| Research Article | ||
Open Vet. J.. 2025; 15(10): 5175-5182 Open Veterinary Journal, (2025), Vol. 15(10): 5175-5182 Research Article The molecular evidence of toxic shock syndrome toxin-1 on methicillin-resistant Staphylococcus aureus isolated from cats in Surabaya, IndonesiaDaniah Ashri Afnani1,2, Aswin Rafif Khairullah3, Mustofa Helmi Effendi4,5*, Wiwiek Tyasningsih6, John Yew Huat Tang5, Budiastuti Budiastuti7, Saifur Rehman8, Ikechukwu Benjamin Moses9, Dea Anita Ariani Kurniasih10, Sheila Marty Yanestria11, Riza Zainuddin Ahmad2, Siti Hamidatul Aliyah12, Alfiana Laili Dwi Agustin13 and Katty Hendriana Priscilia Riwu131Doctoral Program of Veterinary Science, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 2Department of Microbiology and Parasitology, Faculty of Veterinary Medicine, Universitas Pendidikan Mandalika, Mataram, Indonesia 3Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor, Indonesia 4Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 5School of Food Industry, Faculty of Bioresources, and Food Industry, Universiti Sultan Zainal Abidin, Besut, Malaysia 6Division of Veterinary Microbiology, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 7Study Program of Pharmacy Science, Faculty of Health Science, Universitas Muhammadiyah Surabaya, Surabaya, Indonesia 8Department of Pathobiology, Faculty of Veterinary and Animal Sciences, Gomal University, Dera Ismail Khan, Pakistan 9Department of Applied Microbiology, Faculty of Science, Ebonyi State University, Abakaliki, Nigeria 10Research Center for Public Health and Nutrition, National Research and Innovation Agency (BRIN), Bogor, Indonesia 11Department of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Wijaya Kusuma Surabaya, Surabaya, Indonesia 12Center for Biomedical Research, National Research and Innovation Agency (BRIN), Bogor, Indonesia 13Department of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Pendidikan Mandalika, Mataram, Indonesia *Corresponding Author: Mustofa Helmi Effendi. Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia. Email: mhelmieffendi [at] gmail.com Submitted: 10/04/2025 Revised: 14/09/2025 Accepted: 21/09/2025 Published: 31/10/2025 © 2025 Open Veterinary Journal
AbstractBackground: Staphylococcus aureus is a significant microorganism in both human and veterinary medicine due to its ability to colonize mucosal surfaces, particularly the nasal cavity, and its potential to cause opportunistic infections in domestic cats. Methicillin-resistant S. aureus (MRSA) is a new strain of S. aureus with multidrug-resistance (MDR) characteristics and is resistant to β-lactam antibiotics. Companion animals are classified as potential reservoirs of MRSA. In this study, 150 cats were brought to veterinary clinics and animal hospitals in Surabaya for routine check-ups or medical treatment. Cats were not selected based on the presence of clinical symptoms, allowing for the identification of both symptomatic and asymptomatic carriers. Swab samples were collected from the nasal cavities using sterile cotton swabs under aseptic conditions. This approach enabled the assessment of MRSA colonization. Aim: This study aimed to detect the tst gene that produces toxic shock syndrome toxin-1 (TSST-1) from MRSA isolates in cats. Methods: A total of 150 nose swab samples were collected from cats visiting veterinary clinics and hospitals as pet patients spread over Surabaya, Indonesia. Amies Medium Transport was used to swab the noses of the samples, and the microbiological standard procedure was used for identification. Staphylococcus aureus isolates were subjected to an antibiotic resistance test using the Kirby-Bauer diffusion method by streaking a bacterial suspension according to a 0.5 McFarland standard, then five different antibiotic discs on Mueller-Hinton Agar. Cefoxitin-resistant S. aureus isolates grew on Oxacillin Resistance Screening Agar Base (ORSAB) as a confirmatory test for MRSA. Results: There were 18 (12%) S. aureus isolates identified and isolated. The antibiotic resistance test results revealed 7 (38.88%) MDR isolates; there were 3 (16.66%) MDR of S. aureus isolates and 4 (22.22%) MDR of S. aureus isolates that were ORSAB positive; there were MDR and MRSA isolates. 4 MRSA isolates (22.22%) were subjected to molecular detection. Conclusion: Polymerase chain reaction 1 (5.55%) of the MRSA isolate was test gene positive at 143 bp. This indicates that cats may be a potential source of transmission to humans and endanger public health. Keywords: Antibiotic, Cats, MRSA, Public health, TSST-1. IntroductionThe popularity of cats as pets is rapidly rising, particularly in metropolitan areas, and domestication and globalization have increased the risk of bacterial transmission between humans and animals (Afnani et al., 2022). The skin and mucous membrane flora of mammals and birds naturally contain Staphylococcus aureus, a common opportunistic infection in humans and veterinary medicine (Khairullah et al., 2019; Khairullah et al., 2024a). Resistance by mutation is the term used to describe how S. aureus develops drug resistance as a result of genetic changes that change the target DNA gyrase or decrease outer membrane proteins (Ramandinianto et al., 2020a; Pinchuk et al., 2010). Bacteria can become drug-resistant by producing high levels of α-lactamase through plasmid-mediated transduction, transformation, and insertion of drug-resistance genes. This is a type of plasmid-mediated resistance (Ramandinianto et al., 2020b). The primary source of methicillin-resistant S. aureus (MRSA) is drug-resistant gene transmission mediated by plasmids, which can expand the genome and enable the transfer of resistance genes between S. aureus and other bacteria (Waruwu et al., 2023). Human, animal, and environmental antimicrobial resistance (AMR) is increased via intricate interactions between bacterial species from various "environments" (Yunita et al., 2020; Kumar et al., 2016). MRSA is a methicillin-resistant strain of the bacteria that is currently posing a threat to the entire world (Putra et al., 2023). It has spread to the public among people without risk factors for contracting MRSA, posing a fresh danger after becoming well-established in the health care setting (Kaben et al., 2024). The subsequent finding that MRSA can colonize or infect animals and food derived from animals was highly alarming, and more MRSA reservoirs were identified (Tyasningsih et al., 2019). Close contact between pets and their owners fosters an environment that is conducive to MRSA transmission, and pet owners are more likely to become colonized by MRSA than the general population, suggesting that dogs also operate as reservoirs (Ramadhani et al., 2024). However, several studies found that the majority of MRSA in pets comes from humans, and that close contact with people is associated with a higher risk of S. aureus colonization in pets (Khairullah et al., 2023). High population density, the potential for nosocomial transmission from humans, poor cleaning and disinfection practices, the unknown carrier status of multiple animals, or a stressful environment are some potential reasons for this exposure (Khairullah et al., 2024b). Staphylococcal infections from animals can be transmitted to humans, and the most common staphylococcal strain in cats is S. aureus because these germs can occasionally flourish on household animals as opportunistic pathogens (Pantosti, 2012; Haag et al., 2019). The virulence factors that drive the host's innate and adaptive immune responses are the main pathophysiology of S. aureus as a pathogen (Cheung et al., 2021; Fazal et al., 2023). The test gene encodes the toxin known as toxic shock syndrome toxin 1 (TSST-1), which is one of the most significant virulence factors of S. aureus (Zheng et al., 2020). A diverse range of virulence factors determines the pathogenicity of S. aureus, with secreted toxins playing a major role. Numerous toxins produced by S. aureus harm biological membranes, causing cell death (Zhang et al., 2017; Rehab et al., 2016). The most well-known S. aureus superantigen, the 22-kD TSST-1, causes toxic shock syndrome (TSS), a disease that causes the release of cytokines such as TNF-, IL-1, and IL-2 (Ortega et al., 2010). Although TSST does not cause emesis, unlike enterotoxin superantigens, TSS is a dangerous and occasionally fatal condition (Otto, 2014; Chen et al., 2023). MRSA strains have been linked to no instances in Europe, a considerable percentage of TSS cases in Japan, and occasional cases in the United States (Jamart et al., 2005). Historically, TSS was typically linked to menstruation and high-absorbency tampons; however, TSS is more commonly caused by other staphylococcal infections, especially those affecting the skin and soft tissues (Sharma et al., 2019). Staphylococcus aureus toxin-induced fever, rash, desquamation, hypotension, and multiple organ failure are the hallmarks of TSS in both clinical and laboratory settings (Atchade et al., 2024). Because TSST-1-carrying MRSA is difficult to treat, it poses a risk to public health. This study aimed to detect the tst gene that produces TSST-1 in MRSA isolates from cats. Materials and MethodsStudy design and sample collectionThis research was conducted using a cross-sectional study design, which aims to analyze the correlational relationship between potential causes and their associated outcomes. A random sampling technique was employed, in which nasal swabs were collected using Amies transport medium. A total of 150 nasal swabs from both healthy and diseased cats were taken from five regions (North, East, West, Central, and South) of Surabaya, East Java, Indonesia, between May 2022 and July 2023. The required sample size was calculated using the disease prevalence formula described by Martin et al. (1987), expressed as N=4PQ/L². Where N represents the sample size, P is the assumed prevalence, q equals 100% minus P, and L denotes the desired margin of error. Samples were obtained from several laboratories, namely, the Department of Veterinary Microbiology and Mycology, Faculty of Veterinary, Airlangga University, and the Central for Health Laboratory Surabaya. Bacterial isolatesAmies was used as a medium to collect cat nasal swab samples, which were then kept in an icebox at 4°C. Staphylococcus aureus was isolated using a sterilized Amies cotton swab and streaked on MSA. The bacterial inoculum was cultured on MSA medium at 37°C for 24 hours (Effendi et al., 2019). Gram staining, positive catalase, coagulase, and Voges-Proskauer test results, as well as yellow colonies with yellow zones on MSA media, were used to identify S. aureus (Effendi et al., 2018). Antibiotic sensitivity and MRSA confirmation testsAntibiotic sensitivity tests were performed using five different antibiotics disk using Kirby-Bauer diffusion method, there were cefoxitin 5 µg, erythromycin 15 µg, tetracycline 30 µg, ciprofloxacin 5 µg, and chloramphenicol 30 µg. A McFarland equivalent of 0.5 was used to prepare purified cultures in bacterial suspension. The Mueller Hinton Agar (MHA) (Oxoid-CM00337) was streaked with a sterile cotton swab to create the inoculum. Five distinct antibiotic disks were then added, and the mixture was incubated at 35°C for 24 hours. The width of the inhibition zone of the clear region surrounding the antibiotic disk, as specified by the (Clinical and Laboratory Standards Institute, 2020), was measured in millimeters (mm) to determine the results of antibiotic sensitivity tests. The MRSA confirmation test was performed using several cefoxitin-resistant S. aureus isolates from MHA (Oxoid-CM00337), and the outcomes were then streaked on Oxacillin Resistance Screening Agar Base (ORSAB) (HiMedia M1415) and Oxacillin Resistance Selective Supplement (Supplement, HiMedia FD191) (Decline et al., 2020). Detection of the tst geneDNA was extracted from the MRSA isolate using a QIAamp DNA Mini Kit 50 (Qiagen, Germany) according to the manufacturer’s instructions. MRSA isolates were added with 180 µl lysozyme (20 mg/ml) and incubated for 45 minutes at 37°C. Then, the extracted DNA was diluted to 20 µl Proteinase K and 200 µl Buffer kit. 3 µl of DNA solution was used for polymerase chain reaction (PCR) amplification. Pure S. aureus genomic DNA was used as a template for amplification using a Promega GoTag Master Mix in accordance with the manufacturer’s guidelines. PCR amplification was performed using test (F) and test (R) as described in Table 1. The amplification process included an initial denaturation at 95°C for 4 minutes of 1 cycle, denaturation at 95°C for 30 seconds, followed by annealing at 54°C for 30 seconds in 40 cycles, then an extension at 72°C for 1 minutes, and a final extension at 72°C for 1 minute. Following amplification reactions, the PCR products were electrophoretically separated on a 2% agarose gel using Ultrapore Agarose Gel (Invitrogen, USA) and stained with Safe DNA gel stain (Invitrogen, USA) at 100 V for 45 minutes in 10X UltrapureTM TBE buffer (Invitrogen, USA) before being illuminated by a UV transilluminator and visualized. Table 1. The primers used in this study.
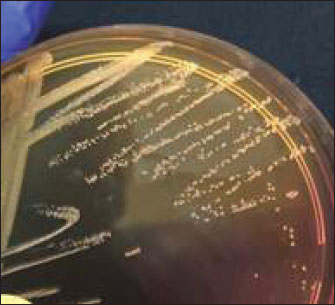
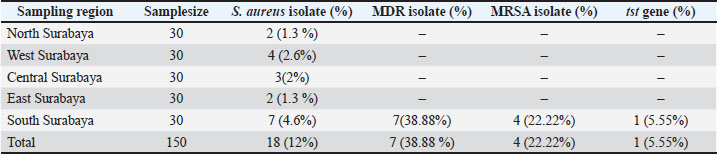
Ethical approvalThis research received ethical approval from the Animal Ethics Committee, Faculty of Veterinary Medicine, Wijaya Kusuma University. The approval was issued through the Animal Care and Use Committee (ACUC) with certificate number 217-KKE. ResultsIsolation and identification of S. aureus strainsA total of 150 nasal swabs from cat samples were analyzed, and 18 (12%) were positive for S. aureus based on morphological culture by the shape, color, and edges of the bacterial colonies (Fig. 1). Gram staining was used as the identification method, and biochemical tests including catalase, coagulase, and Vogues Proskauer were used.
Fig. 1. Staphylococcus aureus colonies in the MSA. Antibiotic sensitivity and MRSA confirmation testsFrom 150 samples, 18 (12%) S. aureus isolates were identified: one resistant to tetracycline, two to tetracycline and erythromycin, and seven classified as multidrug-resistant (MDR). The main MDR patterns were cefoxitin–ciprofloxacin–tetracycline–erythromycin–chloramphenicol (four isolates) and erythromycin–tetracycline–chloramphenicol (three isolates). Seven isolates showed no resistance (Tables 2 and 3). Table 2. Prevalence results from a regional MRSA screening and the presence of the tst gene in MRSA isolates.
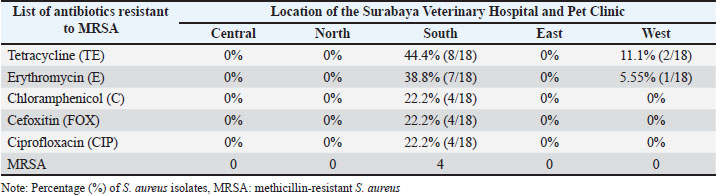
Table 3. Ratio of antibiotic resistance test and region-based MRSA screening from cat nasal swabs (%).
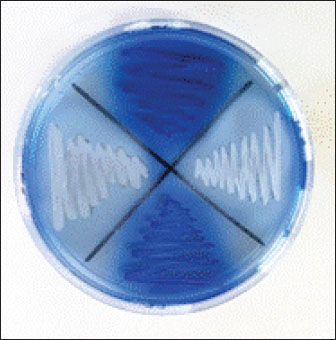
Four S. aureus isolates that were resistant to cefoxitin were classified as presumptive MRSA isolates and then continued to streaked on ORSAB that contained oxacillin resistance selective supplement as a confirmation test of MRSA. The results of the ORSA test showed that the blue color indicated positive and the white color indicated negative confirmation results, as shown in Figure 2. Four presumptive MRSA isolates that tested with ORSA showed all positive results; it can be concluded that they were phenotypically confirmed as MRSA isolates.
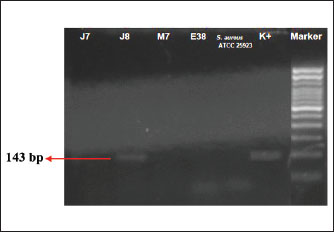
Fig. 2. ORSAB as a confirmation test for MRSA. The blue color indicates positive confirmation results, and the white color indicates negative confirmation results. Detection of the tst gene in MRSA isolatesFour MRSA isolates were further examined molecularly using conventional PCR, and one of four MRSA isolates harbored the tst gene (Fig. 3).
Fig. 3. Gel electrophoresis demonstrating the presence of gen test at a nucleotide length of 143 bp in one of the four MRSA isolates from nasal swabs in cats. Note: K- stands for S. aureus ATCC 25923 as the negative control, K+ stands for positive control, and J7, M7, and E38 are MRSA isolates. DiscussionBased on the prevalence results of multidrug resistant S. aureus isolates were found seven (38.88%) in South Surabaya, four of them were MRSA isolates confirmed with ORSAB phenotypically and there was found one MRSA isolate harbored the test gene. Among the many virulence traits of MRSA are toxins called TSST-1, which are encoded by the test gene (Zarei Koosha et al., 2016). These results are consistent with those of prior research conducted in multiple countries, which similarly identified MRSA isolates harboring the test gene. The detection of the test gene in our MRSA isolate is consistent with the findings of several international studies. For example, Zhao et al. (2019) reported that 198 of 208 MRSA isolates (29.88%) carried the test gene. Similarly, Motamedifar et al. (2015) found that 11.66% of MRSA isolates in Iran were positive for tst. In contrast, a study conducted in Japan reported a significantly higher prevalence, with 75% of MRSA isolates carrying the test gene (Nagao et al., 2009). These variations suggest geographical and epidemiological differences in the distribution of tst-positive MRSA (Fisher et al., 2018). Furthermore, Al Laham et al. (2015) reported a high prevalence of MRSA isolates carrying the test gene in the Gaza region, especially within the CC22 clone ("Gaza strain"), indicating the potential for widespread carriage in community settings. In contrast, our study showed a relatively low prevalence (one isolate) of the test gene among MRSA strains, which may be attributed to differences in sample size, population, or local practices. Numerous studies have also reported the detection of the test gene in MRSA from animal sources. For instance, Farahmand-Azar et al. (2013) detected test in 19.2% of bovine S. aureus isolates, whereas Bruce et al. (2022) identified it in 2 of 7 MRSA isolates from pets (dogs and cats). These findings, along with the report by Crespo-Piazuelo and Lawlor (2021), support the idea that close human–animal contact can facilitate the zoonotic transmission of MRSA strains harboring virulence genes such as test. The widespread resistance of S. aureus strains to multiple antibiotics, particularly in hospital settings, remains a major global health concern. Due to its resistance to methicillin and other commonly used antibiotics, MRSA is categorized as a superbug Khairullah et al., 2022). The high rates of AMR observed in our isolates may reflect the overuse or misuse of antibiotics, which has been reported to reduce the effectiveness of treatments and enhance the survival of resistant strains (Mustapha et al., 2014). Moreover, AMR may influence the expression of virulence factors. The co-occurrence of multidrug resistance and test gene expression, although low in our study, may have serious clinical implications, as TSST-1 is associated with higher mortality rates (Zarei Koosha et al., 2016). TSST-1 acts as a superantigen, leading to excessive T-cell activation and cytokine release, which may result in TSS characterized by rapid fever onset, organ failure, and potentially death (Spaulding et al., 2013; Juwita et al., 2021). Interestingly, while a high percentage of tst-positive isolates have been reported in some studies, the clinical incidence of TSS remains low. Environmental and host factors significantly influence the expression of tst (Deurenberg et al., 2005; Mrochen et al., 2020). TSST-1 production is affected by temperature, pH, oxygen levels, magnesium concentration, and bacterial density (Yarwood and Schlievert, 2000; Khojasteh et al., 2016; Kulhankova et al., 2018). The optimal conditions for TSST-1 expression include a temperature range of 37°C–40°C, pH 6.5–7, and aerobic conditions (Zschöck et al., 2000). Historically, TSS was often associated with tampon use in menstruating women, but recent studies have linked it to nonmenstrual cases, particularly skin and soft tissue infections (Schlievert and Davis, 2020). The potential of tampons to absorb magnesium and create an environment conducive to test gene expression highlights the importance of understanding the host-environment interactions that modulate virulence factor production (Tong et al., 2015). Lastly, the detection of tst-positive MRSA in both humans and animals underscores the importance of the One Health approach to infection control (Nandhini et al., 2022). Close contact with domestic animals, such as cats and dogs, may contribute to the transmission of these strains to humans (Crespo-Piazuelo and Lawlor, 2021; Krakauer and Stiles, 2013). Therefore, proper sanitation and hygiene practices, including routine disinfection, pet hygiene, waste management, and limiting unnecessary contact between animals, are crucial for preventing the spread of MRSA (Haag et al., 2019). ConclusionIn conclusion, this study demonstrated that MRSA is a serious worldwide problem; its occurrences are not limited to human health alone, as it can spread from humans to animals in numerous ways. Among the virulence factors carried by MRSA are toxins known as TSST-1. At trace doses, TSST-1 is hazardous to humans, in contrast to most other secreted toxins produced by S. aureus. TSST-1 is impervious to proteolysis, heat, and acid as a superantigen. The TSST-1 superantigen is essential for controlling the host and preserving a colonizing environment. The frequency of TSST-1 from MRSA isolates can be used as a guide to prevent and control MRSA infections and raise public awareness, particularly among high-risk groups including pet owners, paramedics, and veterinarians. More investigation is required to identify the additional MRSA virulence factors than TSST-1. AcknowledgmentsThe authors would like to thank the Faculty of Veterinary Medicine, Universitas Airlangga. Conflict of interestThe authors declare no conflict of interest. FundingThis study was partly supported by The International Research Consortium, Lembaga Penelitian dan Pengabdian Masyarakat, Universitas Airlangga, Surabaya, Indonesia, in 2024 (Grant number: 171/UN3.LPPM/PT.01.03/2024). Author’s contributionsDAA, WT, and MHE: Conceived, designed, and coordinated the study. JYHT, SMY, and BB: Designed data collection tools, supervised the field sample and data collection, and performed laboratory work and data entry. ALDA, IBM, and KHPR: Validation, supervision, and formal analysis. RZA and DAAK: Contributed reagents, materials, and analysis tools. SHA, SR, and ARK: Carried out the statistical analysis and interpretation and participated in the preparation of the manuscript. All authors have read, reviewed, and approved the final version of the manuscript. Data availabilityAll data are available in the revised manuscript. ReferencesAfnani, D.A., Fatih, N., Effendi, M.H., Tyasningsih, W., Khairullah, A.R., Kurniawan, S.C., Silaen, O.S.M., Ramandinianto, S.C., Widodo, A. and Riwu, K.H.P. 2022. Profile of multidrug resistance and methicillin-resistant Staphylococcus aureus (MRSA) isolated from cats in Surabaya, Indonesia. Biodiversitas 23(11), 5703–5709. Atchade, E., De Tymowski, C., Grall, N., Tanaka, S. and Montravers, P. 2024. Toxic shock syndrome: a literature review. Antibiotics 13(1), 96. Bruce, S.A., Smith, J.T., Mydosh, J.L., Ball, J., Needle, D.B., Gibson, R. and Andam, C.P. 2022. Shared antibiotic resistance and virulence genes in Staphylococcus aureus from diverse animal hosts. Sci. Rep. 12(1), 4413. Chen, Q., Zhao, G., Yang, W., Chen, F., Qi, Y. and Lou, Z. 2023. Investigation into the prevalence of enterotoxin genes and genetic background of Staphylococcus aureus isolates from retain foods in Hangzhou, China. BMC Microbiol. 23(1), 294. Cheung, G.Y.C., Bae, J.S. and Otto, M. 2021. Pathogenicity and virulence of Staphylococcus aureus. Virulence 12(1), 547–569. Clinical and Laboratory Standards Institute. 2020. Performance standards for antimicrobial disk diffusion susceptibility testing; 17th Informational Supplement. Approved Standard M100-17. Clinical and Laboratory Standards Institute, Wyne, PA. Crespo-Piazuelo, D. and Lawlor, P.G. 2021. Livestock-associated methicillin-resistant Staphylococcus aureus (LA-MRSA) prevalence in humans in close contact with animals and measures to reduce on-farm colonisation. Ir. Vet. J. 74(1), 21. Decline, V., Effendi, M.H., Rahmaniar, R.P., Yanestria, S.M. and Harijani, N. 2020. Profile of antibiotic-resistant and presence of methicillin-resistant Staphylococcus aureus from nasal swab of dogs from several animal clinics in Surabaya, Indonesia. Int. J. One Health 6(1), 90–94. Deurenberg, R.H., Nieuwenhuis, R.F., Driessen, C., London, N., Stassen, F.R., Tiel, F.H., Stobberingh, E.E. and Vink, C. 2005. The prevalence of the Staphylococcus aureus tst gene among community- and hospital-acquired strains and isolates from Wegener's Granulomatosis patients. FEMS. Microbiol. Lett. 245(1), 185–189. Effendi, M.H., Hisyam, M.A.M., Hastutiek, P. and Tyasningsih, W. 2019. Detection of coagulase gene in Staphylococcus aureus from several dairy farms in East Java, Indonesia, by polymerase chain reaction. Vet. World 12(1), 68–71. Effendi, M.H., Oktavianto, A. and Hastutiek, P. 2018. Tetracycline resistance gene in Streptococcus agalactiae isolated from bovine subclinical mastitis in Surabaya, Indonesia. Philipp. J. Vet. Med. 55(SI), 115–120. Farahmand-Azar, S., Ahmadi, M., Saei, H.D. and Anassori, E. 2013. Identification of toxic shock syndrome toxin-1 (TSST-1) gene in Staphylococcus aureus isolated from bovine mastitis milk. Arch. Razi Inst. 68(1), 17–22. Fazal, M.A., Rana, E.A., Akter, S., Alim, M.A., Barua, H. and Ahad, A. 2023. Molecular identification, antimicrobial resistance and virulence gene profiling of Staphylococcus spp. associated with bovine sub-clinical mastitis in Bangladesh. Vet. Anim. Sci. 21(1), 100297. Fisher, E.L., Otto, M. and Cheung, G.Y.C. 2018. Basis of Virulence in Enterotoxin-Mediated Staphylococcal Food Poisoning. Front. Microbiol. 9(1), 436. Haag, A.F., Fitzgerald, J.R. and Penadés, J.R. 2019. Staphylococcus aureus in Animals. Microbiol. Spectr. 7(3), 11. Jamart, S., Denis, O., Deplano, A., Tragas, G., Vandergheynst, A., De Bels, D. and Devriendt, J. 2005. Methicillin-resistant Staphylococcus aureus toxic shock syndrome. Emerg. Infect. Dis. 11(4), 636–637. Juwita, S., Indrawati, A., Damajanti, R. and Safika Mayasari. 2021. Multiple antibiotic resistance and virulence factors of Staphylococcus aureus strains isolated from dairy farms in South Sulawesi, Indonesia. Biodiversitas 23(2), 1015–1022. Kaben, S.E., Al-Arif, M.A., Ma’Ruf, A., Effendi, M.H., Kurniawan, S.C., Riwu, K.H.P., Afnani, D.A., Silaen, O.S.M., Millannia, S.K., Ramadhani, S., Widodo, A., Farizqi, T.I. and Khairullah, A.R. 2024. Detection mecA gene and Staphylococcus aureus resistance to several antibiotics isolated from cat ear swabs at a veterinary hospital located at Surabaya – Indonesia. Braz. J. Vet. Res. Anim. Sci. 61(1), e209027. Khairullah, A., Sudjarwo, S., Effendi, M., Ramandinianto, S., Gelolodo, M., Widodo, A., Riwu, K. and Kurniawati, D. 2023. Pet animals as reservoirs for spreading methicillin-resistant Staphylococcus aureus to human health. J. Adv. Vet. Anim. Res. 10(1), 1–13. Khairullah, A.R., Kurniawan, S.C., Sudjarwo, S.A., Effendi, M.H., Widodo, A., Moses, I.B., Hasib, A., Zahra, R.L.A., Gelolodo, M.A., Kurniawati, D.A., Riwu, K.H.P., Silaen, O.S.M., Afnani, D.A. and Ramandinianto, S.C. 2024b. Kinship analysis of mecA gene of methicillin-resistant Staphylococcus aureus isolated from milk and risk factors from the farmers in Blitar, Indonesia. Vet. World 17(1), 216–225. Khairullah, A.R., Raharjo, D., Rahmahani, J., Suwarno., Tyasningsih, W. and Harijani, N. 2019. Antibiotics resistant at Staphylococcus aureus and Streptococcus sp isolated from bovine mastitis in Karangploso, East Java, Indonesia. Indian J. Forensic Med. Toxicol. 13(4), 439–444. Khairullah, A.R., Sudjarwo, S.A., Effendi, M.H., Ramandinianto, S.C., Widodo, A. and Riwu, K.H.P. 2022. A review of horses as a source of spreading livestock-associated methicillin-resistant Staphylococcus aureus to human health. Vet. World 15(8), 1906–1915. Khairullah, A.R., Widodo, A., Riwu, K.H.P., Yanestria, S.M., Moses, I.B., Effendi, M.H., Fauzia, K.A., Fauziah, I., Hasib, A., Kusala, M.K.J., Raissa, R., Silaen, O.S.M., Ramandinianto, S.C. and Afnani, D.A. 2024a. Spread of livestock-associated methicillin-resistant Staphylococcus aureus in poultry and its risks to public health: a comprehensive review. Open Vet. J. 14(9), 2116–2128. Khojasteh, V.J., Alfakhri, S. and Foster, H.A. 2016. Effects of silver sulphadiazine on production of extracellular proteins by strains of Staphylococcus aureus isolated from burns wound. Iran. J. Pharm. Res. 15(2), 653–660. Krakauer, T. and Stiles, B.G. 2013. The staphylococcal enterotoxin (SE) family: sEB and siblings. Virulence 4(8), 759–773. Kulhankova, K., Kinney, K.J., Stach, J.M., Gourronc, F.A., Grumbach, I.M., Klingelhutz, A.J. and Salgado-Pabón, W. 2018. The superantigen toxic shock syndrome toxin 1 alters human aortic endothelial cell function. Infect. Immun. 86(3), e00848-17. Kumar, K., Gill, J., Arora, S. and Khanna, S. 2016. Prevalence of Multidrug-resistant, Extensively Drug-resistant, and Pandrug-resistant Pseudomonas aeruginosa from a Tertiary Level Intensive Care Unit. J. Glob. Infect. Dis. 8(4), 155–159. Laham, N.A., Mediavilla, J.R., Chen, L., Abdelateef, N., Elamreen, F.A., Ginocchio, C.C., Pierard, D., Becker, K. and Kreiswirth, B.N. 2015. MRSA clonal complex 22 strains harboring toxic shock syndrome toxin (TSST-1) are endemic in the primary hospital in Gaza, Palestine. PLos One 10(3), 120008. Martin, S.W., Meek, A.H. and Willeberg, P. 1987. Veterinary epidemiology: principles and methods. Ames: Iowa State University Press. Motamedifar, M., Ebrahim-Saraie, H.S., Alfatemi, S.M., Zalipour, M., Kaveh, M. and Khoshkharam-Roodmajani, H. 2015. Frequency of the toxic shock syndrome toxin-1 gene in methicillin-suspectible and methicillin-resistant Staphylococcus aureus isolates from teaching hospitals in Shiraz, Iran. J. Braz. Soc. Trop. Med. 48(1), 90–93. Mrochen, D.M., Fernandes De Oliveira, L.M., Raafat, D. and Holtfreter, S. 2020. Staphylococcus aureus host tropism and its implications for murine infection models. Int. J. Mol. Sci. 21(19), 7061. Mustapha, M., Kolo, Y.M.B., Geidam, Y.A. and Gulani, I.A.G. 2014. Review on Methicillin-resistant Staphylococcus aureus (MRSA) in dogs and cats. J. Anim. Vet. Adv. 6(2), 61–73. Nagao, M., Okamoto, A., Yamada, K., Hasegawa, T., Hasegawa, Y. and Ohta, M. 2009. Variations in amount of TSST-1 produced by clinical methicillin resistant Staphylococcus aureus (MRSA) isolates and allelic variation in accessory gene regulator (agr) locus. BMC Microbiol. 9(1), 52. Nandhini, P., Kumar, P., Mickymaray, S., Alothaim, A.S., Somasundaram, J. and Rajan, M. 2022. Recent developments in methicillin-resistant Staphylococcus aureus (MRSA) treatment: a review. Antibiotics (Basel). 11(5), 606. Ortega, E., Abriouel, H., Lucas, R. and Gálvez, A. 2010. Multiple roles of Staphylococcus aureus enterotoxins: pathogenicity, superantigenic activity, and correlation to antibiotic resistance. Toxins (Basel). 2(8), 2117–2131. Otto, M. 2014. Staphylococcus aureus toxins. Curr. Opin. Microbiol. 17(1), 32–37. Pantosti, A. 2012. Methicillin-resistant Staphylococcus aureus associated with animals and its relevance to human health. Front. Microbiol. 3(1), 127. Pinchuk, I.V., Beswick, E.J. and Reyes, V.E. 2010. Staphylococcal enterotoxins. Toxins 2(8), 2177–2197. Putra, G.D.S., Khairullah, A.R., Effendi, M.H., Lazuardi, M., Kurniawan, S.C., Afnani, D.A., Silaen, O.S.M., Waruwu, Y.K.K., Millannia, S.K., Widodo, A., Ramadhani, S., Farizqi, M.T.I. and Riwu, K.H.P. 2023. Detection of multidrug-resistant Staphylococcus aureus isolated from dairy's milk in Medowo Village of Kediri District, Indonesia. Biodiversitas 24(1), 423–430. Ramadhani, S., Khairullah, A.R., Effendi, M.H., Sukmanadi, M., Tyasningsih, W., Millannia, S.K., Afnani, D.A., Moses, I.B., Farizqi, M.T.I. and Kaben, S.E. 2024. Detection of the methicillin-resistant Staphylococcus aureus gene encoding mecA in nasal swabs of cats in Surabaya, Indonesia. Open Vet. J. 14(11), 3026–3036. Ramandinianto, S.C., Khairullah, A.R., Effendi, M.H. and Hestiana, E.P. 2020a. Profile of multidrug resistance (MDR) and methicillin resistant Staphylococcus aureus (MRSA) on dairy farms in east Java province, Indonesia. Indian J. Forensic Med. Toxicol. 14(4), 3439–3445. Ramandinianto, S.C., Khairullah, A.R., Effendi, M.H., Tyasningsih, W. and Rahmahani, J. 2020b. Detection of enterotoxin type B gene on methicillin resistant Staphylococcus aureus (MRSA) isolated from raw milk in East Java, Indonesia. Syst. Rev. Pharm. 11(7), 290–298. Rehab, M.E., Dina, E.R. and Shaymaa, H.A.R. 2016. Toxin gene profile and antibiotic resistance of Staphylococcus aureus isolated from clinical and food samples in Egypt. Afr. J. Microbiol. Res. 10(13), 428–437. Schlievert, P.M. and Davis, C.C. 2020. Device-associated menstrual toxic shock syndrome. Clin. Microbiol. Rev. 33(3), e00032. Sharma, H., Turner, C.E., Siggins, M.K., El-Bahrawy, M., Pichon, B., Kearns, A. and Sriskandan, S. 2019. Toxic shock syndrome toxin 1 evaluation and antibiotic impact in a transgenic model of Staphylococcal soft tissue infection. mSphere 4(5), e00665-19. Spaulding, A.R., Salgado-Pabón, W., Kohler, P.L., Horswill, A.R., Leung, D.Y.M. and Schlievert, P.M. 2013. Staphylococcal and streptococcal superantigen exotoxins. Clin. Microbiol. Rev. 26(3), 422–447. Tong, S.Y.C., Davis, J.S., Eichenberger, E., Holland, T.L. and Fowler, V.G. 2015. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 28(3), 603–661. Tyasningsih, W., Effendi, M.H., Budiarto, B. and Syahputra, I.R. 2019. Antibiotic resistance to Staphylococcus aureus and methicillin resistant Staphylococcus aureus (MRSA) isolated from dairy farms in Surabaya, Indonesia. Indian Vet. J. 96(11), 27–31. Waruwu, Y.K.K., Khairullah, A.R., Effendi, M.H., Lukiswanto, B.S., Afnani, D.A., Kurniawan, S.C., Silaen, O.S.M., Riwu, K.H.P., Widodo, A. and Ramandinianto, S.C. 2023. Detection of methicillin-resistant Staphylococcus aureus and multidrug resistance isolated from cats in animal clinic at Sidoarjo District, East Java, Indonesia. Biodiversitas 24(1), 106–111. Yarwood, J.M. and Schlievert, P.M. 2000. Oxygen and carbon dioxide regulation of toxic shock syndrome toxin 1 production by Staphylococcus aureus MN8. J. Clin. Microbiol. 38(5), 1797–1803. Yunita, M.N., Effendi, M.H., Rahmaniar, R.P., Arifah, S. and Yanestria, S.M. 2020. Identification of spa gene for strain typing of methicillin-resistant Staphylococcus aureus (MRSA) isolated from nasal swab of dogs. Biochem. Arch. 20(1), 2999–3004. Zarei Koosha, R., Mahmoodzadeh Hosseini, H., Mehdizadeh Aghdam, E., Ghorbani Tajandareh, S. and Imani Fooladi, A.A. 2016. Distribution of tsst-1 and mecA genes in Staphylococcus aureus isolated from clinical specimens. Jundishapur J. Microbiol. 9(3), e29057. Zhang, X., Hu, X. and Rao, X. 2017. Apoptosis induced by Staphylococcus aureus toxins. Microbiol. Res. 205(1), 19–24. Zhao, H., Xu, S., Yang, H., He, C., Xu, X., Hu, F., Shu, W., Gong, F., Zhang, C. and Liu, Q. 2019. Molecular typing and variations in amount of tst gene expression of TSST-1-producing clinical Staphylococcus aureus isolates. Front. Microbiol. 10(1), 1388. Zheng, Y., Qin, C., Zhang, X., Zhu, Y., Li, A., Wang, M., Tang, Y., Kreiswirth, B.N., Chen, L., Zhang, H. and Du, H. 2020. The tst gene associated Staphylococcus aureus pathogenicity island facilitates its pathogenesis by promoting the secretion of inflammatory cytokines and inducing immune suppression. Microb. Pathog. 138(1), 103797. Zschöck, M., Botzler, D., Blöcher, S., Sommerhäuser, J. and Hamann, H.P. 2000. Detection of genes for enterotoxins (ent) and toxic shock syndrome toxin-1 (tst) in mammary isolates of Staphylococcus aureus by polymerase-chain-reaction. Int. Dairy J. 10(8), 569–574. | ||
| How to Cite this Article |
| Pubmed Style Afnani DA, Khairullah AR, Effendi MH, Tyasningsih W, Tang JYH, Budiastuti B, Rehman S, Moses IB, Kurniasih DAA, Yanestria SM, Ahmad RZ, Aliyah SH, Agustin ALD, Riwu KHP. The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Vet. J.. 2025; 15(10): 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 Web Style Afnani DA, Khairullah AR, Effendi MH, Tyasningsih W, Tang JYH, Budiastuti B, Rehman S, Moses IB, Kurniasih DAA, Yanestria SM, Ahmad RZ, Aliyah SH, Agustin ALD, Riwu KHP. The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. https://www.openveterinaryjournal.com/?mno=251997 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i10.34 AMA (American Medical Association) Style Afnani DA, Khairullah AR, Effendi MH, Tyasningsih W, Tang JYH, Budiastuti B, Rehman S, Moses IB, Kurniasih DAA, Yanestria SM, Ahmad RZ, Aliyah SH, Agustin ALD, Riwu KHP. The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Vet. J.. 2025; 15(10): 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 Vancouver/ICMJE Style Afnani DA, Khairullah AR, Effendi MH, Tyasningsih W, Tang JYH, Budiastuti B, Rehman S, Moses IB, Kurniasih DAA, Yanestria SM, Ahmad RZ, Aliyah SH, Agustin ALD, Riwu KHP. The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Vet. J.. (2025), [cited January 25, 2026]; 15(10): 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 Harvard Style Afnani, D. A., Khairullah, . A. R., Effendi, . M. H., Tyasningsih, . W., Tang, . J. Y. H., Budiastuti, . B., Rehman, . S., Moses, . I. B., Kurniasih, . D. A. A., Yanestria, . S. M., Ahmad, . R. Z., Aliyah, . S. H., Agustin, . A. L. D. & Riwu, . K. H. P. (2025) The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Vet. J., 15 (10), 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 Turabian Style Afnani, Daniah Ashri, Aswin Rafif Khairullah, Mustofa Helmi Effendi, Wiwiek Tyasningsih, John Yew Huat Tang, Budiastuti Budiastuti, Saifur Rehman, Ikechukwu Benjamin Moses, Dea Anita Ariani Kurniasih, Sheila Marty Yanestria, Riza Zainuddin Ahmad, Siti Hamidatul Aliyah, Alfiana Laili Dwi Agustin, and Katty Hendriana Priscilia Riwu. 2025. The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Veterinary Journal, 15 (10), 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 Chicago Style Afnani, Daniah Ashri, Aswin Rafif Khairullah, Mustofa Helmi Effendi, Wiwiek Tyasningsih, John Yew Huat Tang, Budiastuti Budiastuti, Saifur Rehman, Ikechukwu Benjamin Moses, Dea Anita Ariani Kurniasih, Sheila Marty Yanestria, Riza Zainuddin Ahmad, Siti Hamidatul Aliyah, Alfiana Laili Dwi Agustin, and Katty Hendriana Priscilia Riwu. "The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia." Open Veterinary Journal 15 (2025), 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 MLA (The Modern Language Association) Style Afnani, Daniah Ashri, Aswin Rafif Khairullah, Mustofa Helmi Effendi, Wiwiek Tyasningsih, John Yew Huat Tang, Budiastuti Budiastuti, Saifur Rehman, Ikechukwu Benjamin Moses, Dea Anita Ariani Kurniasih, Sheila Marty Yanestria, Riza Zainuddin Ahmad, Siti Hamidatul Aliyah, Alfiana Laili Dwi Agustin, and Katty Hendriana Priscilia Riwu. "The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia." Open Veterinary Journal 15.10 (2025), 5175-5182. Print. doi:10.5455/OVJ.2025.v15.i10.34 APA (American Psychological Association) Style Afnani, D. A., Khairullah, . A. R., Effendi, . M. H., Tyasningsih, . W., Tang, . J. Y. H., Budiastuti, . B., Rehman, . S., Moses, . I. B., Kurniasih, . D. A. A., Yanestria, . S. M., Ahmad, . R. Z., Aliyah, . S. H., Agustin, . A. L. D. & Riwu, . K. H. P. (2025) The molecular evidence of toxic shock syndrome toxin-1 on methicillinresistant Staphylococcus aureus isolated from cats in Surabaya, Indonesia. Open Veterinary Journal, 15 (10), 5175-5182. doi:10.5455/OVJ.2025.v15.i10.34 |