it possible role in transmission of infection from birds to human especially in urban regions with high density. The outcomes of this study are agree with results of the study cited earlier, as C. meleagridis was also recorded from Columba livia domestica pigeon's fecal samples. This agreement corroborates the hypothesis that pigeons may play as a significant reservoir for this species in Iraq and shed light on the necessity for further scientific work into its epidemiological dynamics and its effects on public health. C. meleagridis was diagnosed in four regional samples (No 1,No 4, No 8, No10).Despite this parasite mainly infect birds but the risk of it transmission to human is highly documented.
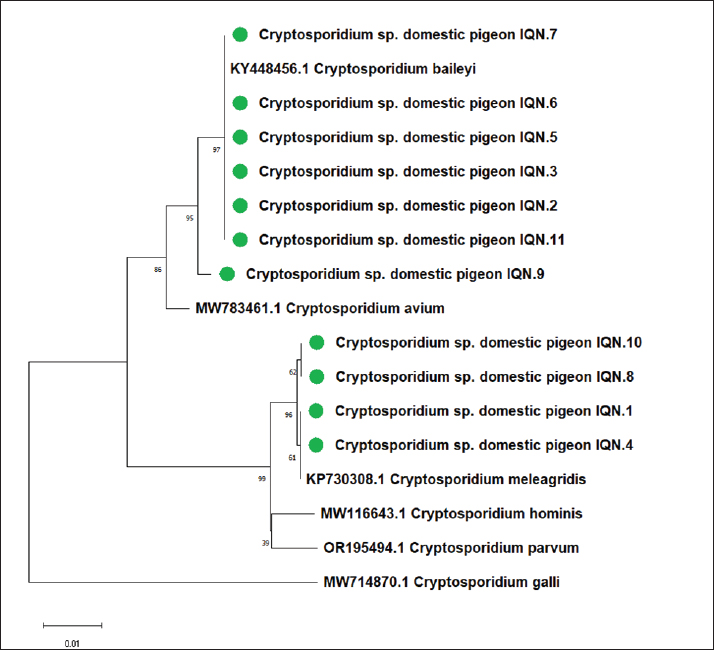
Fig. 3. Phylogenetic tree analysis based on 18S ribosomal RNA gene partial sequence in 18 local Cryptosporidium species from domestic pigeon used for genetic species typing analysis. The phylogenetic tree was constructed using the neighbor-joining method (MEGA 6.0 version). Local Cryptosporidium sp. IQN-No.1, No.4, No.8, and No.10 isolates showed close related to NCBI-BLAST Cryptosporidium meleagridis (KP730308.1), the local Cryptosporidium sp. IQN-No.2, No.3, No.4, No.5, No.6, No.7, and No.11 isolates were closely related to NCBI-BLAST Cryptosporidium baileyi (KY448456.1), and the local Cryptosporidium sp. IQN-No.9 isolate was closely related to NCBI-BLAST Cryptosporidium avium (MW783461.1) at a total genetic change of 0.01%. The genetic sequencing of the specimens showed a 99.65%match with reference sequence from Gen Bank ( accession number : K p730308.1) transmission from birds to humans, particularly in urban environments where domestic pigeons are in close contact with the human population. Which suggesting a strong genetic like with human - pathogenic strains. This apprehensions of its transmission to human. Cryptosporidium baileyiCryptosporidium baileyi was identified in seven samples from this study (IQN-No.2, No.3, No.4, No.5, No.6, No.7, and No.11). This species is primarily associated with domestic birds, particularly chickens and turkeys. Although it is not a common cause of human cryptosporidiosis, it has been reported to cause infections in immunocompromised individuals. Genetic sequencing of the local isolates revealed a 99.75% identity with a poultry-derived reference sequence (GenBank accession: KY448456.1), indicating a high degree of genetic similarity. This genetic closeness reinforces the possibility of bird-to-human transmission, particularly in shared environments. Cryptosporidium baileyi is a well-documented avian Cryptosporidium species that infects the respiratory, gastrointestinal, and reproductive tracts of birds, particularly poultry (Nakamura and Meireles, 2015). Although not considered a primary zoonotic species, its presence in birds living near humans—especially in urban areas—warrants epidemiological attention. In Iraq, Al-Tamimi and Al-Zubaidi (2020) first reported C. baileyi in domestic pigeons Columba livia domestica in Babil Governorate, where their nested PCR-based study confirmed a 33.3% infection rate. This investigation represented the first molecular confirmation of C. baileyi in domestic pigeons in the country, highlighting their potential role as reservoir hosts for veterinary-relevant and possibly zoonotic Cryptosporidium species. Our findings are in agreement with those of the previous study, further supporting the credibility of the detection and emphasizing the importance of pigeons as potential carriers of multiple Cryptosporidium species, including C. baileyi. Beyond pigeons, C. baileyi has been detected in other Iraqi avian species. Hashim and Al Zubaidi (2023) conducted a study in Baghdad Governorate to investigate. the presence of Cryptosporidium spp. in quails (Coturnix coturnix) using both conventional microscopy and molecular techniques. The results revealed that 367% was positive and 18s rRNA gene sequencing verified the detection of C. Baileyi and GenBank accession No was MN410723.1. Moreover, Al- Ghezy and Al-Zubaidi (2020) demonstrated the presence of C. baileyi in domestic chickens Gallus gallus domesticus in Iraq. And this parasite was isolated from chickens which exhibited the digestive and respiratory signs and verified by nested polymerase chain reaction (PCR) and molecular sequence analysis. The combined results indicate that the C. baileyi occurs in more one bird species. This wide host range and adaptive response to environmental conditions the regard to the need for uninterrupted epidemiological monitoring in particular among birds in the nearby human and those reared in dense agricultural systems. Cryptosporidium aviumOur results disclosed C. avium in rock pigeon Columba livia domestica which help in a substantial contribution to perception of Cryptosporidium infections in host birds in Iraq. This species infect a broad spectrum of birds species and has formerly been recorded in both non domestic birds and captive birds in numerous countries. According to our knowledge C. avium has not been reported in any scientific publication in Iraq hence, its recorded our study represent the first evidence of this species in Iraq. The detection of C. avium in domestic pigeons prompts significant epidemiological implication notably urban areas which pigeon live near humans in spite of this parasite ,it no liked to human parasitism however ,its genetic homology with another species of animal- derived parasite refers to latent risk for interspecies infection especially immunocompromised persons. This increase our understanding of the variety of this parasitic genus in Iraq. And stand out the value of molecular tracking in both free- living birds and farm birds. Study considers the first record of C. avium in Iraq, this species found in sample No. 9. The results of genetic sequencing showed a 99.45% identify with C. avium isolate from birds and Gen Bank accession number was MW 783461.1 in spite of C. avium predominantly infects birds, it has been record variety reported in human. This high level of genetic similarity of C. avium points to this parasite has zoonotic transmission in particular in immunocompromised people. These results which record this species in pigeons are entirely match with many prior studies that used molecular diagnostic approaches. These outcomes contribute and aiding markedly in the growing epidemiological dataset about C. avium in metropolitan and peri- metropolitan regions and reinforce the hypothesis that pigeons may act as main host for this parasitic species. Wang et al. (2021) noted that C. avium is widespread in pigeons and parrots and their review in health framework emphasized the relevance if surveillance of aviam species in environments co- inhabited with humans owing to their probable role as vectors or reservoirs of Cryptosporidium species. This compatible with our results points to pigeon's contribution an important role in the transmission of C. avium in urban regions in this regard, Struthes (2024), in trusted source on diseases of ornamental birds, pointed out the pigeon's C. avium infections can lead to minor tissue pathological changes in liver , intestine and respiratory tract, a few cases despite lack of symptoms in most of time. This note assert the value of molecular diagnosis for detection of subclinical infection as is it in our study The recording of C. meleagridis in domestic pigeons refer to likely peril of infection transmission to human which necessitated more studies and tracking to evaluate its contribution in human infections. Genetic identity between local isolates and human-infecting speciesMolecular sequences of local Cryptosporidium isolates were matched with reference sequences which recorded in NCBI GenBank. Analytical results indicated high level genetic similarity with recognizes species that infect humans among which C. meleagridis, C. baileyi and C. avium. The percentage of genetic match varied and C. avium. The percentage of genetic match varied from 98.50% and 99.75% which implies a tight evolutionary relatedness between the regional isolates and species with zoonotic potential in spite of the C. avium is not usually infect human nevertheless, the considerable genetic identity with human infected isolates under lights its likely cross- species transmission mainly in immunocompromised people . Nucleotide variationsEven though the results showed strong genetic resemblance, minimal genetic variation were detected in the 18s rRNA gene loci. The single – nucleotide mutations refer a minor degree of genetic similarity between the isolates obtained in current study and references sequences in GenBank. Even though this mutations were minor, it may represent adaptive mechanisms that facilitate the parasite to remain alive in changed environmental circumstances in another different host. These modifications impact parasite transmission dynamics, host range, its zoonotic. This work provides a foundation for prospective studies concentrates on genetic mutation of Cryptosporidium in various birds species in Iraq. Extended studies about genetic relationship between the locals isolates and human- infecting isolates can offer meaningful evidence - based insights on routes of infection between humans and birds by shedding light on genetic variation of cryptosporidium in Iraq. This study adds more comprehensive understanding the parasite's epidemiology at the molecular level, and aids in the development of strategies designed in preventing and combating infection. This investigation is an important step in order to identify the characterization of Cryptosporidium species in pigeons Columba domestica livia in Iraq. It is noteworthy that C. avium is documents for the first time. ConclusionThis investigation is an important step in order to identify the characterization of Cryptosporidium species in pigeons Columba domestica livia in Iraq. It is noteworthy that C. avium is documents for the first time. AcknowledgmentsWe extend our sincere thanks to the biology department at the science college in university of Thi- Qar for the support, facilities to conduct this work. Conflict of interestAll authors declare no conflicts of interest that might influence the study results or their interpretation. FundingThis research did not receive any funding from funding agencies in the public, commercial, or not-for-profit sectors. Authors' contributionsFirst author: study design, sample collection, analysis, and data analysis. Second author: contributed to laboratory analyses, scientific review of the manuscript. Third author: contributed to data analysis, drafted and revised the final manuscript. Data availabilityAll data used in this study are available from the corresponding author and will be made available upon justified request. ReferencesAbdullah, S. 2024. Detection of Natural Infection with Cryptosporidiosis in Domestic Poultry from Sulaymaniyah Province/Iraq. Egypt. J. Vet. Sci. 55(1), 59–67; doi:10.21608/ejvs.2023.223117.1541 Al-Ataby, F.H. and Radhi, L.Y. 2025. Cryptosporidiosis: a comprehensive review of intestinal parasitic infection. Manuscr. Evol. Med. Nat. Sci. 2(1), 62–76. Al-Ghezey, M.R.K. and Al-Zubaidi, M.T.S. 2020. Molecular detection of Cryptosporidium parasite in chickens (broiler and layer) in. Plant. Arch. 20(2), 7680–7684. Al-Tamimi, M.K. and Al-Zubaidi, M.T.S. 2020. High prevalence of Cryptosporidium meleagridis in domestic pigeons (Columba livia domestica) raises the prospect of zoonotic transmission in Babylon Province, Iraq. Iraqi. J. Vet. Med. 44(E0), 7–13; doi:10.30539/iraqijvm.v44iE0.1012 Hashim, .F. and Al Zubaidi, M.T.S. 2024. Detection of Cryptosporidium infection in domestic pigeons in Baghdad City Using Conventional PCR Technique. Iraqi J. Ind. Res. 11(1), 118–122; doi: 10.53523/ijoirVol11I1ID444 Hashim, Y.F. and Al Zubaidi, M.T.S. 2023. Cryptosporidium species diagnosis in handlers of domestic pigeons in Baghdad City: molecular and microscopic approaches. J. Adv. Vet. Anim. Res. 10(4), 744–750;doi: http://doi.org/10.5455/javar.2023.j730 J. Yahya, K. and T. S. Al-zubaidi, M. 2023. Traditional and molecular detection of Cryptosporidium parasite in quails (Coturnix coturnix) in Baghdad province, Iraq. Adv. Anim. Vet. Sci. 11(6), 864–869; doi: 10.1016/j.advvs.2011.09.010 Liao, C., Wang, T., Koehler, A.V., Hu, M. and Gasser, R.B. 2021. Cryptosporidium of birds in pet markets in Wuhan city, Hubei, China. Curr. Res. Parasitol. Vector. Borne. Dis. 1, 100025; doi:10.1016/j.crpvbd.2021.100025 Maktoof, A.A., Alrikaby, N.J. and Abul-Doanej, H.A.I. 2023. Trace element concentration in Raillietina echinobothrida and Aporina sp. in comparison to their host tissues dove, Streptopelia senegalensis, collected from Nasiriya city, southern Iraq. Iran. J. Ichthyol. (Special. Issue. 1). (1), 1–4. Mcconkey, G.A. 2024. Editorial: rising stars in parasite and host: 2023. Front. Cell. Infect. Microbiol. 14, 1509107; doi:10.3389/fcimb.2024.1509107 Meinhardt, P.L., Casemore, D.P. and Miller, K.B. 1996. Epidemiologic aspects of human cryptosporidiosis and the role of waterborne transmission. Epidemiol. Rev. 18(2), 118–136; doi:10.1093/oxfordjournals.epirev.a017920 Nakamura, A.A. and Meireles, M.V. 2015. Cryptosporidium infections in birds - a review. Rev. Bras. Parasitol. Vet. 24, 253–267. Quadros R.M. de., Marques, S.M.T., Amendoeira, C.R., Souza LA de., Amendoeira, P.R. and Comparin, C.C. 2006. Detection of Cryptosporidium oocysts using by auramine and Ziehl–Neelsen staining. Parasitol. Latinoam. 61(3-4), 117–120; doi: 10.4067/S0717-77122006000200003 Rafiei, A., Rashno, Z., Samarbafzadeh, A. and Khademvatan, S. 2014. Molecular characterization of Cryptosporidium spp. Isolated from immunocompromised patients and children. Jundishapur. J. Microbiol. 7(4), e9183; doi:10.5812/jjm.9183 Struthers, J.D. 2024. Doves and pigeons. In Pathology of pet and aviary birds. Tully, T.N. and Dorrestein, G.M Hoboken, NJ: Wiley-Blackwell; doi: 10.1002/9781119650522.ch15 Wang, Y., Zhang, K., Chen, Y., Li, X. and Zhang, L. 2021. Cryptosporidium and cryptosporidiosis in wild birds: a one health perspective. Parasitol. Res. 120(5), 1615–1628; doi:10.1007/s00436-021-07289-3 | ||
| How to Cite this Article |
| Pubmed Style Alrikaby NJ, Hami HJ, Al-gorani AF. Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Vet. J.. 2026; 16(2): 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 Web Style Alrikaby NJ, Hami HJ, Al-gorani AF. Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. https://www.openveterinaryjournal.com/?mno=258756 [Access: February 27, 2026]. doi:10.5455/OVJ.2026.v16.i2.46 AMA (American Medical Association) Style Alrikaby NJ, Hami HJ, Al-gorani AF. Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Vet. J.. 2026; 16(2): 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 Vancouver/ICMJE Style Alrikaby NJ, Hami HJ, Al-gorani AF. Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Vet. J.. (2026), [cited February 27, 2026]; 16(2): 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 Harvard Style Alrikaby, N. J., Hami, . H. J. & Al-gorani, . A. F. (2026) Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Vet. J., 16 (2), 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 Turabian Style Alrikaby, Nuha Jabbar, Hasan Jasim Hami, and Amal F. Al-gorani. 2026. Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Veterinary Journal, 16 (2), 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 Chicago Style Alrikaby, Nuha Jabbar, Hasan Jasim Hami, and Amal F. Al-gorani. "Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq." Open Veterinary Journal 16 (2026), 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 MLA (The Modern Language Association) Style Alrikaby, Nuha Jabbar, Hasan Jasim Hami, and Amal F. Al-gorani. "Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq." Open Veterinary Journal 16.2 (2026), 1289-1296. Print. doi:10.5455/OVJ.2026.v16.i2.46 APA (American Psychological Association) Style Alrikaby, N. J., Hami, . H. J. & Al-gorani, . A. F. (2026) Molecular detection and genetic characterization of Cryptosporidium species in domestic pigeons (Columba livia domestica) in Nasiriyah, Iraq. Open Veterinary Journal, 16 (2), 1289-1296. doi:10.5455/OVJ.2026.v16.i2.46 |