Open Veterinary Journal, (2026), Vol. 16(3): 1636-1645
Research Article
10.5455/OVJ.2026.v16.i3.21
Rapid visual detection of Staphylococcus epidermidis in bovine mastitis milk using LAMP–lateral flow dipstick
Mingfeng Chu1†, Xiaoyu Chong1†, Wei Cheng1, Huiying Zhang1, Yuchen Liang1, Mengke Si1, Yiwei Wang1,
Junxue Qiu1, Baolei Yang1, Xuelong Chen1,2*, Yanping Qi1,2* and Haixia Wang3
1Anhui Province Key Laboratory of Animal Nutritional Regulation and Health, Anhui Science and Technology University, Fengyang, China
2Anhui Engineering Technology Research Center of Pork Quality Control and Enhance, Anhui Science and Technology University, Fengyang, China
3Daqing Agricultural and rural Bureau, Daqing, Heilongjiang, China
*Corresponding Author: Yanping Qi. Anhui Province Key Laboratory of Animal Nutritional Regulation and Health, Anhui Science and Technology University, Fengyang, China. Email: qiyanping2018 [at] vip.163.com
Submitted: 08/10/2025 Revised: 15/02/2026 Accepted: 27/02/2026 Published: 31/03/2026
© 2026 Open Veterinary Journal
This is an Open Access article distributed under the terms of the Creative Commons Attribution-Non Commercial-No Derivatives License (http://creativecommons.org/licenses/by-nc-nd/4.0/), which permits non-commercial re-use, distribution, and reproduction in any medium, provided the original work is properly cited, and is not altered, transformed, or built upon in any way.
Abstract
Background: The use of Loop-mediated Isothermal Amplification (LAMP) technology in combination with Lateral Flow Dipstick (LFD) for the rapid detection of Staphylococcus epidermidis (S. epidermidis) offers a technical solution for early disease monitoring in dairy farms.
Aim: This method is intended to enable early and accurate identification of S. epidermidis infections in dairy farms, facilitating timely intervention and treatment to reduce economic losses and ensure milk quality and safety.
Methods: A pair of primers specific to the aaa gene of S. epidermidis was designed, and the aaa-FIP primer was 5'-end labeled with biotin for LAMP amplification, followed by hybridization with a 6-FAM-labeled probe, and the result was read out on an LFD. The specificity, sensitivity, and clinical applicability of the method were evaluated.
Results: The results demonstrated that the optimal reaction conditions consisted of incubation at 64°C for 30 minutes, followed by probe hybridization for 5 minutes and subsequent chromogenic reaction on the LFD, which enabled the specific and visual detection of S. epidermidis. The established Loop-mediated Isothermal Amplification-Lateral Flow Dipstick (LAMP-LFD) method, which exhibits high specificity and sensitivity with a detection limit 100-fold higher than that of conventional polymerase chain reaction, achieved 100% consistency when applied to 20 clinical samples suspected of S. epidermidis infection.
Conclusion: This study pioneers the development of a LAMP-LFD method that is simple to operate and can visually detect S. epidermidis associated with bovine mastitis with high specificity and sensitivity, making it suitable for rapid on-site detection of early infections in grassroots farms and holding great promise for broad application.
Keywords: Mastitis in dairy cows, Loop-mediated isothermal amplification, Lateral flow dipstick, Rapid test, Staphylococcus epidermidis.
Introduction
Mastitis, the most prevalent disease in dairy cows, caused by an inflammatory reaction of the mammary gland due to bacterial infection, can be triggered by over 20 species of pathogenic bacteria, with Staphylococcus epidermidis (S. epidermidis) emerging as one of the most common coagulase-negative staphylococci associated with bovine mastitis (Grzeczka and Niewitecki, 2021; Velasco Bolaños et al., 2021; Costa et al., 2024; Cho et al., 2025). Currently, there is still a lack of effective therapeutic agents for S. epidermidis infections, and early detection and prevention remain the key strategies for controlling the disease (Manandhar et al., 2021; Serra et al., 2023). Therefore, the establishment of rapid and accurate detection methods for S. epidermidis is of great significance for the sustainable development of bovine mastitis control.
Traditional detection techniques like pathogen isolation and culture are laborious, time-consuming, and technically difficult, while real-time fluorescence quantitative PCR is also challenging to apply directly in the field due to its reliance on expensive instruments and the need for skilled operators(Hiitiö et al., 2018). Notomi et al. (2000) developed the loop-mediated isothermal amplification (LAMP) technology, which can rapidly and efficiently amplify the target nucleic acid sequences under a constant temperature of 60 °C–65°C by meticulously designing 4–6 specific primers. LAMP has become a powerful tool for rapid and reliable on-site detection of pathogens, thanks to its high sensitivity and specificity, low cost, and ease of operation(Velders et al., 2018).
Loop-mediated Isothermal Amplification-Lateral Flow Dipstick (LAMP-LFD) is a method that combines LAMP amplification detection technology with lateral flow test strips, which interprets LAMP results by detecting a colorimetric change on the strip (Kong et al., 2017). With no requirement for sophisticated equipment or toxic dyes, the entire LAMP detection can be completed within 5–10 minute using this method, highlighting its considerable application potential (Bian et al., 2022; Wang et al., 2025). Currently, LAMP-LFD detection technology has been applied to the detection of various pathogens, including orf virus (Yang et al., 2022), Listeria monocytogenes in raw chicken meat (Wachiralurpan et al., 2017), avian adenovirus serotype 4 (Zhai et al., 2019), Mycoplasma hyopneumoniae (Sun et al., 2020), Staphylococcus aureus isolated from bovine mastitis samples (Sheet et al., 2016), and Streptococcus uberis in milk (Cornelissen et al., 2016), and Streptococcus agalactiae in bovine milk (Bosward et al., 2016). In this study, we employed the LAMP-LFD technique to design primers targeting the aaa gene of S. epidermidis, thereby establishing a rapid, specific, sensitive, and practically applicable detection method. This method provides technical support for the early diagnosis of S. epidermidis-associated bovine mastitis.
Materials and Methods
The strains tested in this study
All strains were isolated from quarter-milk samples of Holstein cows with clinical mastitis at Heping Dairy Farm, Bengbu, Anhui, China, from September 2021 to May 2024. Klebsiella pneumoniae, Escherichia coli, Serratia marcescens, Staphylococcus chromogenes, S. agalactiae, Staphylococcus simulans, Staphylococcus haemolyticus, and 20 presumptive S. epidermidis isolates recovered from bovine mastitis milk were all maintained in our laboratory.
Experimental materials
The Bst DNA polymerase and dNTP mixture were purchased from Beijing Baozhi Physics Technology Co., Ltd., Beijing, China. SYBR Green I nucleic acid dye and Super Red nucleic acid dye were obtained from Biosharp, Hefei, Anhui, China, and betaine dry powder was sourced from Merck Chemical Technology (Shanghai) Co., Ltd., Shanghai, China. The DL 2,000 DNA Marker was purchased from Beijing Polymerization Biotechnology Co., Ltd., Beijing, China.
Bacterial culture and DNA extraction
After centrifuging the milk samples from cows with mastitis, the sediment was evenly spread on Columbia blood agar plates, incubated at 37°C in an inverted position for 18–24 hours, and then single colonies were purified, Gram-stained, and examined under a microscope for preliminary identification of suspected bacterial species. A 1.5 ml aliquot of the bacterial suspension was centrifuged at 12,000 rpm for 10 minutes, the supernatant was discarded, the resulting bacterial pellet was resuspended in 100 μl of ddH₂O, incubated in a boiling water bath at 100°C for 10 minutes, immediately transferred to an ice bath for 5 minutes to cool, subjected to a second centrifugation under the same conditions (12,000 rpm, 10 minutes), and the final supernatant was carefully transferred to a new sterile microcentrifuge tube for subsequent experiments. The concentration and purity (A260/A280 ratio) of the extracted DNA were determined using a NanoDrop UV spectrophotometer.
16S rRNA sequencing
To ensure the accuracy of the bacterial strain used in this study, 16S rRNA sequencing was performed for species identification (note: the 16S rRNA is only used for confirming the bacterial species, while the aaa gene is the actual functional target gene of subsequent experiments). Genomic DNA was extracted from purified single bacterial colonies using the TIANGEN Bacterial Genomic DNA Extraction Kit strictly following the manufacturer’s instructions. The full-length 16S rRNA gene was amplified by PCR with universal primers (e.g., 27F: 5'-AGAGTTTGATCMTGGCTCAG-3' and 1492R: 5'-GGTTACCTTGTTACGACTT-3') in a 50 μl reaction system (containing 25 μl 2 × Taq PCR Master Mix, 2 μl genomic DNA template, 2 μl of each primer, and 19 μl sterile ddH₂O) with the following program: pre-denaturation at 95°C for 5 minutes, 35 cycles of 95°C for 30 seconds, 55°C for 30 seconds, 72°C for 1.5 minutes, and final extension at 72°C for 10 minutes. The PCR products were verified by 1.5% agarose gel electrophoresis, purified using a PCR purification kit, and then sent to a commercial sequencing company for bidirectional Sanger sequencing. The obtained 16S rRNA sequences were edited, aligned with the reference sequences in the National Center for Biotechnology Information (NCBI) GenBank database using BLAST, and subjected to phylogenetic analysis to confirm the species identity as S. epidermidis, which ensured the reliability of the subsequent experiments targeting the aaa gene of S. epidermidis.
Primer and probes
One set of specific LAMP primers targeting the aaa gene of S. epidermidis was designed using PrimerExplorer V5 software, and the detailed sequences are shown in Table 1. These primers include the forward inner primer (FIP), reverse inner primer (BIP), forward outer primer (F3), and reverse outer primer (B3). The BLAST tool from the NCBI was employed to align the synthesized primers against the nt database in GenBank. All primers were synthesized by Sangon Biotech (Shanghai) Co., Ltd.
Table 1. Sequences of primers and probe used in this study.

Establishment and optimization of the LAMP reaction system
The 25 μl reaction system consisted of 2.5 μl of 10 × buffer, 6 mM MgSO₄, 1.4 mM dNTPs, 1.6 μM FIP, 1.6 μM BIP, 0.2 μM F3, 0.2 μM B3, 1.6 U/μl Bst DNA polymerase, 1 μl of S. epidermidis DNA template, and 11.5 μl of ddH₂O. The components were sequentially added to a 200 μl centrifuge tube. The reaction mixture was incubated in a 60°C water bath for 50 minutes, followed by heat inactivation at 80°C for 5 minutes. The reaction products were subsequently verified by 1.5% agarose gel electrophoresis.
Subsequently, the concentration of Mg²⁺ was tested in the range of 2–14 mmol/l, with an increment of 2 mmol/l for each LAMP reaction. Agarose gel electrophoresis was employed to detect the amplification products, thereby determining the optimal Mg²⁺ concentration. Building on the previously optimized reaction system, the concentration of betaine in the reaction was further optimized. The concentration of betaine was tested in the range of 0.2–1.0 mmol/l, with an increment of 0.2 mmol/l for each LAMP reaction. The concentration of the dNTP mixture was tested in the range of 0.2–1.4 mmol/l, with an increment of 0.2 mmol/l for each LAMP reaction. The reaction temperature was tested at 1°C intervals between 60°C and 65°C, while the reaction time was set from 1–60 minute, with intervals of 5 minutes. The optimal reaction temperature and time were determined by 1.5% agarose gel electrophoresis, aiming to achieve the most distinct electrophoretic bands.
Establishment of LAMP-LFD
The optimized LAMP reaction system, incorporating biotin-labeled FIP primers and 6-FAM-labeled probes, was subjected to hybridization at 65°C for 5 minutes, followed by heat treatment at 80°C for 5 minutes. Subsequently, 5 µl of the reaction product was diluted with 90 µl of ddH₂O, and the diluted sample was applied to the sample zone of the LFD for detection. During migration to the sample binding pad, test line, and control line, biotin and 6-FAM-labeled DNA probes effectively bind to streptavidin-coated gold nanoparticles on the sample binding pad. If amplification of the biotin 6-FAM probe of the target gene occurs, its product will be captured by the 6-FAM antibody to form a ternary complex, causing the detection line to show a red band, and conversely, the biotin- and 6-FAM-labeled probe, which does not form a hybridization product, migrates past the test line and binds to the biotinylated bovine serum albumin at the quality control line, showing a red band only at the quality control line.
Analysis of specificity, sensitivity, and repeatability of LAMP-LFD
To verify the specificity of the LAMP-LFD assay, S. epidermidis, K. pneumoniae, E. coli, S. marcescens, S. chromogenes, S. agalactiae, S. simulans, S. hemolyticus, were selected. Amplification was performed using the optimized LAMP protocol. The amplified products were then detected by both LFD and 1.5% agarose gel electrophoresis.
The genomic DNA samples of bovine-derived S. epidermidis were serially10-fold diluted from 1.93 × 101 ng/ul–1.93 × 10-5 ng/ul for evaluating thesensitivity of the LAMP-LFD method. LAMP amplification reactions were then performed using this dilution series as templates, and the amplification products were detected using both LFD and 1.5% agarose gel electrophoresis.
Similarly, genomic DNA at various dilutions was used as the template, and the outer primers F3 and B3 of S. epidermidis were employed as specific PCR primers for the reaction. The PCR reaction system consisted of a 12 µl mixture, including 2 × Es Taq premix (with dye), 2 μl of outer primers, 1 μl of template DNA, and 8 μl of ddH₂O. The cycling conditions were as follows: initial denaturation at 94°C for 2 minutes, followed by 30 cycles of 94°C for 30 seconds (denaturation), 56°C for 30 seconds (annealing), and 72°C for 30 seconds (extension), with a final extension at 72°C for 2 minutes. The amplified products were detected using 1.5% agarose gel electrophoresis.
The reproducibility of the method was evaluated with triplicate samples. First, S. epidermidis DNA was serially diluted with ddH₂O to the minimum concentration required for LAMP amplification and then used as the template. These samples were then amplified using the optimized LAMP reaction system. Finally, the amplified products in the three groups of samples were detected by both LFD and 1.5% agarose gel electrophoresis.
The detection of clinically isolated strains by LAMP-LFD
Genomic DNA was extracted from 20 clinically suspected S. epidermidis samples, which were initially identified as presumptive S. epidermidis isolates based on a combination of clinical symptom screening and preliminary phenotypic characterization—these isolates were recovered from quarter-milk samples of Holstein cows with clinical mastitis and exhibited Gram-positive cocci arranged in clusters by Gram staining—followed by separate LAMP-LFD and PCR assays with negative controls (sterile double-distilled water) and positive controls included in the experiments.
Statistical analysis
All experiments included 3 biological and 3 technical replicates. Data were analyzed using SPSS 26.0, and graphs were plotted with GraphPad Prism 9.0. Specificity, positive predictive value (PPV), negative predictive value (NPV), and accuracy were calculated with 95% confidence intervals (95%CI) via the Wilson score method. Sensitivity was evaluated by the coefficient of variation (CV) at the limit of detection (CV<10% for stability). Repeatability was assessed by one-way ANOVA and intraclass correlation coefficient (ICC≥0.8 for excellent repeatability). Kappa test and McNemar test were used to analyze the agreement and difference between LAMP-LFD and conventional PCR.
Ethical approval
Not needed for this study.
Results
Optimization of the LAMP reaction system
During optimization of the LAMP reaction system, the Mg²⁺ concentration was varied from 0–14 mmol/l, with the most distinct band observed at 10 mmol/l (Fig. 1A). When the final concentration of betaine was 0.2–1.0 mmol/l, the reaction products exhibited bright ladder-like bands, with the most distinct bands observed at 0.8 mmol/l (Fig. 1B). The concentration of dNTPs was optimized. Amplification of the target sequence was achieved within a dNTP concentration range of 0.2–1.4 mmol/l (Fig. 1C), with the optimal amplification efficiency and the clearest target band observed at a final dNTP concentration of 1 mmol/l. Positive bands were detectable across the reaction temperature range of 60°C–65°C, with the most intense band polymorphism observed at 64°C (Fig. 1D). Finally, the optimized reaction time results showed that distinct bands could be generated when the reaction time exceeded 30 minutes, and no significant differences in band intensity were observed across different reaction temperatures (Fig. 1E).
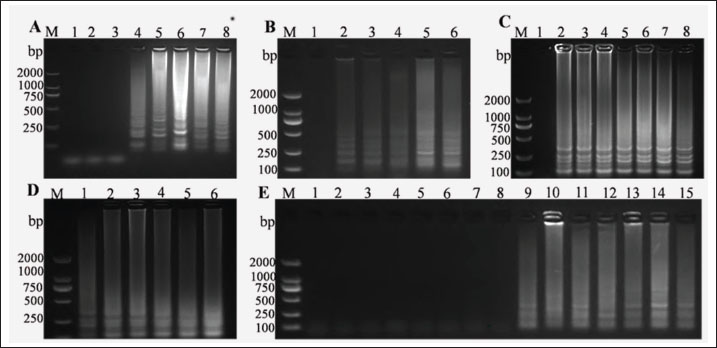
Fig. 1. Optimization of the LAMP reaction. (A) Optimization of Mg²⁺ content, M: DNA Maker DL2000, Numbers 1–8 represent the negative control, 2, 4, 6, 8, 10,12 and 14 mmol/l respectively. (B) Optimization of betaine concentration, M: DNA Maker DL2000, Numbers 1–6 represent the negative control, 0.2, 0.4, 0.6, 0.8, and 1.0 mmol/l respectively. (C) Optimization of dNTPs concentration, M: DNA Maker DL2000, Numbers 1–8 represent the negative control, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, and 1.4 mmol/l respectively.(D) Optimization of reaction temperature, M: DNA Maker DL2000, Numbers 1–6 represent 60°C, 61°C, 62°C, 63°C, 64°C, and 65°C respectively. (E) Optimization of reaction time , M: DNA Maker DL2000, Numbers 1–15 represent the negative control, 1, 3, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, and 60 minutes respectively.
Establishment of LAMP-LFD
Positive samples display a red band on the test line and a blue band on the control line, whereas negative samples show only a blue band on the control line (Fig. 2), indicating successful establishment of the LAMP-LFD assay.

Fig. 2. Establishment of the LAMP-LFD for S. epidermidis, lane 1: sample detection, lane 2: Negative control.
Specificity of LAMP-LFD
The established LAMP-LFD detection method was used to detect S. epidermidis, K. pneumoniae, E. coli, S. marcescens, S. chromogenes, S. agalactiae, S. simulans, and S. hemolyticus (Fig. 3). When S. epidermidis DNA was used as a template, the control line on the horizontal gel strip showed a blue band, the test line showed a red band, and the electrophoresis results displayed typical trapezoidal bands, indicating a positive result. In contrast, no electrophoresis bands were observed when ddH₂O and other bacterial DNA samples were used as templates. Additionally, no red bands were observed on the detection line of the horizontal chromatography test strip, indicating a negative test result. These results demonstrate that the established LAMP-LFD has good specificity.
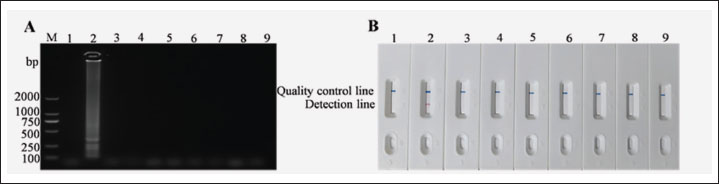

Fig. 3. Specific analysis of LAMP-LFD for S. epidermidis. (A) Detection of agarose gel electrophoresis. (B) Detection by LAMP-LFD, Numbers 1-9 represent the negative control, S. epidermidis, K. pneumoniae, E. coli, S. marcescens, S. chromogenes, S. agalactiae, S. simulans, S. hemolyticus respectively.
Sensitivity of LAMP-LFD
To assess the sensitivity of the LAMP-LFD method developed in this study, S. epidermidis DNA was serially diluted 10-fold from 1.93 × 101–1.93 × 10⁻5 ng/µl (Fig. 4). The lowest detection limit of the LAMP-LFD method was found to be 10⁻4 ng/µl. In contrast, using the same DNA concentrations for PCR detection (Fig. 4), the lowest detection limit was 10⁻2 ng/µl. This indicates that the LAMP-LFD method is 100-fold more sensitive than PCR for the detection of S. epidermidis DNA. Therefore, the LAMP-LFD method established in this study can be used for the detection of S. epidermidis.
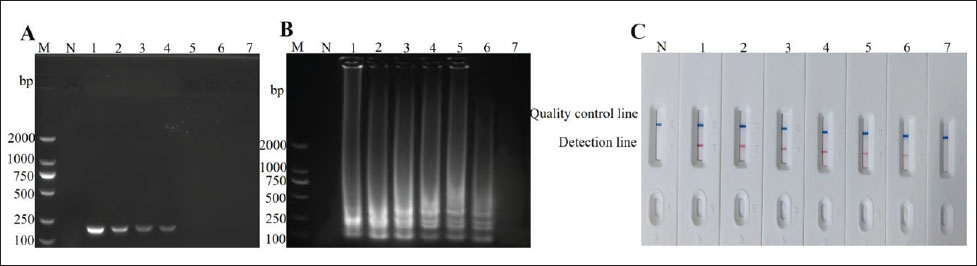
Fig. 4. Detection of sensitivity of LAMP-LFD for S. epidermidis. (A) Sensitivity of PCR. (B) Sensitivity of LAMP-AGE. M, DNA Marker DL2000, 1–7 represent1.93 × 101 –1.93×10−5ng/μl. (C) Sensitivity of LAMP-LFD.
Repeatability of LAMP-LFD
In total, three samples of S. epidermidis DNA were separately diluted to 10⁻4 ng/µl and used to perform LAMP assays. The amplified products were detected using LFD and electrophoresis. Positive results were obtained for the three parallel reactions (Fig. 5), indicating that LAMP-LFD was repeatable.
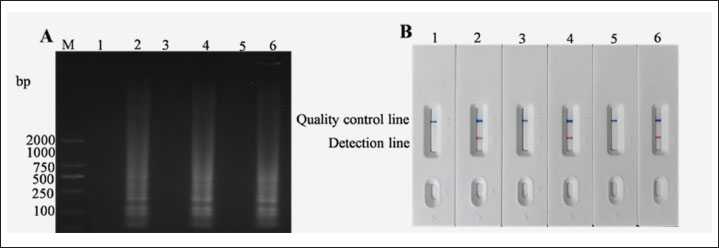
Fig. 5. Test of repeatability of LAMP-LFD for S. epidermidis. (A) Test of LAMP agarose gel electrophoresis repeatability. (B) Repeatability detection by LAMP-LFD, M: DNA Marker DL2000, lane 1: Negative control, lane 2: S. epidermidis DNA, lane 3: Negative control, lane 4: S. epidermidis DNA, lane 5: Negative control, lane 6: S. epidermidis DNA.
Clinical sample detection by LAMP-LFD
In this study, 20 clinical samples from suspected S. epidermidis infections were tested using both the LAMP-LFD method and the PCR method. The results showed that the detection results of the two methods were consistent, with 7 positive samples detected by each method (Fig. 6). The concordance rate of detection between the two methods reached 100%, which fully indicates that the LAMP-LFD method established in this study has the potential to be a fast and reliable tool for the detection of S. epidermidis.
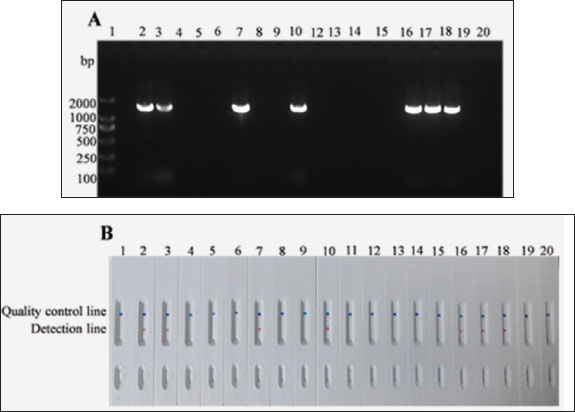
Fig. 6. Clinical results S. epidermidis for detection by LAMP-LFD. (A) Sensitivity of PCR. (B) Study on clinical specimen detection by LAMP-LFD,1, Negative control; 2–20. Clinical samples.
Statistical evaluation of sensitivity, specificity, and agreement
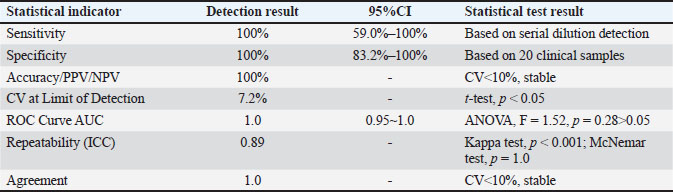
The statistical performance of the LAMP-LFD method is summarized in Table 2. Based on the detection of serial dilutions (down to 1.93 × 10⁻4 ng/µl) and eight non-target bacterial strains, the method demonstrated 100% sensitivity and 100% specificity. The PPV, NPV, and accuracy were also 100%. Due to the limited number of non-target strains tested (n=8), the 95% (95% CI) for specificity ranged from 66.3% to 100%. The CV at the limit of detection was 7.2%, indicating good assay stability. Repeatability, assessed by the ICC from triplicate experiments, was excellent (ICC=0.89). The agreement between LAMP-LFD and conventional PCR for 20 clinical samples was perfect, with a Cohen's Kappa value of 1.0 (p < 0.001). The area under the receiver operating characteristic (ROC) curve, using bacterial culture and sequencing as the reference standard, was 1.0 (95% CI: 0.95–1.0), confirming the high diagnostic accuracy of the method.
Table 2. Statistical evaluation of the LAMP-LFD method for detection of S. epidermidis.
Discussion
In this study, specific LAMP primers targeting the aaa gene of S. epidermidis were designed, and a visual detection method incorporating a probe and LFD was established after optimization of the reaction conditions. The results showed that the entire detection process could be observed with the naked eye in as little as 35 minutes. Zhang et al. (2019) reported a LAMP-LFD method for detecting M. pneumoniae in sheep, with the entire process taking as little as 65 minutes. Wachiralurpan et al. (2017) reported a LAMP-LFD detection method for detecting L. monocytogenes in raw chicken meat, which requires 90 minutes to achieve visual detection. Liu et al. (2023) reported that the LAMP-LFD test for detecting Helicobacter pylori requires 1.5 hours to achieve visual detection. Compared with the aforementioned rapid detection methods, the S. epidermidis LAMP-LFD detection method used in this study is more time-efficient, operates at a constant temperature, does not require expensive equipment, and provides directly visualizable results.
The study demonstrated that the S. epidermidis LAMP-LFD detection method established herein exhibits excellent specificity, with no cross-reactivity with DNA templates from other negative samples. Furthermore, the LAMP-LFD used in this experiment has a detection limit as low as 1.93 × 10⁻4 ng/µl, which is in complete agreement with the results reported by Liu et al. (2023), and its sensitivity is 100 times that of conventional PCR. Yang et al. (2022) reported that the minimum detection concentration of the LAMP-LFD assay for the sheep poxvirus B2L gene was 4.95 × 10−2 copies/μl, which is 1,000 times higher than the minimum detection limit of the PCR assay. Zou et al. (2018) reported that the minimum detection limit for LAMP-LFD detection of the genome DNA of the East China Sea dinoflagellate was 47 pg/μl, which is 10 times higher than that of the PCR detection method. Kiatpathomchai et al. (2008) reported that the minimum detection limit of the LAMP-LFD method for shrimp Taura syndrome virus was 1 × 10‒7 ng/µl, which was 10 times higher than the minimum detection limit of 1 × 10‒6 ng/µl for PCR detection. These studies have all demonstrated that the sensitivity of LAMP-LFD is higher than that of conventional PCR. In addition, LAMP-LFD also has the advantages of a relatively simple operating procedure, no requirement for complex equipment, and a shorter detection time (Wang et al., 2020). The combination of LAMP technology with LFD test strips enables specific hybridization between 6-FAM-labeled probes and biotin on the amplification products, allowing visual detection within a short time while ensuring specificity and sensitivity, and overcoming the drawbacks of other detection methods(Liu et al., 2023).
Despite the promising performance of the established LAMP-LFD assay for detecting S. epidermidis from purified DNA and clinical isolates under controlled laboratory conditions, this study has several limitations that need to be acknowledged and addressed for its practical application: regarding the potential limitations in real milk samples, raw milk is a complex matrix containing various inhibitors such as fats, proteins, and calcium ions, which may bind to nucleic acids or inhibit DNA polymerase activity, thereby interfering with DNA extraction efficiency and the accuracy of subsequent LAMP-LFD detection (Ramesh et al., 2002); another critical concern is the contamination risk in LAMP assays, as isothermal amplification techniques are highly susceptible to aerosol contamination during reaction setup and product handling that could lead to false-positive results, and although strict laboratory zoning (with separate areas for reagent preparation, sample processing, and product detection) and overlaying mineral oil on the reaction mixture were adopted to minimize such risks in the present study, these measures only mitigate rather than completely eliminate contamination risks, underscoring the need for more robust strategies like closed-tube detection formats in future kit development; furthermore, a notable limitation lies in the lack of validation across different farms and regions, since the current validation was based on samples from a limited number of sources, and we have not yet carried out validation experiments on S. epidermidis isolates from bovine mastitis across diverse geographical locations and breeding systems—a gap that may restrict the generalizability of the assay, given the significant variations in S. epidermidis genetic diversity and milk matrix composition across different regions and farming models, and to address this issue, subsequent research will expand the sample collection scope to include multiple pastures covering different geographical regions and breeding models, conducting validation experiments to comprehensively evaluate the method’s applicability and reliability under diverse strain backgrounds and breeding environments, thereby further improving the practical application value of the research results.
There are still some key challenges to be overcome in the research and industrialization of LAMP-LFD technology, especially in the stage of primer design. The high GC content regions in the genome of S. epidermidis are prone to causing primer dimer formation and reducing amplification efficiency, increasing the risk of non-specific binding and affecting the sensitivity and specificity of the LAMP reaction (Velasco-Bolaños et al., 2021). The length and structure of LAMP amplification products may affect their binding efficiency with LFD probes, necessitating the optimization of the concentration and positioning of labeled primers (Ci et al., 2023). Optimizing these technical details is a crucial step for the commercialization of LAMP-LFD. During the industrialization process, the large-scale production of the S. epidermidis detection kit still faces many challenges. For example, the transportation stability of Bst DNA polymerase in high-temperature environments in tropical regions (such as dairy cow breeding areas) is insufficient, necessitating the development of a lyophilized protective agent formulation (Holz et al., 2024). In addition, the actual application scenarios in breeding farms require that the detection methods must adapt to rudimentary conditions. Therefore, it is necessary to develop portable constant-temperature devices to facilitate grassroots detection applications.
Conclusion
In summary, this study has successfully established a LAMP-LFD method for the detection of S. epidermidis, enabling rapid and straightforward identification of the bacteria without the need for costly equipment or reagents, and holds broad application prospects. The entire process can be completed within 35 minutes using only a water bath and test strips, specifically designed for rapid on-site detection of S. epidermidis in bovine mastitis. This approach offers an efficient and convenient new strategy for the detection of the bacterium and holds significant value for promotion.
Acknowledgments
We sincerely thank personnel from the College of Anhui Province Key Laboratory of Animal Nutritional Regulation and Health, Anhui Science and Technology University, and MJE editor (www.mjeditor.com) for their linguistic assistance while preparing this manuscript.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
This work was supported by Major Scientific Research Project of Anhui Provincial Department of Education (2025AHGXZK20143), Anhui Provincial Quality Engineering Project for the New Era of Educating People (Postgraduate Education) (2024cxcysj188), Grass Feeding Livestock Resource Utilization and Health Science and Technology Innovation Team (2023AH010061), Veterinary Science Peak Discipline Project of Anhui Science and Technology University (XK-XJGF002).
Authors' contributions
Conceptualization: Mingfeng Chu, Xiaoyu Chong. Formal analysis: Wei Cheng. Funding acquisition: Xuelong Chen, Yanping Qi. Investigation: Huiying Zhang. Methodology: Yuchen Liang Junxue Qiu Baolei Yang. Project administration: Mengke Si. Resources: Xuelong Chen. Supervision: Yanping Qi. Validation: Yiwei Wang. Writing - original draft: Mingfeng Chu. Writing - review & editing: Xuelong Chen.
Data availability
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
References
Bian, Z., Liu, W., Jin, J., Hao, Y., Jiang, L., Xie, Y. and Zhang, H. 2022. Development of a recombinase polymerase amplification assay with lateral flow dipstick (RPA-LFD) for rapid detection of Shigella spp. and enteroinvasive Escherichia coli. PLos One. 17, 278869.
Bosward, K.L., House, J.K., Deveridge, A., Mathews, K. and Sheehy, P.A. 2016. Development of a loop-mediated isothermal amplification assay for the detection of Streptococcus agalactiae in bovine milk. J. Dairy Sci. 99, 2142–2150.
Cho, J.H., Lee, G.M., Ko, S., Kim, Y. and Kim, D. 2025. Characterization and therapeutic potential of newly isolated bacteriophages against Staphylococcus species in bovine mastitis. J. Virol. 99, 190124.
Ci, L., Gu, Q.L., Xian, S.M., Zhou, P., Liang, Q., Zheng, W. and W, Q. 2023. Establishment and Preliminary Application of a Rapid LAMP-LFD Detection Method for Duck Plague. Casp. J. Vet. Sci. 43, 2050–2055.
Cornelissen, J.B.W.J., De Greeff, A., Heuvelink, A.E., Swarts, M., Smith, H.E. and Van Der Wal, F.J. 2016. Rapid detection of Streptococcus uberis in raw milk by loop-mediated isothermal amplification. J. Dairy. Sci. 99, 4270–4281.
Costa, A., Bovenhuis, H., Egger-Danner, C., Fuerst-Waltl, B., Boutinaud, M., Guinard-Flament, J., Obritzhauser, W., Visentin, G. and Penasa, M. 2024. Mastitis has a cumulative and lasting effect on milk yield and lactose content in dairy cows. J. Dairy Sci. 108, 635–650.
Grzeczka, A. and Niewitecki, W. 2021. Properties of Coagulase-Positive Staphylococcal Cells that Make it Difficult to Diagnose and Treat Mastitis in Cows. Med. J. Cell. Biol. 9, 100–104.
Hiitiö, H., Pyörälä, S., Taponen, S., Rajala-Schultz, P. and Simojoki, H. 2018. Elimination of experimentally induced bovine intramammary infection assessed by multiplex real-time PCR and bacterial culture. J. Dairy. Sci. 101, 5267–5276.
Holz, N., Wax, N., Oest, M. and Fischer, M. 2024. REASSURED Test System for Food Control—Preparation of LAMP Reaction Mixtures for In-Field Identification of Plant and Animal Species. Appl. Sci. 14, 10946.
Kiatpathomchai, W., Jaroenram, W., Arunrut, N., Jitrapakdee, S. and Flegel, T.W. 2008. Shrimp Taura syndrome virus detection by reverse transcription loop-mediated isothermal amplification combined with a lateral flow dipstick. J. Virol. Methods. 153, 214–217.
Kong, C., Wang, Y., Fodjo, E.K., Yang, G.X., Han, F. and Shen, X.S. 2017. Loop-mediated isothermal amplification for visual detection of Vibrio parahaemolyticus using gold nanoparticles. Microchimica Acta 185, 35.
Liu, W., Lu, G., Wang, Y., Chen, Z., Gao, Y., Yin, Z., Wu, Y., Lv, X., Guo, P. and Zhao, Y. 2023. A novel loop-mediated isothermal amplification-lateral flow dipstick method for Helicobacter pylori detection. Front. Microbiol. 14, 1094600.
Manandhar, S., Singh, A., Varma, A., Pandey, S. and Shrivastava, N. 2021. Phenotypic and genotypic characterization of biofilm producing clinical coagulase negative staphylococci from Nepal and their antibiotic susceptibility pattern. Ann. Clin. Microbiol. 20, 41.
Notomi, T. 2000. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, 63.
Ramesh, A., Padmapriya, B.P., Chrashekar, A. and Varadaraj, M.C. 2002. Application of a convenient DNA extraction method and multiplex PCR for the direct detection of Staphylococcus aureus and Yersinia enterocolitica in milk samples. Mol. Cell. Probes. 16, 307–314.
Serra, N., Di Carlo, P., Andriolo, M., Mazzola, G., Diprima, E., Rea, T., Anastasia, A., Fasciana, T. M. A., Pipitò, L., Capra, G., Giammanco, A. and Cascio, A. 2023. Staphylococcus aureus and Coagulase-Negative Staphylococci from Bloodstream Infections: Frequency of Occurrence and Antimicrobial Resistance, 2018-2021. LIFE-BASEL. 13.
Sheet, O.H., Grabowski, N.T., Klein, G. and Abdulmawjood, A. 2016. Development and validation of a loop mediated isothermal amplification (LAMP) assay for the detection of Staphylococcus aureus in bovine mastitis milk samples. Mol. Cell. Probes 30, 320–325.
Sun, P., Peng, Z., Wang, S., Wang, Y., Wu, X., Xu, S. and Li, J. 2020. Establishment and preliminary application of a rapid detection method for porcine mycoplasma pneumonia using LAMP-LFD. J. Vet. Med. 51, 1419–1428.
Velasco-Bolaños, J., Ceballes-Serrano, C.C., Velásquez-Mejía, D., Riaño-Rojas, J.C., Giraldo, C.E., Carmona, J.U. and Ceballos-Márquez, A. 2021. Application of udder surface temperature by infrared thermography for diagnosis of subclinical mastitis in Holstein cows located in tropical highlands. J. Dairy Sci. 104, 10310–10323.
Velders, A.H., Schoen, C. and Saggiomo, V. 2018. Loop-mediated isothermal amplification (LAMP) shield for Arduino DNA detection. BMC Res. Notes 11, 93.
Wachiralurpan, S., Sriyapai, T., Areekit, S., Kaewphinit, T., Sriyapai, P., Santiwatanakul, S. and Chansiri, K. 2017. Development of a rapid screening test for Listeria monocytogenes in raw chicken meat using loop-mediated isothermal amplification (LAMP) and lateral flow dipstick (LFD). Food Anal. Methods 10, 3763–3772.
Wang, H., Dong, C., Tian, X., Pan, Y., Wang, L., An, T. and Zhu, L. 2025. Development and application of a dual LAMP-LFD assay for the simultaneous detection of Streptococcus suis and Glaesserella parasuis. Front. Cell. Infect. Microbiol. 15, 1575365.
Wang, L., Chen, G., Zhang, C., Wang, Y. and Sun, R. 2020. Application of loop-mediated isothermal amplification combined with lateral flow dipstick to rapid and sensitive detection of Alexandrium catenella. Environ. Sci. Pollut. Res. 27, 4246–4257.
Yang, Q., Li, Z., Xia, S., Bao, T., Zhang, Y., Liang, Q. and Gu, Q. 2022. Establishment and preliminary application of a rapid detection method for sheep mouth ulcer virus using LAMP-LFD. Chin. J. Vet. Med. 42, 507–513.
Zhai, X., Mei, X., Wu, X., Zuo, L., Zhou, L., Tian, Y., Han, X., Yang, X. and Wang, H. 2019. A loop-mediated isothermal amplification coupling with a lateral flow dipstick for rapid and specific detection of fowl adenovirus serotype-4. J. Virol. Methods 270, 79–86.
Zhang, J., Cao, J., Zhu, M., Xu, M. and Shi, F. 2019. Loop-mediated isothermal amplification-lateral-flow dipstick (LAMP-LFD) to detect Mycoplasma ovipneumoniae. World. J. Microbiol. 35, 31.
Zou, X., Huang, H., Qiao, L., Dang, C., Pang, J., Lv, P., Wang, J. and Zhu, P. 2018. Rapid detection method for harmful red tide algae Prorocentrum donghaiense based on LAMP-LFD technology. Oceans Lakes 49, 586–593.
















