| Research Article | ||
Open Vet. J.. 2025; 15(12): 6602-6621 Open Veterinary Journal, (2025), Vol. 15(12): 6602-6621 Research Article Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samplesKhilood Hamdan Fahad, Jenan Nadhim Sadeq* and Azhar Abdulsada NeamahDepartment of Microbiology, College of Veterinary Medicine, University of Al-Qadisiyah, Al Diwaniyah, Iraq *Corresponding Author: Jenan Nadhim Sadeq. Department of Microbiology, College of Veterinary Medicine, University of Al-Qadisiyah, Al Diwaniyah, Iraq. Email: jinan.sadeq [at] qu.edu.iq Submitted: 13/05/2025 Revised: 29/10/2025 Accepted: 11/11/2025 Published: 31/12/2025 © 2025 Open Veterinary Journal
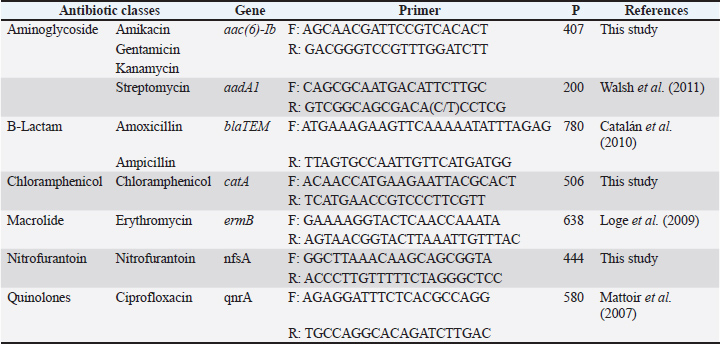
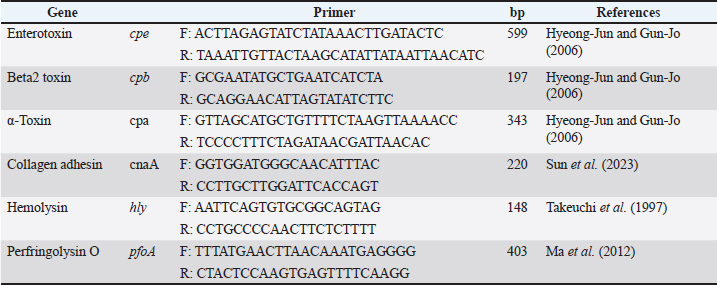
AbstractBackground: Clostridium perfringens is a common cause of foodborne and hospital infections. It produces several toxins and shows increasing antibiotic resistance. Aim: This study aimed to detect C. perfringens in human clinical samples and meat from local markets. This study also aimed to evaluate their antibiotic resistance and virulence gene profiles. Methods: A total of 188 samples were collected. Ninety-five came from humans. Ninety-three came from meat. Standard laboratory methods have been used to isolate bacteria. Identification of the 16S rRNA gene has been confirmed using biochemical tests and polymerase chain reaction (PCR). Ten samples were sequenced. MEGA6 software was used for the phylogenetic tree. Antibiotic testing was performed on 17 drugs. PCR was also used to find six virulence genes and eight resistance genes. Results: Of 188 samples, 76 were positive for C. perfringens. Thirty-six samples were from humans. Forty were from meat. Most patients showed strong resistance to erythromycin (90.78%) and tetracycline (84.21%). The top resistance genes were ermB (86.84%), tetA (81.57%), and blaTEM (65.78%). Some strains had up to eight resistance genes. The most prevalent virulence gene was perfringolysin O gene (76.31%). Hemolysin gene came next (73.68%), followed by cpe (50%). Both meat and human samples contained hypervirulent strains. RAPD-PCR showed that some meat and human strains were very similar. This may indicate cross-transmission. Conclusion: This study shows that meat and clinical samples share harmful C. perfringens strains that carry dangerous combinations of resistance and virulence genes. The results highlight a serious public health risk and call for stricter control of antibiotic use and better food handling hygiene to reduce transmission. Keywords: Anaerobe, Foodborne, Plasmid, Zoonosis. IntroductionSoils, water, raw animal meat, and even the intestines of various vertebrates, including humans, are all habitats of Clostridium perfringens, a bacterium of notable concern. These perfringens are infamous for the illnesses they cause, such as food poisoning and severe necrotic enteritis. Both animals and humans are susceptible to infections caused by the perfringens. Clostridium perfringens, like others of its ilk, generates formidable tissues that damage toxins and spores, the two features of its anatomy that are best suited for survival in ambient conditions. In perfringens, various toxins result in different diseases. Perfringens necrotic enteritis is ubiquitous and far from trivial in poultry, resulting in slowed developmental rates throughout the fowl population, increased mortality, and a financial burden from the exorbitant cost of treatment. The poultry industry suffers because of the exorbitant cost of treatment (García-Vela et al., 2023). For years, farmers in the poultry industry have incessantly used antibiotic growth promoters (AGPs) to control and promote growth. However, C. perfringens-infected chickens became resistant to treatment (Mohiuddin et al., 2023). While AGPs were banned, there was also a notable increase in the outbreaks of necrotic enteritis (Mohiuddin et al., 2023). Clostridium perfringens, like some other bacteria, is remarkably adaptable and has been discovered in rivers and lakes, some of which possess toxins and other resistant genes. It can survive in various environments and still be considered a threat (Fourie et al., 2020). One such study discovered such bacterial strains in the water. These bacteria are resistant to numerous antibiotics and possess a range of toxin genes (Li et al., 2024). These genes are associated with critical disease (Milton et al., 2020). In chickens, strains were identified during the food manufacturing process, providing insight into their transmission to humans (Xu et al., 2021a,b). Both C. perfringens and Clostridioides difficile pose a threat to healthcare facilities. They lead to gastrointestinal infections and demonstrate resistance to multiple antibiotics (Venhorst et al., 2022). Some meat specimens contained strains that produce a potent alpha-toxin (Ali et al., 2024). This particular toxin devastates cells, inflicts tissue harm, and causes severe diarrhea. Even companion animals may harbor resistant strains. In China, a study reported numerous resistance genes in strains associated with domestic cats and dogs (Fang et al., 2025). Infections are associated with a decline in milk and meat yield in cattle and sheep, leading to adverse health and food supply implications (Xiaoting et al., 2021). For instance, in Pakistan, the economic burden associated with C. perfringens rests with its diminished meat and milk productivity (Khan et al., 2021). Some strains associated with outbreaks of enterotoxin-induced food poisoning during periods of increased laxity in hospital caregivers, particularly in the context of digestive disorders, were identified. Scientists are currently investigating advancements in therapies. Phage therapy, where isolated bacteriophages are administered to specifically target and eradicate C. perfringens in a culture, completely bypassing antibiotics (Thanki et al., 2024). The ginger plant, and others that are similarly classified as ‟natural,” are also effective in decreasing the colonization of poultry by pathogens (Valdez et al., 2023). The genes that directly relate to infection severity are cpe, cpb, and netB. Clostridium perfringens has been found to be threatening with the ability to change expression patterns. (Milton et al., 2020; Li et al., 2024). The bacterium may vary between different meat strains and those taken from humans (Hassani et al., 2022; Duc et al., 2024). Furthermore, ‟C. perfringens employs mobile genetic elements, such as plasmids and phages, to horizontally transfer toxin and resistance genes to different bacterial strains” (Fourie et al., 2020; Aljindan et al., 2023). It is greatly spread due to food negligence. Primarily, unsanitary practices permit the progression of this organism. Precooked or preheated foods containing this spore from their unprocessed state will be difficult to kill. These spores have exhibited the ability to withstand extreme heat and flourish under such conditions (Fourie et al., 2020; AlJindan et al., 2023). In offering management to C. perfringens, there is a wider span, such as humans and animals, that must be coordinated. A One Health approach should be incorporated to support food safety and the welfare of the public (Valdez et al., 2023). The current study aimed to investigate the presence of C. perfringens in chicken meat and meat samples purchased from the local market. This study also aimed to determine their antibiotic resistance and virulence genes. Materials and MethodsSampling and collection of samplesFor this study, 188 samples were collected, of which 95 were from patients and 93 were of unprocessed meat obtained from Al-Diwaniyah city, Iraq. Out of the 95 samples collected, the predominant samples were feces and wound swabs of patients in Al-Diwaniyah Teaching Hospital, Al-Diwaniyah City, Iraq. Samples were collected under strictly controlled and sterile conditions. Clinical specimens were taken using sterile swabs, which were placed in sterile and appropriately labeled containers. Such containers were tightly closed and subsequently stored in cooled transport boxes with ice packs to retrieve and maintain bacterial viability until the designated time for laboratory examination. Identical procedures were followed for meat samples obtained from various market sites, which comprised beef, lamb, and chicken cuts. Meat samples were obtained by swab testing using sterile gloves and sterile plastic bags to maintain aseptic conditions. Each sample collection containing the sample bags was recorded using distinctive sample labels. Each sample code was recorded in a logbook to provide constant tracking of the sample from collection to the laboratory for analysis. Tracking the sample with respect to collection and laboratory testing helped in correlating the sample data with the obtained resistance and identification results. All samples were packed in transit cold boxes to protect them from moist heat and temperature excess. All samples were in the laboratory by the end of the collection day. The majority of responses were received within 4 hours of sampling. Any samples that were kept longer than 4 hours were stored in refrigeration at 4°C for 24 hours at most. The sampling strategy included negative controls for every trip. These controls were obtained by laying sterile swabs and containers in an environment that was not to be touched by any actual samples. This ensures that contamination was not present in the samples during the collection or handling stages. All the collection and transport methods were performed at a biosafety level of two. Protective gloves, laboratory coats, and disinfectants were used to perform these procedures. The work surfaces and tools were cleaned and disinfected before and after use. Participants from the study provided samples along with the attending medical personnel. Every minute detail, such as the time and location of the samples, was recorded and kept in a safe location. Meat samples were sourced from different geographical areas in the city to cover a wide area for the assessment of the contamination level within the city. The study was designed to evaluate the presence of C. perfringens in specimens obtained from both human and animal origins. This also facilitated the study of the similarities of C. perfringens, which are of a genetic or resistance nature from both origins. This was vital to apprehend the potential spread of infection from the meat products to humans. Broad sampling enhanced the results’ dependability while reducing the potential sampling bias. It also established the framework for all subsequent laboratory analyses. Isolation and identification of C. perfringens strainsAfter receiving the specimens as samples, the bacteria were placed in the cooked meat. This step helps Clostridia and other anaerobes grow. The cooked meat is kept in an anaerobic condition at 37°C for 24 hours. The samples were then placed in blood agar with 5% sheep blood. They were kept in specially designed anaerobic jars with gas-generating kits at 37°C for 24 hours. Clostridium perfringens forms huge, irregular, grayish colonies with a double alignment of hemolysis zones and a complete inner and partially outer zone. The rest is for identifying and then confirming the presence of the suspected bacteria. They were later confirmed to be differentiated by the shape of the colonies, the scent, and the pattern of hemolysis (Rana et al., 2023). Other tests confirming the presence of the bacterium were gelatin liquefaction, nitrate reduction, lactose fermentation, and lecithinase activity. These were all steps for achieving positive identification. When all was done, the suspected DNA was isolated and purified with the set boiling lysis methods. The 16S rRNA gene was amplified using the universal primers 27F and 1492R. Polymerase chain reaction (PCR) was performed in a 50 μl reaction volume, which comprised the template DNA, Taq polymerase, the four dNTPs, MgCl2, the buffer, and the primers. The amplification cycles began with an ‟initial denaturation step” set at 94˚ for 10 minutes, followed by 35 cycles of denaturation at 94°C for 1 minute, annealing at 55°C for 1 minute ‟annealing” and at 72°C for 1 minute ‟extension.” The PCR sample was subjected to 1% agarose gel and ethidium bromide to visualize the results. Examination of the results was performed using ultraviolet (UV) light. A band that illuminated at 1,500 bp indicated the 16S rRNA gene that was amplified during the PCR process. From the samples, 10 were selected for sequencing. Sequences were aligned with the GenBank database using the BLAST program. Remarkably, all isolated and sequenced samples were similar to Clostridium perfringens reference strains (et al., 2023). Antibiotic susceptibility testingAll bacterial cultures were grown overnight before the application of the disk diffusion method of Isolation of Antibiotic-Resistant Bacteria on Mueller-Hinton agar plates (EUCAST, 2024). Antibiotic resistance testing was carried out using Antibiotic discs (ADS) on agar plates, and the results were recorded per the guidelines prescribed by the European Committee on Antimicrobial Susceptibility Testing (EUCAST, 2024). Bacterial suspensions were prepared at a concentration of 1.5 × 108 CFU/ml (or 0.5 MacFarland standard), and sterile saline was used for the dilution. Antibiotic suspensions, which were prepared in accordance with the Antibiotic Susceptibility Testing, were placed on the surface of Mueller-Hinton Agar using sterile forceps. Amikacin, Gentamycin, Kanamycin, Streptomycin, Amoxicillin, Ampicillin, Cefotaxime, Ceftriaxone, Erythromycin, Azithromycin, Chloramphenicol, Nitrofurantoin, Ciprofloxacin, Tetracycline, and Doxycycline were the different antibiotics used during the experiment. The plates were incubated for 24 hours at a temperature of 37°C, and the results were analyzed under anaerobic conditions. The separated colonies were measured in millimeters and assessed as sensitive, intermediate, or resistant. All antibiotic resistance profiles were recorded, and a multiple antibiotic resistance index was calculated. The MAR Index is defined as the ratio of the number of antibiotics to which an isolate was considered resistant over the total number of antibiotics tested, as resistance in a proportionate manner. More than three classes of resistant antibiotics were defined as MDR. Some applied samples that were completely resistant to the antibiotics were defined as pan-resistant. Data collected from both clinical and meat isolates were weighed against each other. This indicated possible cross-resistance between pathogenic and foodborne microorganisms. The parallels indicated the cardinal possibilities of a common origin and horizontal gene transfer. To document reproducibility, every set of tests was completed with negative controls and QC strains. All tests were executed again to answer certain ambiguous results. All of the resistance data were subsequently analyzed in the extraneous electronic databases for comprehensive statistical interpretation. Molecular detection of genes with virulenceTo assess the pathogenicity of each isolate, virulence genes were screened using PCR amplification. DNA was isolated from confirmed colonies using boiling lysis. The supernatant was the DNA template. Primers were designed to amplify six virulence genes (Table 1). These were cpe (enterotoxin), cpb (beta2 toxin), cpa (alpha-toxin), pfoA (perfringolysin O), hly (hemolysin), and cnaA (collagen adhesin). The primer design and amplification conditions, including annealing temperatures, were derived from established protocols. Multiplex PCR amplification was performed for each gene. The 25 μl reaction was composed of template DNA, primers, Taq polymerase, buffer, dNTPs, and MgCl2. A set of PCR conditions was defined as optimal for each gene. DNA amplification products were analyzed through electrophoresis on 1.5% agarose gels. The gels were subjected to ethidium bromide staining and UV light visualization. A 100 bp DNA ladder was used to determine the band size. The presence of the genotype was based on the presence of specific amplification products (bands). Accuracy was maintained in each run using positive and negative controls. Only unambiguous, distinct bands were diagnosed as positive (Rana et al., 2023). Table 1. Primers used for detecting antibiotic resistance genes in C. perfringens.
Among the virulence genes analyzed, cpe and pfoA were more frequently detected in clinical specimens than in meat and human samples. Other isolates, such as cpb and hly, were found in greater frequencies in meat isolates. Some isolates contained four or more genes associated with virulence. These were termed hypervirulent and analyzed separately. Their presence in both meat and human samples is alarming from a public health perspective. Gene association patterns were also studied. Some patterns were found multiple times, indicating either clonal dissemination or selective environmental pressure favoring certain sets within the gene pool. This portion of the investigation displayed the diversity of virulence in the C. perfringens population. It also illuminated why the potential for disease varies among the isolates. Every gel image was captured and is cataloged for easy access. Detection of antibiotic-resistance genesThe PCR technique was also used to assess the resistance of the genes. The individual isolates were subjected to a modified boiling lysis protocol for DNA extraction. Primers were designed and synthesized to target C. perfringens’ critical and widely described resistance genes (Table 2) (Li et al., 2021). The genes included ermB (conferring resistance to macrolides and related compounds), tetA (conferring resistance to tetracycline), blaTEM (conferring resistance to β-lactams), aadA1 (conferring resistance to streptomycin), aac(6)-Ib (conferring resistance to aminoglycosides), catA (conferring resistance to chloramphenicol), qnrS (conferring resistance to quinolones), and nfsA (conferring resistance to nitrofurantoin). Each reaction was performed in a total volume of 25μl and contained the DNA template, primers, buffer, MgCl2, Taq polymerase, and dNTPs. The PCR condition was adjusted for the genes, and the rest of the parameters were kept fixed. The annealing temperatures were determined from the literature (Table 2). Table 2. Primers used for detecting virulence genes in C. perfringens isolates.
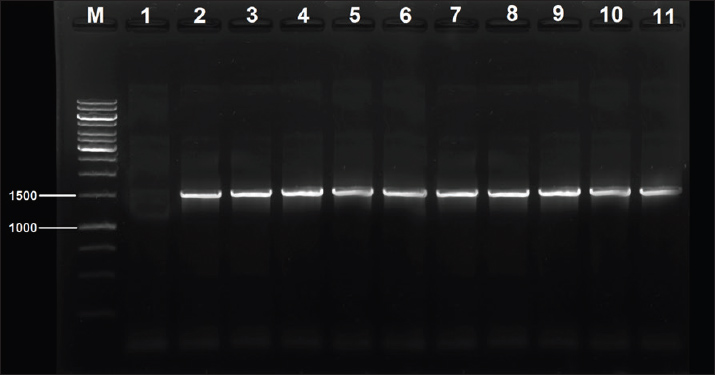
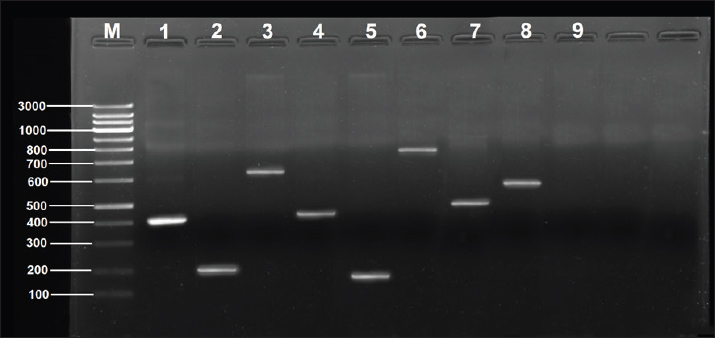
The amplification products were resolved by electrophoresis on 1.5% agarose gels and subsequently stained with ethidium bromide. Migrations were compared using a 100-bp DNA ladder. The presence of the bands was positive only when their dimensions were expected. The proportion of each observed trait was determined and compared between the different cohorts. Certain PCR procedures were performed on random samples to check the methods’ reproducibility. Gels and documents were prepared for later verification. Molecular typing and genetic analysis (GTA)To assess genetic diversity, molecular typing was conducted using random amplified polymorphic DNA–polymerase chain reaction (RAPD-PCR) and enterobacterial repetitive intergenic consensus–polymerase chain reaction (ERIC-PCR). These techniques enabled the investigators to pinpoint genetic variances among isolates representing distinct geographical locations. RAPD-PCR was performed using random sequence primers. Each isolate displayed distinct patterns of amplified DNA bands. These patterns were recorded as images and scanned onto 2% agarose gel. ERIC-PCR relied on repetitive genome sequences (Abdel-Glil et al., 2021). This technique enabled the classification of isolates based on genetic similarities. All reaction components were set at 25 ml volumes per reaction. UV light was cast within the gel containing the emitted DNA bands. The captured images were archived as scanned images. Specialized pattern analysis band software was used to evaluate the captured patterns. Dendrograms were created using the unweighted pair group method with arithmetic mean method. The phylogenetic tree was constructed using MEGA 6 software, and the genetic relatedness among the isolates was analyzed. Some isolates from human and meat sources showed >80% resemblance. This indicated a possible linkage or source crossing over. The remains were more distantly related, suggesting separate genetic origins. The outlier strains were also readily apparent from the most closely related strains’ cohesive genetic structure. The most closely related isolates also tended to share the same sets of virulence and resistance genes. This result reinforced the hypothesis of the clonal expansion of successful strains. Statistical analysisSPSS software, version 25, was used to conduct the analyses of all the data sets, which to a large extent were filled in various spreadsheets. The data sets were then subjected to expository and inferential statistical analyses. The chi-square test was used to evaluate the frequencies of resistance and virulence genes isolated from human and meat sources. A p value of 0.05 was the cutoff point in the calculations that determined the statistical significance of the data. Confidence intervals (95% confidence interval) were determined when appropriate. The association between phenotypic resistance and resistance genes was assessed using the Pearson correlation test. Ethical approvalThe current project was approved by the committee for Research Ethics at the College of Veterinary Medicine, University of Al-Qadisiyah, Iraq (May 12, 2024; No. 2046). ResultsSampling and collection of samplesMicroscopy after Gram staining revealed large Gram-positive rods occurring as singles or in chains. A total of 188 samples were analyzed, of which 95 were sourced from human clinics and 93 from locally sourced meat. Of the 188 samples, 76 were positive for C. perfringens. Among the human samples, 36 (37.89%) were positive. Among the meat samples, 40 isolates (43.01%) were positive, indicating that raw meat products are clinically more contaminated than clinical specimens. Three types of meat samples were taken from various locations to improve the representation of market contamination: beef, chicken, and lamb. The positive samples did not pertain to a single meat type, suggesting the absence of confinement to a particular animal, thus likely existing within the entire global meat supply chain. In the human specimens, the positive cases were mainly associated with gastrointestinal tract issues or infected wounds. The patients showed signs of C. perfringens infection, marked by abdominal pain or soft tissue damage. This high positivity rate indicates a high prevalence of infection with this bacterium among humans in the study area. Negative controls with no growth were achieved, and it was confirmed that the sampling and the following steps of the procedure were meticulously performed. The samples’ transport and processing did not show signs of contamination. This bolstered the credibility of the findings, thereby strengthening the defense of the employed sampling technique. Identification and molecular techniques were applied shortly after to isolate and characterize a few positive samples collected from meat and human sources. Such comparisons were vital in assessing potential commonalities in the source and distribution of the bacterial strains, suggestive of foodborne or environmental transmission. Isolation and identification of C. perfringens strainsEach of the 76 blood agar culture plates was isolated through anaerobically enriched primary cultures. Colonies demonstrate the unparalleled double-zone hemolysis associated with C. perfringens. Presumptive identification was performed on the colonies that were staged to subcultures and stained with Gram. All samples contained large gram-positive rods (Table 3). All isolates were subjected to PCR testing to assess the 16S rRNA gene. Each of the 76 isolates was amplified with the expected 1,500-bp fragment. All samples analyzed via gel electrophoresis exhibited distinct thin bands with absent non-specific amplification and background noise (Fig. 1).
Fig. 1. Amplification of 16S rRNA of 10 Clostridium perfringens isolates was fractionated on 1% agarose gel electrophoresis stained with ethidium bromide, Lane M: 1kb ladder marker; Lane1: negative control; Lane2: HC05; Lane3: HC16; Lane4: HC22; Lane5: HC34; Lane6: MC23; Lane7: HM41; MC32: MC15; Lane9: MC03; Lane9: HC27; Lane10: MC39. Table 3. Distribution of positive C. perfringens isolates from different sample types.
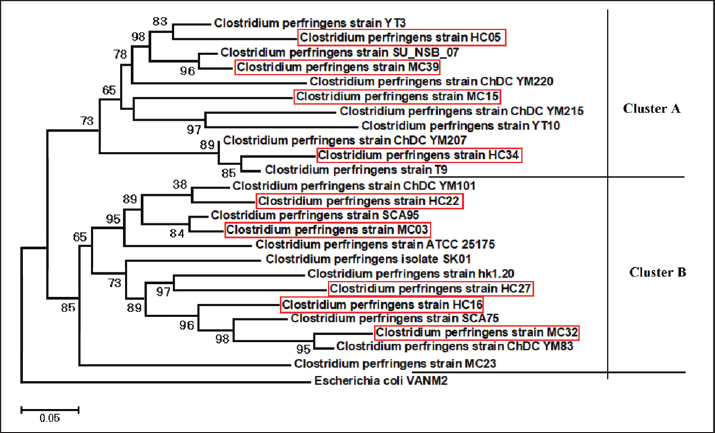
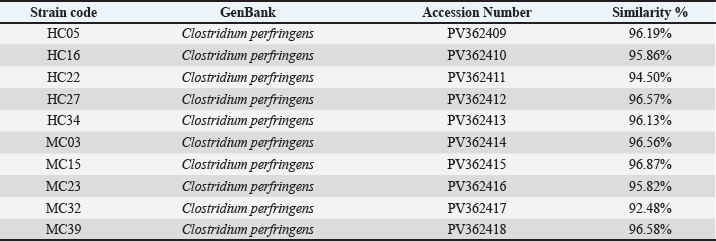
The isolates were, per the judge, C. perfringens. We randomly selected 10 isolates for 16S rRNA gene sequencing. BLAST analysis indicated that the sequences corresponded with C. perfringens varieties within GenBank, ranging in identity score from 92.48% to 96.87%. This not only affirmed the species classification but also indicated a strong relationship with C. perfringens strains from various geographic locations. These sequences contributed to a phylogenetic analysis in which the tested isolates comprised two primary clusters. Closer association was noted among certain meat and human isolates, suggesting a potential common vector for circulation or parallel evolutionary adaptation. Some strains were observed in different branches, which may illustrate strain diversity from different sources (Table 4 and Fig. 2).
Fig. 2. Phylogenetic tree showing the evolutionary relationships of the 16S rRNA gene of eight Clostridium perfringens species. Table 4. Comparison between isolate sequences with available GenBank sequences by similarity percentage using the BLAST function.
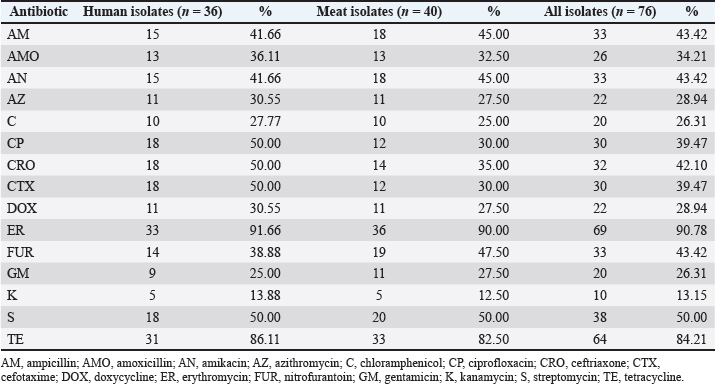
Antibiotic susceptibility testingEvidence from the disk diffusion method has indicated that several antibiotics have minimal effectiveness. Erythromycin demonstrated the greatest resistance (no inhibition, 91.66% in human isolates; 90% in meat isolates), with the highest absolute resistance recorded. The widespread tetracycline resistance (with 86.11% human isolates and 82.5% meat-derived strains) was also noted. These findings raise the important issue of antibiotics’ reckless use—even abuse—in human health care and agriculture (Table 5). Table 5. Antibiotic susceptibility of C. perfringens isolates.
Clinical isolates demonstrated the greatest resistance to ciprofloxacin, cefotaxime, and ceftriaxone. Human strains demonstrated moderate resistance, with 50% of isolates exhibiting resistance from meat (30%–35%). This suggests that antibiotics are more resistant in clinical strains (which are possibly subjected to more extreme conditions). The resistance to furazolidone in meat samples (47.5%) has been linked with the historical use of this drug in livestock and is much greater than that in human samples (38.88%). The observation of low resistance to the more active beta-lactam antibiotics, including ampicillin, amikacin, and amoxicillin, was consistent with moderate resistance (32.5%–45%), with little differences for human and meat isolates. Very low resistance to kanamycin and streptomycin was observed. Discrepancies, such as the 50% resistance rate reported in the subgroup analyses and the approximately 13% in the general combination totals, recorded in the results of the streptomycin type of analyses, arouse serious concerns. Such inconsistencies are indicative of technical deficiencies, complications in resolving batch differences, and other unconsidered research elements. Regarding the overall resistance profile, the greatest concern came from the sheer number of isolates attributed to the resistance of multiple agents. With many strains proving to be MDR and a few displaying resistance to all of the agents tested, the threat posed from a public health and treatment standpoint was indeed alarming. Molecular detection of genes with virulenceThe 76 isolates revealed that over 80% of the clinical samples and 70% of the meat samples contained the pfoA virulence gene under strain, and the pfoA was the most dominantly detected variation associated with its strain. The hemolysin gene (hly) came second to pfoA at 66.66% prevalence with clinical samples and considerably more at 80% of the meat isolates. The elevated presence of meat isolates might be a consequence of the selective pressure exerted by the husbandry environment and the surrounding biome. The enterotoxin gene (cpe) was detected in 63.88% of human isolates but only in 37.5% of meat isolates. Enterotoxigenic strains are more common in human infections than in raw meat, possibly due to adaptation or host-specific factors. Alpha-toxin (cpa) was present in 50% of meat isolates and 41.66% of clinical isolates. The differences were not extreme, indicating that this gene is widely distributed in various environments and hosts. The beta2 toxin gene (cpb) and the collagen adhesin gene (cnaA) were less common. The cpb gene was found in 27.63% of all isolates, more frequently in meat samples. Overall, cnaA appeared in only 30.26% of isolates, slightly higher in clinical strains. Co-occurrence of multiple virulence genes, especially pfoA with hly or cpe, was common and identified in the majority of isolates (Figs. 3 and 4).
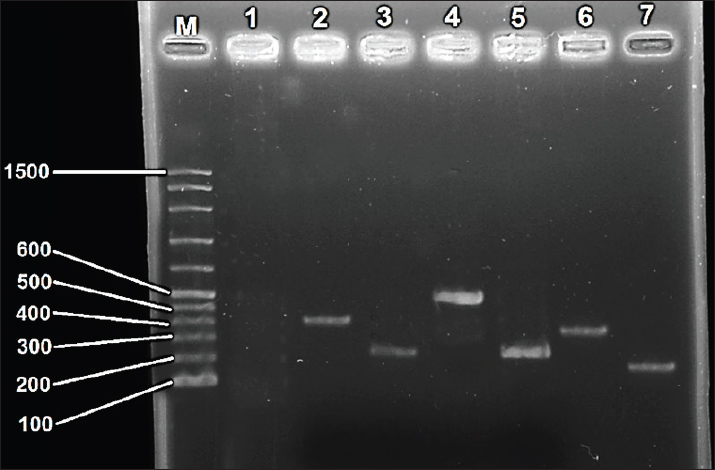
Fig. 3. Agarose gel image of virulence genes in Clostridium perfringens isolate HC16 by polymerase chain reaction (PCR) technique, electrophoresed on 1.5 % (w/v) agarose gel. LaneM: 100 bp DNA ladder. Lane 1: Negative control. Lane 2: pfoA gene. Lane 3: cnaA gene. Lane 4: cpe gene Lane 5: cpb gene Lane 6: cpa gene Lane 7: hly gene.
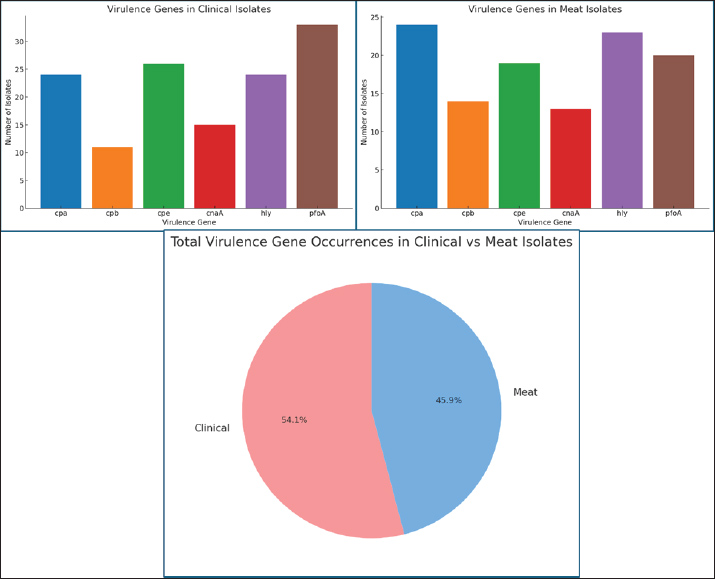
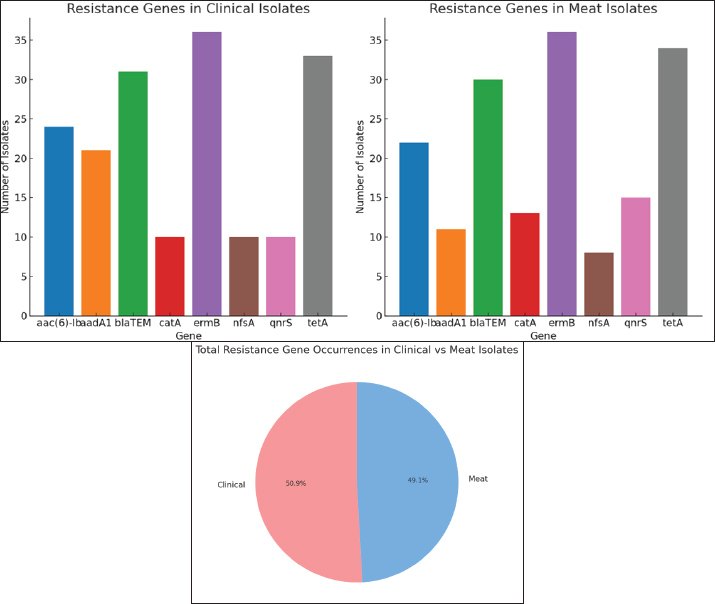
Fig. 4. Virulence gene in C. perfringens isolates from clinical sources and meat from markets. Detection of antibiotic-resistance genesThe PCR detection of resistance genes showed a wide distribution of multiple genetic determinants across the isolates. The macrolide resistance gene ermB was the most prevalent, found in 80.55% and 92.5% of human and meat isolates, respectively. This gene confers resistance to erythromycin and related antibiotics, matching the results of phenotypic resistance. The tetracycline resistance gene tetA was also widely present in 75% of clinical isolates and 87.5% of meat isolates. The β-lactamase gene blaTEM was found in 69.44% and 62.5% of human and meat isolates, respectively. This supports the continued decline in the effectiveness of penicillin-related drugs. Aminoglycoside resistance genes showed varying results. aac(6)-Ib appeared in approximately 43% of all isolates, whereas aadA1, which confers resistance to streptomycin, was found in 50% of human isolates but only 22.5% of meat isolates. This difference may be due to the selective use of human medicine over animal production. Other genes, such as qnrS (quinolone resistance), catA (chloramphenicol resistance), and nfsA (nitrofurantoin resistance), had intermediate to low prevalence. These genes were more common in meat isolates, suggesting that food supply resistance is emerging. Overall, 68% of isolates carried multiple resistance genes, often linked together, indicating a strong potential for horizontal gene transfer. ermB and tetA were the most common resistance genes and appeared in most isolates. These genes often occur together. Some isolates carried six or more resistance genes. These were defined as highly resistant and further analyzed. Many of them were obtained from meat samples, raising concerns about the use of agricultural antibiotics. Comparison between phenotypic resistance and the presence of genes showed high agreement. For example, erythromycin-resistant isolates almost always carried ermB. Statistical analysis helped identify gene clusters and common resistance patterns. These findings demonstrate that C. perfringens carries various resistance genes. This has serious implications for food safety and treatment (Figs. 5 and 6).
Fig. 5. Agarose gel image of antibiotic resistance genes in Clostridium perfringens isolate HC22 by polymerase chain reaction (PCR) technique, electrophoresed on 1.5 % (w/v) agarose gel. LaneM : 100 bp DNA ladder. Lane 1: aac(6)-Ib gene. Lane 2: aadA1 Lane 3: ermB Lane 4: nfsAgene. Lane 5: tetM gene. Lane 6: blaTEM gene. Lane 7: CatA gene. Lane 8: qnrA gene Lane 9: Negative control.
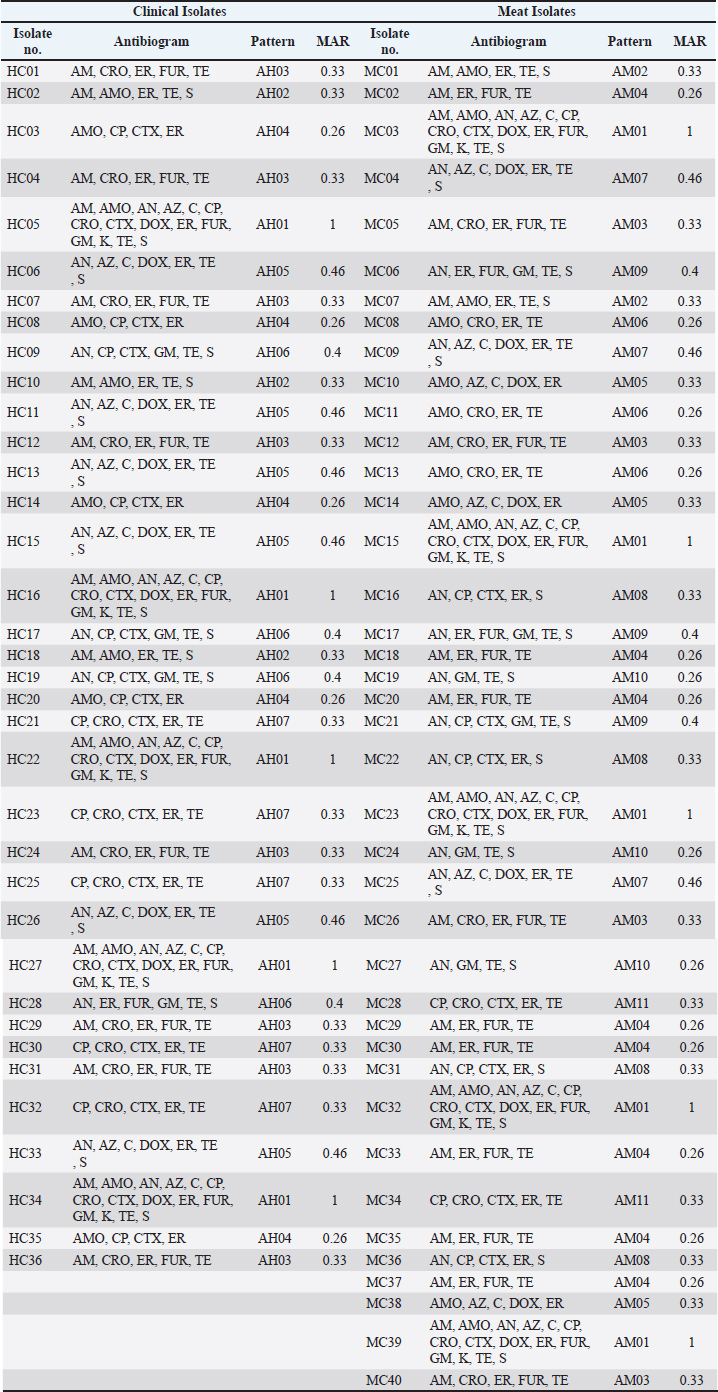
Fig. 6. Antibiotic resistance gene profile in C. perfringens isolates isolated from clinical and meat samples. Molecular typing and genetic analysisThe C. perfringens-type isolates analyzed using RAPD-PCR and ERIC-PCR revealed considerable genetic variance. The classification of multiple isolates as clusters suggests the presence of certain associates. The fact that isolates from human and meat sources are grouped with more than 80% similarity is of interest, indicating possible transmission or a shared source. The dendrograms resulting from the typing data consist of 2 major clusters. Cluster A contained a collection of clinical isolates with a high load of virulence genes. Cluster B contained a mixture of meat and clinical specimens, many of which had similar resistance gene content. The data supported the concept of intermingling strains from the food chain and human infections. Several isolates exhibited no phylogenetic relationships and unique genetic patterns. These may reflect more novel or rare strains. The rest were phylogenetically aligned, and their resistance and virulence profiles were dissimilar, indicating divergence in the species. The virulent and resistant strains were comparable. Hyper-resistant isolates are more likely to be in the same assemblage as those with multiple virulence genes. Indeed, the typing results further indicate that the C. perfringens strains in this study were not only cross-diverse but also adept at spanning across human and alimentary sources. This validates the necessity of a One Health perspective, in which bacterium oversight and management should integrate human, veterinary, and food safety (Table 6). Table 6. Antibiotic resistance pattern and MAR of C. perfringens isolates.
Molecular fingerprinting of C. perfringens by RAPD-PCR and ERIC–PCR typingIn this study, the genetic distinction for the C. perfringens isolates was completed using RAPD-PCR and ERIC-PCR. Both techniques for each of the 76 isolates used in the study resulted in distinct and distinct banding patterns. The number of bands in RAPD-PCR ranged from 3 to 10, depending on the primers used. Other ERIC-PCR had a more stable banding profile, with the majority of isolates showing 5–7 bands. Each isolate was then compared to establish mutational similarity. The cluster analysis of the banding patterns revealed many potential groupings. Some of the meat and human isolates were in close association with each other, suggesting that over 80% of the isolates were genetically similar, promoting the concept of copious cross-contamination or a shared environmental source. Several of the closely related isolates also contained common antibiotic resistance and virulence, which strengthens the case for transmission from different sources. Not all isolates were in close association with each other, with many clinical and meat isolates being distinct and separate, showing unassociated strains. These strains may have been sourced from disparate environmental sources or have been acquired through diverse mechanisms. The plethora of diverse banding patterns indicates that C. perfringens is not a clonal expression and has multiple environmental sources that are the result of food and clinical contamination. Overall, the meat isolates contained a marginally higher genetic variation than the clinical isolates. This might be due to the multiple species and environments in which meat is produced and disseminated. However, the overlap in some clusters showed that the transmission of some strains between meat and humans is not possible, but rather probable. The combination of the RAPD and ERIC-PCR techniques proved to be very useful for fingerprinting. The two provided sufficient strain differential analysis, confirming that C. perfringens strains share close genetic relationships in both raw meat and human infection cases (Fig. 7). This underscores the need for molecular typing to trace the infection pathways and analyze the peripheral infection mechanism of this bacterium within the population.
Fig. 7. Dendrogram of typeable Clostridium perfringens isolates produced from RAPD analysis using the average linkage unweighted group pair method with arithmetic averages (UPGMA). The dendrogram shows two main C. perfringens isolate clusters. The first large cluster includes strains from HC13 to MC32. The second main cluster starts at MC01 and ends at MC20. Both clusters contain strains from humans and meat sources. This mix suggests possible transmission between the two. Several subgroups have high similarity (>90%). For example, HC13, HC35, and HC24 are closely grouped. Another tight group includes MC25, MC37, and MC15. Some human and meat isolates cluster together. HC05 and MC21 are in the same branch. HC28, HC27, and MC40 also share a close position. These patterns indicate that meat and clinical isolates may be genetically linked. DiscussionIdentification of C. perfringens strainsThe ability of C. perfringens to be found in both human and meat samples indicates a risk of zoonotic transmission. The observation that nearly 43% of meat samples and almost 38% of clinical samples came out positive points to the routine exposure of people to this bacterium within a given population. This is consistent with the abundant documentation on the environmental distribution of C. perfringens in foods and clinics in Pakistan and China (Sárvári and Schoblocher, 2020; Xu et al., 2021a,b; Haider et al., 2022; Wu et al., 2023). The close range of detection rates in the clinical and meat samples reinforces the hypothesis that the clinical meat samples are a principal source of infection, especially if the meat is poorly cooked or handled improperly. Previous studies corroborate the observation that meat and poultry dominate the supply of livestock products and are a common source of antibiotic-resistant C. perfringens (Walsh et al., 2011; Xu et al., 2021a,b). The general perspective on C. perfringens, particularly in relation to public health, especially with the unavoidable ease of shedding the spores after improperly cooked meat, is worrying. Spore formation under conditions such as inadequate refrigeration results in vegetative growth, which leads to rapid multiplication and subsequent foodborne diseases (Catalán et al., 2010; Rajput et al., 2020). Double-zone hemolysis pattern on blood agar and various biochemical tests validated the isolates. Even in low-resource settings where molecular approaches are less commonplace, these conventional approaches are still reliable and inexpensive. Identification was confirmed using 16S rRNA PCR. Isolates yielded a 1,500-bp band, and alignment with C. perfringens strains was confirmed through sequencing. This approach assists with the larger goal of enhancing molecular epidemiology to strengthen pathogen surveillance of foodborne diseases (Ma et al., 2012; Sun et al., 2023). Interestingly, a number of sequenced isolates demonstrated >96% identity to reference strains, indicating a close genetic relationship to C. perfringens defined global strains. On the other hand, some of these sequences displayed 96% similarity, suggesting that they were locally adapted or emergent variants. This phenomenon is also supported by the genetic configurations of wild birds and duck farms, wherein distinct lineages have also been observed (Revitt-Mills et al., 2021; Xiu et al., 2022; Azevedo et al., 2024; Song et al., 2024; Wang et al., 2024a,b; Fang et al., 2025). Antibiotic susceptibility testing revealed alarming resistance patterns. More than 80% of examined subjects exhibited resistance to erythromycin and tetracycline. This corresponds to the resistance rates of macrolide and tetracycline class antibiotics among poultry in Pakistan and duck farms across Asia (Haider et al., 2022; Xiu et al., 2022). The authors suggested that the rampant use of antibiotics in both human and veterinary medicine could explain the high rates of resistance. The widespread use of antibiotics in agriculture is a noted factor in the development of resistant populations (Walsh et al., 2011; Rajput et al., 2020), as evidenced by the meat samples from this study. Resistance to ciprofloxacin and beta-lactams, both of which are used to treat gastrointestinal tract infections, is also a concern (Haider et al., 2022). The resistance patterns described in this study have also been noted in Hungary and other European countries, which is alarming (Sárvári and Schoblocher, 2020; Sun et al., 2023). Antibiotic resistance and gene profilingMost of the tested isolates showed untreatable resistance to almost all antibiotics. These strains that are resistant to multiple drugs are quite alarming. They demonstrate how foodborne pathogens transfer from the farm to the table (Gambino et al., 2021; Haider et al., 2022). The genetic determinants of ermB and tetA, which are the most prevalent among resistant isolates, demonstrate resistance to erythromycin and tetracycline. These genes are known to be present in macrolide and tetracycline resistance strains and in poultry, goats, and even wild birds (Ma et al., 2012; Wu et al., 2023; Song et al., 2024). The gene bla template, commonly referred to as the beta-lactamase gene, was detected in more than 60% of the isolates, suggesting that lateral gene transfer resulted in both pule and meat strains. This gene has also been identified within the mobile constituents of C. perfringens and related pathogenic organisms (Revitt-Mills et al., 2021; Wu et al., 2023). Strains aac (6)-Ib and aadA1 showed a higher prevalence in clinical strains because of rampant antibiotic use in hospitals. These aminoglycoside-modifying enzymes are still sparsely documented in food sources, but their occurrence in the present meat samples suggests their dispersal across different environments (Haider et al., 2022; Song et al., 2024). The presence of catA and qnrS in meat isolates demonstrates the circulation of both older and newer mechanisms of resistance (Watts et al., 2022). This situation creates difficulty in selecting the appropriate treatment, particularly when multiple resistance genes are present in the same isolate (Wu et al., 2023). Revitt-Mills et al. (2021) and Watts et al. (2022) found nfsA in both human and meat isolates, indicating the presence of a plasmid-bearing gene associated with the transfer of NF resistance, albeit rarely used in veterinary medicine (Revitt-Mills et al., 2021; Watts et al., 2022). The virulence gene profiling (VGM) study revealed that pfoA and hly are the most prevalent virulence genes. The degree of virulent strain distribution outside the clinical setting is corroborated by immune system evasion coupled with tissue destruction (Ma et al., 2012; Haider et al., 2022). The higher occurrence of enterotoxin cpe in human isolates is a well-known phenomenon, as it is frequently linked with cases of foodborne illness (Wu et al., 2023). However, cpe detection in meat isolates signifies a potential risk of infection via cross-contaminated food (Catalán et al., 2010). The clinical relevance of these cnaA and cpb genes, even though their occurrence is much rarer, is of interest. Their presence in foodborne isolates bolsters the case in meat as a reservoir of highly virulent strains (Haider et al., 2022) alongside the cnaA and cpb genes, which encode severe tissue-invasive components like beta2 toxin and collagen binding proteins (Song et al., 2024). Virulence gene profilingFour or more virulence genes were detected in select isolates. Hypervirulent types were confirmed in both the human and meat sources. Such types are even more worrisome in relation to immunocompromised persons and mirror what has previously been reported in goats and poultry (Ma et al., 2012; Rajput et al., 2020). The presence of resistance and virulence genes in a single isolate is a concern. This results in bacteria that are difficult to treat and can cause severe infections. Evidence from wild and domestic birds indicates that such strains are already prevalent in the environment (Gambino et al., 2021; Song et al., 2024). Some human and meat isolates showed unambiguous relationships through RAPD and ERIC-PCR molecular typing. These cross-contaminated networks of nearly identical strains from diverse origins indicate transmission in both directions. Other studies have also revealed strain similarity across different production stages (Xiu et al., 2022). Molecular typing and transmissionNot all isolates shared genetic relationships, with some being distinctly different and forming different clusters. This suggests that C. perfringens has multiple non-food sources. There may be environmental and animal reservoirs that contribute different strains to the population (Xu et al., 2021a,b; Gambino et al., 2021). Band patterns were more consistent with ERIC-PCR than with RAPD. This implies that ERIC elements may be more reliable genomic indicators for the typing of this particular bacterium. However, it must be noted that the use of both approaches together yielded a more comprehensive understanding of strain diversity, which is important for epidemiological tracing (Xiu et al., 2022; Song et al., 2024). Some clusters were made up exclusively of hyper-resistant and highly virulent strains. This association provides evidence that these genetic lineages are more harmful than those of other lineages. These results align with the theory that some C. perfringens clades are more dominant for surviving antibiotics and penetrating hosts (Watts et al., 2022; Wu et al., 2023). There are some C. perfringens strains that are anaerobic; however, these strains are infrequently encountered. Anaerobic isolates of C. perfringens that have some tolerance to oxygen are thought to survive for longer periods than the majority of C. perfringens strains that exclusively survive in hosts (Song et al., 2024). Oxygen exposure can also trigger mutation and stress responses (Sun et al., 2023). These phenomena may contribute to resistance development under anaerobic conditions. This demonstrates the increased versatility of C. perfringens in the development of resistance (Hyeong-Jun and Gun-Jo, 2006). The ongoing circulation of such a strain continues to support the necessity for new forms of treatment. Phytochemicals, such as ethanolic thyme extract and cinnamaldehyde, have been shown to reduce the load of C. perfringens and enhance gut immunity in broilers (Cattoir et al., 2007; Walsh et al., 2011). Phage therapy is also in progress to advance the control of C. perfringens. Certain G-type C. perfringens-targeting bacteriophages disinfect infected chicken meat (Catalán et al., 2010; Wang et al., 2024a,b). The use of postbiotics and specific probiotics, such as Lactococcus garvieae, has been proposed to enhance the intestinal barrier and immune response to improve resistance to C. perfringens infection (Ozma et al., 2025). These methods have been proposed for use in contemporary livestock systems (Wang et al., 2024a,b). A notable example is the pCW3 conjugative plasmid family. These plasmids are readily accepted in the community due to their ability to import many mobile genetic elements, thereby providing the recipient cell with increased versatility. These plasmids also bear the feared toxin and antibiotic resistance fusion, making them hazardous vectors (Revitt-Mills et al., 2021; Wu et al., 2023). Inherent limitations in the advancement of techniques to slow the spread of plasmid-mediated vectors. Recent investigations have indicated that the compatibility of plasmid replication proteins with plasmids and their particular plasmid-coexistence-associated proteins can be exploited to inhibit plasmid gene flow (Revitt-Mills et al., 2021). Wild birds and feral mammals have also been considered C. perfringens reservoirs that advance the spread of certain resistant strains along the ecosystems and food chains (Watts et al., 2022). There is a need to strengthen C. perfringens surveillance systems. The genetic and functional diversity of the organism requires enhanced surveillance. Monitoring the evolution and spread of virulent strains is increasingly reliant on multilocus sequence typing and whole-genome sequencing (Karim et al., 2018; Yaseen et al., 2020; Qui, 2023; Maty, 2023; Hassan et al., 2024; Rahawi et al., 2024). The strong antagonism of foodborne pathogens, including C. perfringens, displayed by some lactic acid bacteria isolated from fermented products further demonstrates the ability of natural antagonists to serve as food biopreservatives or feed to biopreserve food systems (Karim et al., 2018; Habeeb, 2018; Yaseen et al., 2020; Al-Deresawi et al., 2022; Ghazi et al., 2024; Habeeb, 2024). Taking all the information above into account, we can conclude that C. perfringens is an organism of public health and food safety concern. Foodborne outbreaks and possibly direct contact transmission emphasize the need for a unified health approach to mitigate their spread across humans, animals, and the environment. These profiles lacked associations with others, suggesting that they originated from more uncommon sources or represent novel lineages. The typing results contributed to the deepening of the study’s understanding. It unified previously observed genetic frameworks with the phenotype exhibited in other assay components. The C. perfringens population structure of the region is insightful, and the typing techniques applied are reproducible, economical, and useful. The high frequency of ermB and tetA genes in meat isolates is possibly associated with the use of antibiotics in animal husbandry. Erythromycin and tetracycline are widely used in livestock for growth promotion and disease control. However, the long-term application of these antibiotics to animals creates a selection pressure for the development of resistance. These resistance genes can persist in the gastrointestinal microbiota of animals and be transferred to the meat. This may explain the higher proportions of ermB and tetA in meat isolates than in clinical specimens. The results of this study confirmed previous work done on chicken and beef farms, emphasizing the need for prudent antibiotic control in livestock farming. The combination of human and meat samples into the same RAPD clusters suggests possible cross-transmission. However, RAPD-PCR is a relatively low-resolution technique and cannot adequately establish direct connections between the sources. While other techniques, such as whole genome sequencing, offer increased precision, the information presented suggests a finite set of sources or contacts that remain unaccounted for. Contaminated meat, unsterilized tools, or poor kitchen sanitization are plausible transmission pathways that need further validation. It is relatively uncontroversial to state that the resistance gene and phenotypic resistance alignment with the presence of resistance determinants is exceptionally high. For instance, ermB-positive strains displayed resistance to erythromycin, whereas the same trend was observed with tetA and tetracycline. Nevertheless, the trend is not universal. A number of strains with the resistance gene emerged and remained sensitive, whereas some sensitive strains exhibited resistance despite the absence of the corresponding gene. This could be accounted for by controlling the expression of the genes, as well as genetic elements that are unnoticed in the comprehensive PCR panel. Resilience mechanisms, such as efflux pumps or uncharacterized resistance genes, cannot be dismissed. These results are a threat to public health. Clostridium perfringens isolated from meat showed numerous resistance and virulence genes, which is highly concerning. Microorganism strains can pose a potential threat if transmitted to humans. This reinforces the need to implement strict hygiene measures and safety protocols during meat product handling and selling. Governments and relevant authorities need better training to enhance inspection and kitchen hygiene standards. Furthermore, better surveillance of resistant strains of microorganisms in food and humans is required. These measures may reduce an outbreak’s likelihood. This study is not without its limitations. Indeed, RAPD-PCR is a low-resolution technique compared with much more advanced methods, such as WGS. While it may suggest direct transmission, it cannot ascertain it. Of the 10 isolates that were sequenced—which does not encapsulate the total diversity—barely any were postulated to exist. For some of these, the 16S rRNA, these are identical gene sequences, of which 16S rRNA is 3 or more copies, and mutations exist in more copies or sequences. These figures are significantly below the 97% benchmark, which serves as an identity cutoff at the species level. This may imply that some isolates are either divergent strains of C. perfringens or closely related species. The low levels of similarity may also stem from mutations, poor sequencing quality, and the limitations of the database used. These situations require the use of specific primers or whole-genome sequencing. This exemplifies the lack of caution to be exercised with 16S rRNA sequencing, as it is often insufficient in providing conclusive evidence of a species. ConclusionThis study highlights the widespread presence of C. perfringens in human illness and local market meats, reinforcing the importance of this zoonotic and foodborne pathogen. The detection rates in both scenarios imply that unhygienic and undercooked, mis-stored meat may directly infect and facilitate the transmission of the disease to people. Isolates were abundant and contained the widest range of virulent genes. Isolates of cpe, pfoA, and hla genes had the greatest potential to cause disease because many of the isolates had multiple virulence factors and were, therefore, hypervirulent. The dual presence of multiple virulence and resistance to many antibiotics is even more disturbing. Of the phenotypes tested, resistance was noted for the three most common agents: erythromycin, tetracycline, and beta-lactams. These phenotypes were linked to respective ermB, tetA, and blaTEM gene copies in both human and meat isolates, reinforcing the notion of a common origin and lateral gene transfer. The close genetic relatedness between some meat-derived and clinical strains provides further evidence for foodborne transmission and emphasizes the need for monitoring frameworks, while the noted genetic diversity indicates adaptation to clinical and environmental settings. The lack of gene expression information stands as a critical limitation, as it makes associating genotype with functional pathogenicity almost impossible. Future research should employ sophisticated attempts along with expression analysis to address this gap. AcknowledgmentThe authors thank the College of Veterinary Medicine, University of Al-Qadisiyah, for their support in this study. Conflict of interestThe authors declare no conflicts of interest. FundingThe authors self-funded the study with no external funding. Authors’ contributionsAll authors participated in the study. Data availabilityData are available when requested by the corresponding author. ReferencesAbdel-Glil, M.Y., Thomas, P., Linde, J., Jolley, K.A., Harmsen, D., Wieler, L.H., Neubauer, H. and Seyboldt, C. 2021. Establishment of a Publicly Available Core Genome Multilocus Sequence Typing Scheme for Clostridium perfringens. Microbiol. Spectr. 9(2), e0053321; doi:10.1128/Spectrum.00533-21 Al-Deresawi, T.S., Mohammed, M.K. and Khudhair, S.H. 2022. The use of bacterial cellulose produced from the local isolate Komagataeibacter xylinus TELE as an antibacterial for wounds. Int. J. Drug Deliv. Technol. 12(4), 1825–1830. doi: 10.25258/ijddt.12.4.56 Ali, T., Sarwar, A. and Anjum, A.A. 2024. Alpha toxin production potential and antibiotic resistance patterns of Clostridium perfringens isolates from meat samples. Anim. Bioscience 37(11), 1970–1978. Aljindan, R., Aleraky, D.M., Farhat, M., Almandil, N.B., Abdulazeez, S. and Borgio, J.F. 2023. Genomic insights into virulence factors and multi-drug resistance in Clostridium perfringens IRMC2505A. Toxins 15(6), 359. Azevedo, I., Barbosa, J., Albano, H., Nogueira, T. and Teixeira, P. 2024. Lactic acid bacteria isolated from traditional and innovative alheiras as potential biocontrol agents. Food. Microbiol. 119, 104450. Catalán, A., Espoz, M.C., Cortés, W., Sagua, H., González, J. and Araya, J.E. 2010. Tetracycline and penicillin resistant Clostridium perfringens isolated from the fangs and venom glands of Loxosceles laeta: its implications in loxoscelism treatment. Toxicon 56(6), 890–896. Cattoir, V., Poirel, L., Rotimi, V., Soussy, C.J. and Nordmann, P. 2007. Multiplex PCR for detection of plasmid-mediated quinolone resistance qnr genes in ESBL-producing enterobacterial isolates. J. Antimicrob. Chemotherapy. 60(2), 394–397. Duc, H.M., Hoa, T.T.K., Ha, C.T.T., Van Hung, L., Van Thang, N., Minh Son, H. and Flory, G.A. 2024. Prevalence and antibiotic resistance profile of Clostridium perfringens isolated from pork and chicken meat in Vietnam. Pathogens 13(5), 400. European Committee on Antimicrobial Susceptibility Testing (EUCAST). 2024. Breakpoint tables for interpretation of MICs and zone diameters. Version 14.0, valid from 2024-01-01. Available via www.eucast.org Fang, M., Yuan, Y., Tian, X., Liu, Y., Wu, K., Zhu, Y., Zhang, L., Fox, E.M., Li, R., Bai, L., Li, M., Macori, G., Wang, X., Ye, D., Yang, Z., Zhang, R. and Wang, J. 2025. Genomic and antimicrobial resistance profiles of Clostridium perfringens isolated from pets in China. Vet. Microbiol. 304, 110490. Fourie, J.C.J., Bezuidenhout, C.C., Sanko, T.J., Mienie, C. and Adeleke, R. 2020. Inside environmental Clostridium perfringens genomes: antibiotic resistance genes, virulence factors and genomic features. J. Water Health 18(4), 477–493. Gambino, D., Vicari, D., Vitale, M., Schirò, G., Mira, F., Giglia, M.L., Riccardi, A., Gentile, A., Giardina, S., Carrozzo, A., Cumbo, V., Lastra, A. and Gargano, V. 2021. Study on bacteria isolates and antimicrobial resistance in wildlife in Sicily, Southern Italy. Microorganisms 9(1), 203. García-Vela, S., Martínez-Sancho, A., Said, L.B., Torres, C. and Fliss, I. 2023. Pathogenicity and antibiotic resistance diversity in Clostridium perfringens isolates from poultry affected by necrotic enteritis in Canada. Pathogens 12(7), 905. Ghazi, A.M., Ali Al-bayati, M.A. and Janabi, A.H. 2024. Metabolomics-detected alterations generated by phytosomal propolis and phytosomal lycopene in male rats with induced benign prostatic hyperplasia. Iraqi. J. Vet. Sci. 38(Suppl I–IV), 7–15. Habeeb, A.A. 2018. Detection of drug resistance gene expression in Candida albicans isolated from oral thrush of children via real-time PCR technique. J. Pharm. Sci. Res. 10(3), 594–596. Haider, Z., Ali, T., Ullah, A., Basit, A., Tahir, H., Tariq, H., Ilyas, S.Z., Hayat, Z. and Rehman, S.U. 2022. Isolation, toxinotyping and antimicrobial susceptibility testing of Clostridium perfringens isolated from Pakistan poultry. Anaerobe 73, 102499. Hassan, W.S., Abdulrazzaq, K.M., Al-Obaidi, Q.T. and Al-Azow, K.A. 2024. Molecular detection of Anaplasma platys in dogs in Nineveh province, Iraq. Iraqi J. Vet. Sci. 38(3), 677–682. Hassani, S., Pakbin, B., Brück, W.M., Mahmoudi, R. and Mousavi, S. 2022. Prevalence, antibiotic resistance, toxin-typing and genotyping of Clostridium perfringens in raw beef meats obtained from Qazvin City, Iran. Antibiotics 11(3), 340. Hyeong-Jun, L. and Gun-Jo, W. 2006. PCR-based detection and molecular genotyping of enterotoxigenic Clostridium perfringens isolates from swine diarrhea in Korea. J. Microbiol. Biotechnol. 16(2), 291–294. Karim, S., Mansour, K., Janabi, A. and Al-Nakeeb, N. 2018. First phylogenetic characterization of pseudocowpox virus from cattle in Al-Qadisiyah province, Iraq. Iraqi J. Vet. Sci. 33(1), 123–126. Khan, M.U.Z., Humza, M., Yang, S., Iqbal, M.Z., Xu, X. and Cai, J. 2021. Evaluation and optimization of antibiotics resistance profile against Clostridium perfringens from buffalo and cattle in Pakistan. Antibiotics 10(1), 59. Li, J., Zhou, Y., Yang, D., Zhang, S., Sun, Z., Wang, Y., Wang, S. and Wu, C. 2021. Prevalence and antimicrobial susceptibility of Clostridium perfringens in chickens and pigs from Beijing and Shanxi, China. Vet. Microbiol. 252, 108932; doi:10.1016/j.vetmic.2020.108932 Li, M., Wang, Y., Hou, B., Chen, Y., Hu, M., Zhao, X., Zhang, Q., Li, L., Luo, Y., Liu, Y. and Cai, Y. 2024. Toxin gene detection and antibiotic resistance of Clostridium perfringens from aquatic sources. Int. J. Food. Microbiol. 415, 110642. Ma, M., Li, J. and Mcclane, B.A. 2012. Genotypic and phenotypic characterization of Clostridium perfringens isolates from Darmbrand cases in post-World War II Germany. InfectImmunity 80(12), 4354–4363. Maty, H.N. 2023. Impact of sorbitol and L-carnitine on stimulating thyroid hormone, triiodothyronine, and adenosine triphosphate levels in broilers. Iraqi. J. Vet. Sci. 37(3), 589–590. Milton, A.A.P., Sanjukta, R., Gogoi, A.P., Momin, K.M., Priya, G.B., Das, S., Ghatak, S., Sen, A. and Kandpal, B.K. 2020. Prevalence, molecular typing and antibiotic resistance of Clostridium perfringens in free range ducks in Northeast India. Anaerobe 64, 102242. Mohiuddin, M., Song, Z., Liao, S., Qi, N., Li, J., Lv, M., Lin, X., Cai, H., Hu, J., Liu, S., Zhang, J., Gu, Y. and Sun, M. 2023. Animal model studies, antibiotic resistance and toxin gene profile of NE reproducing Clostridium perfringens type A and type G strains isolated from commercial poultry farms in China. Microorganisms 11(3), 622. Ozma, M.A., Fadaee, M., Hosseini, H.M., Ataee, M.H. and Mirhosseini, S.A. 2025. A critical review of postbiotics as promising novel therapeutic agents for clostridial infections. Probiotics. Antimicrob. Proteins. 17(2), 656–667. Qui, N.H. 2023. Baker’s yeast (Saccharomyces cerevisiae) and its application on poultry’s production and health: a review. Iraqi J. Vet. Sci. 37(1), 213–221. Rahawi, A.M., Al-Taee, S.K., Ali, F.F., Altaey, O.Y. and Abdullah, D.A. 2024. Protective role of biosynthetic silver nanoparticles in broilers with aflatoxicosis through histopathological study of spleen. Iraqi J. Vet. Sci. 38(3), 565–572. Rajput, D.S., Zeng, D., Khalique, A., Rajput, S.S., Wang, H., Zhao, Y., Sun, N. and Ni, X. 2020. Pretreatment with probiotics ameliorate gut health and necrotic enteritis in broiler chickens, a substitute to antibiotics. AMB. Express 10(1), 220. Rana, E.A., Nizami, T.A., Islam, M.S., Barua, H. and Islam, M.Z. 2023. Phenotypical Identification and Toxinotyping of Clostridium perfringens Isolates from Healthy and Enteric Disease-Affected Chickens. Vet. Med. Int. 2023, 2584171; doi:10.1155/2023/2584171 Revitt-Mills, S.A., Watts, T.D., Lyras, D., Adams, V. and Rood, J.I. 2021. The ever-expanding tcp conjugation locus of pCW3 from Clostridium perfringens. Plasmid 113, 102516. doi: 10.1016/j.plasmid.2020.102516 Sárvári, K.P. and Schoblocher, D. 2020. The antibiotic susceptibility pattern of gas gangrene-forming Clostridium spp. clinical isolates from South-Eastern Hungary. Infect. Dis. 52(3), 196–201. Song, X., Zhong, Z., Bai, J., Pu, T., Wang, X., He, H., Chen, Y., Yang, C. and Zhang, Q. 2024. Emergence of genetic diversity and multi-drug resistant Clostridium perfringens from wild birds. BMC. Vet. Res. 20(1), 300. Sun, Z., Lu, M., Lillehoj, H., Lee, Y., Goo, D., Yuan, B., Yan, X. and Li, C. 2023. Characterization of collagen binding activity of Clostridium perfringens strains isolated from broiler chickens. Pathogens 12(6), 778. Takeuchi, S., Hashizume, N., Kinoshita, T., Kaidoh, T. and Tamura, Y. 1997. Detection of Clostridium septicum hemolysin gene by polymerase chain reaction. J. Vet. Med. Sci. 59(9), 853–855. Thanki, A.M., Osei, E.K., Whenham, N., Salter, M.G., Bedford, M.R., Masey O’neill, H.V. and Clokie, M.R.J. 2024. Broad host range phages target global Clostridium perfringens bacterial strains and clear infection in five-strain model systems. Microbiol. Spectr. 12(5), 378423; e0378423 The European Committee on Antimicrobial Susceptibility Testing. (2024). Breakpoint tables for interpretation of MICs and zone diameters, version 14.0. EUCAST. https://www.eucast.org Valdez, G., Shyur, L.F., Wang, S.Y. and Chen, S.E. 2023. Phytogenics in ginger, Origanum vulgare, and Syzygium aromaticum and their potential as a feed additive against Clostridium perfringens in broiler production. Animals 13(23), 3643. Venhorst, J., Van Der Vossen, J.M.B.M. and Agamennone, V. 2022. Battling enteropathogenic clostridia: phage therapy for Clostridioides difficile and Clostridium perfringens. Front. Microbiol. 13, 891790. Walsh, F., Ingenfeld, A., Zampicolli, M., Hilber-Bodmer, M., Frey, J.E. and Duffy, B. 2011. Real-time PCR methods for quantitative monitoring of streptomycin and tetracycline resistance genes in agricultural ecosystems. J. Microbiological Methods 86(2), 150–155. Wang, H., Wang, L., Zhang, F., Li, X., Wang, S., Gao, D., Li, X. and Qian, P. 2024. ParalichenysinDY4, a novel bacteriocin-like substance, is employed to control Clostridium perfringens. Int. J. Biol. Macromolecules 279(Pt 4), 135412. Wang, X.Y., Meng, F.H., Zhang, M.Y., Li, F.X., Lei, Y.X., Ma, Z.G., Li, J.Q., Lou, Y.N., Chu, Y.F., Ma, K. and Yu, S.X. 2024. Gut Lactococcus garvieae promotes protective immunity to foodborne Clostridium perfringens infection. Microbiol. Spectr. 12(10), 402523. Watts, T.D., Traore, D.A.K., Atkinson, S.C., Lao, C., Caltabiano, N., Rood, J.I. and Adams, V. 2022. The specificity of ParR binding determines the incompatibility of conjugative plasmids in Clostridium perfringens. mBio 13(4), 135622. Wu, K., Li, Z., Fang, M., Yuan, Y., Fox, E.M., Liu, Y., Li, R., Bai, L., Zhang, W., Zhang, W.M., Yang, Q., Chang, L., Li, P., Wang, X., Wang, J. and Yang, Z. 2023. Genome characteristics of the optrA-positive Clostridium perfringens strain QHY-2 carrying a novel plasmid type. mSystems 8(4), 53523. Xiaoting, W., Chengcheng, N., Chunhui, J., Yan, L., Jing, L., Qingling, M., Jun, Q., Lixia, W., Kuojun, C., Jinsheng, Z., Zaichao, Z., Weiwei, Y., Yelong, P. and Xuepeng, C. 2021. Antimicrobial resistance profiling and molecular typing of ruminant-borne isolates of Clostridium perfringens from Xinjiang, China. J. Global Antimicrob. Resist. 27, 41–45. Xiu, L., Zhu, C., Zhong, Z., Liu, L., Chen, S., Xu, W. and Wang, H. 2022. Prevalence and multilocus sequence typing of Clostridium perfringens isolated from different stages of a duck production chain. Food. Microbiol. 102, 103901. Xu, W., Wang, H., Chen, S., Chen, Y., Liu, L. and Wu, W. 2021b. Tracing Clostridium perfringens strains along the chicken production chain from farm to slaughter by multilocus sequence typing. Zoonoses Public Health 68(5), 431–442. Xu, W., Wang, H., Liu, L., Miao, Z., Huo, Y. and Zhong, Z. 2021a. Prevalence and characterization of Clostridium perfringens isolated from different chicken farms in China. Anaerobe 72, 102467. Yaseen, M.M., Karawan, A.C., Alfatlawi, M.A.A. and Janabi, A.H.D. 2020. The role of gut bacterial cytochrome-P450 of mosquito larvae in degradation of temephos insecticide. Ann. Trop. Med. Public. Health. 23(1), S412. | ||
| How to Cite this Article |
| Pubmed Style Fahad KH, Sadeq JN, Neamah AA. Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Vet. J.. 2025; 15(12): 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 Web Style Fahad KH, Sadeq JN, Neamah AA. Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. https://www.openveterinaryjournal.com/?mno=258075 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i12.43 AMA (American Medical Association) Style Fahad KH, Sadeq JN, Neamah AA. Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Vet. J.. 2025; 15(12): 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 Vancouver/ICMJE Style Fahad KH, Sadeq JN, Neamah AA. Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Vet. J.. (2025), [cited January 25, 2026]; 15(12): 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 Harvard Style Fahad, K. H., Sadeq, . J. N. & Neamah, . A. A. (2025) Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Vet. J., 15 (12), 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 Turabian Style Fahad, Khilood Hamdan, Jenan Nadhim Sadeq, and Azhar Abdulsada Neamah. 2025. Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Veterinary Journal, 15 (12), 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 Chicago Style Fahad, Khilood Hamdan, Jenan Nadhim Sadeq, and Azhar Abdulsada Neamah. "Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples." Open Veterinary Journal 15 (2025), 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 MLA (The Modern Language Association) Style Fahad, Khilood Hamdan, Jenan Nadhim Sadeq, and Azhar Abdulsada Neamah. "Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples." Open Veterinary Journal 15.12 (2025), 6602-6621. Print. doi:10.5455/OVJ.2025.v15.i12.43 APA (American Psychological Association) Style Fahad, K. H., Sadeq, . J. N. & Neamah, . A. A. (2025) Molecular tracking of antibiotic-resistant and virulent Clostridium perfringens strains in Iraqi human and meat samples. Open Veterinary Journal, 15 (12), 6602-6621. doi:10.5455/OVJ.2025.v15.i12.43 |